

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.2 + phase: 0
(53 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
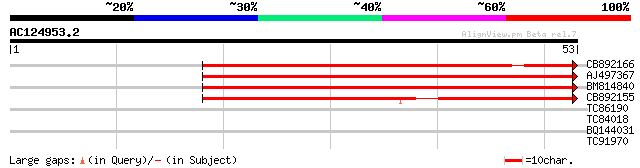
Score E
Sequences producing significant alignments: (bits) Value
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 60 1e-10
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 55 6e-09
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 50 2e-07
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 45 6e-06
TC86190 weakly similar to PIR|T08454|T08454 hypothetical protein... 26 2.8
TC84018 similar to PIR|F86150|F86150 hypothetical protein AAF764... 26 2.8
BQ144031 similar to GP|3132295|dbj| long ORF {TT virus}, partial... 26 3.6
TC91970 similar to GP|6002766|gb|AAF00131.1| class II chitinase ... 25 6.2
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 60.5 bits (145), Expect = 1e-10
Identities = 30/35 (85%), Positives = 32/35 (90%)
Frame = -1
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV+SR GLKIL+ DENGDCIDNTTNVVYK VFQNV
Sbjct: 238 RVSSRNGLKILMIDENGDCIDNTTNVVYK-VFQNV 137
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 543
Score = 55.1 bits (131), Expect = 6e-09
Identities = 24/35 (68%), Positives = 32/35 (90%)
Frame = +1
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTSRKGLKIL+ E+G+C++ T+NVVYKEVF+N+
Sbjct: 52 RVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRNL 156
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 50.1 bits (118), Expect = 2e-07
Identities = 24/35 (68%), Positives = 27/35 (76%)
Frame = +3
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV SR GLKILITDENG +T NVVY+EVFQ +
Sbjct: 303 RVKSRSGLKILITDENGSPSSSTVNVVYQEVFQKI 407
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 45.1 bits (105), Expect = 6e-06
Identities = 23/37 (62%), Positives = 32/37 (86%), Gaps = 2/37 (5%)
Frame = -2
Query: 19 RVTSRKGLKILITDENG--DCIDNTTNVVYKEVFQNV 53
RVTSR+GLKILI++++G DC+ T+NVVY+EVF N+
Sbjct: 253 RVTSREGLKILISNDDGEDDCV--TSNVVYREVFHNL 149
>TC86190 weakly similar to PIR|T08454|T08454 hypothetical protein F22O6.170
- Arabidopsis thaliana, partial (60%)
Length = 1159
Score = 26.2 bits (56), Expect = 2.8
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = +1
Query: 22 SRKGLKILITDENGDCIDNTTNVVYKEVFQN 52
S GLK L+ NG+ I++ T V+ K F++
Sbjct: 661 SEAGLKKLLAFRNGEFIESLTKVMQKGFFES 753
>TC84018 similar to PIR|F86150|F86150 hypothetical protein AAF76467.1
[imported] - Arabidopsis thaliana, partial (38%)
Length = 711
Score = 26.2 bits (56), Expect = 2.8
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = -1
Query: 23 RKGLKILITDENGDCIDNTTN 43
RK L IL D NGD + +TTN
Sbjct: 417 RKNLAILACDFNGDGVPSTTN 355
>BQ144031 similar to GP|3132295|dbj| long ORF {TT virus}, partial (20%)
Length = 1411
Score = 25.8 bits (55), Expect = 3.6
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +2
Query: 1 MLSVCCQQALILFFWADYRVTSRKGL 26
++S+CC +L LFF+ R R GL
Sbjct: 605 VISICCSISLFLFFFRLCRCVIRVGL 682
>TC91970 similar to GP|6002766|gb|AAF00131.1| class II chitinase {Fragaria x
ananassa}, partial (69%)
Length = 681
Score = 25.0 bits (53), Expect = 6.2
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +2
Query: 21 TSRKGLKILITDENGDCIDNTTNVVYKEVFQN 52
TSR+GL+I +E+ DCI + + FQN
Sbjct: 497 TSREGLRI*WAEESRDCIKQ-----FSDCFQN 577
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,755,711
Number of Sequences: 36976
Number of extensions: 14829
Number of successful extensions: 106
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 106
length of query: 53
length of database: 9,014,727
effective HSP length: 29
effective length of query: 24
effective length of database: 7,942,423
effective search space: 190618152
effective search space used: 190618152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 51 (24.3 bits)
Medicago: description of AC124953.2


