

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
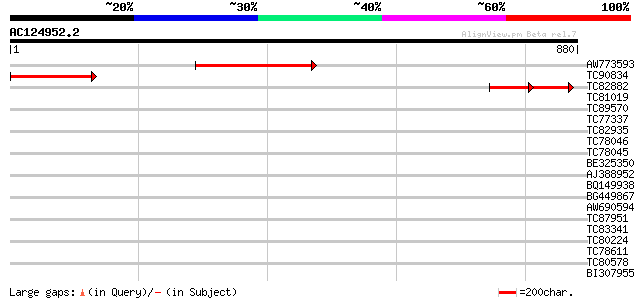
Query= AC124952.2 - phase: 0
(880 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW773593 374 e-103
TC90834 271 8e-73
TC82882 similar to GP|20258832|gb|AAM13898.1 unknown protein {Ar... 142 5e-68
TC81019 similar to GP|5103823|gb|AAD39653.1| Contains PF|00787 P... 37 0.025
TC89570 similar to GP|13936308|gb|AAK40307.1 putative methyl-bin... 37 0.025
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 36 0.073
TC82935 similar to GP|15150635|gb|AAK85442.1 Hypothetical protei... 35 0.096
TC78046 similar to GP|21952848|dbj|BAC06263. putative calreticul... 35 0.13
TC78045 similar to GP|15810563|gb|AAL07169.1 putative calreticul... 35 0.13
BE325350 weakly similar to GP|13702821|gb| putative snRNP protei... 35 0.13
AJ388952 similar to PIR|T04526|T045 hypothetical protein F16A16.... 35 0.16
BQ149938 similar to GP|21166135|gb| hypothetical protein {Dictyo... 34 0.21
BG449867 similar to GP|4127878|emb NDX1 homeobox protein {Glycin... 34 0.21
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 34 0.28
TC87951 similar to PIR|C86418|C86418 hypothetical protein AAG126... 34 0.28
TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A... 33 0.36
TC80224 similar to GP|12324597|gb|AAG52258.1 putative coatomer p... 33 0.36
TC78611 similar to GP|15450599|gb|AAK96571.1 AT5g17440/K3M16_10 ... 33 0.36
TC80578 similar to GP|9759287|dbj|BAB09752.1 arginine-aspartate-... 33 0.36
BI307955 weakly similar to GP|17064958|gb Unknown protein {Arabi... 33 0.62
>AW773593
Length = 567
Score = 374 bits (959), Expect = e-103
Identities = 187/187 (100%), Positives = 187/187 (100%)
Frame = +1
Query: 289 RDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQREN 348
RDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQREN
Sbjct: 7 RDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQREN 186
Query: 349 LDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSST 408
LDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSST
Sbjct: 187 LDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSST 366
Query: 409 STNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPS 468
STNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPS
Sbjct: 367 STNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPS 546
Query: 469 RDGGLTA 475
RDGGLTA
Sbjct: 547 RDGGLTA 567
>TC90834
Length = 488
Score = 271 bits (693), Expect = 8e-73
Identities = 134/134 (100%), Positives = 134/134 (100%)
Frame = +2
Query: 1 MNGSPDPPLDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVT 60
MNGSPDPPLDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVT
Sbjct: 86 MNGSPDPPLDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVT 265
Query: 61 VFHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEE 120
VFHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEE
Sbjct: 266 VFHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEE 445
Query: 121 SDGNENGGEVGERE 134
SDGNENGGEVGERE
Sbjct: 446 SDGNENGGEVGERE 487
>TC82882 similar to GP|20258832|gb|AAM13898.1 unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 526
Score = 142 bits (358), Expect(2) = 5e-68
Identities = 68/68 (100%), Positives = 68/68 (100%)
Frame = +2
Query: 745 LDSIHEHPMLCVTAVNPFLVSKVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRY 804
LDSIHEHPMLCVTAVNPFLVSKVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRY
Sbjct: 8 LDSIHEHPMLCVTAVNPFLVSKVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRY 187
Query: 805 LLESNDFF 812
LLESNDFF
Sbjct: 188 LLESNDFF 211
Score = 135 bits (339), Expect(2) = 5e-68
Identities = 65/69 (94%), Positives = 67/69 (96%)
Frame = +3
Query: 806 LESNDFFALRDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSD 865
L++ FFALRDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSD
Sbjct: 192 LKAMTFFALRDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSD 371
Query: 866 PSSLIFPFQ 874
PSSLIFPFQ
Sbjct: 372 PSSLIFPFQ 398
>TC81019 similar to GP|5103823|gb|AAD39653.1| Contains PF|00787 PX (phox)
domain. {Arabidopsis thaliana}, partial (18%)
Length = 1106
Score = 37.4 bits (85), Expect = 0.025
Identities = 52/199 (26%), Positives = 83/199 (41%), Gaps = 15/199 (7%)
Frame = +3
Query: 415 KSHVISKIEDSE-LSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGL 473
K+H+ ++E S LS +V+ + L S A F R++ G
Sbjct: 18 KTHMWQEVERSSFLSGDGQDVLSSSKSHLNSDESSDDADFERSGRIYS----------GA 167
Query: 474 TASTSSTD--DACLLVKRPRR----ID-----RIEVVGARQKRGDVSFSERLVGVKEYTV 522
AS+SS ++ L P R +D R EV+GA + G + + V
Sbjct: 168 AASSSSISKSESGSLAANPLRGSSAVDSFYKLRCEVLGANIVKS---------GSRTFAV 320
Query: 523 YKIKVWS-GKDQWEVEKRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKESKIFRSASLD 581
Y I V + W +++R+R F LHR +K F E LP K F S+ LD
Sbjct: 321 YSISVTDVNNNSWSIKRRFRHFEELHRRLKE-FPEYNLHLP---------PKHFLSSGLD 470
Query: 582 I--IAKRSVLIQECLQSIL 598
+ I +R L+ + L+ ++
Sbjct: 471 VATIQERCELLDKYLKKLM 527
>TC89570 similar to GP|13936308|gb|AAK40307.1 putative methyl-binding domain
protein MBD105 {Zea mays}, partial (9%)
Length = 728
Score = 37.4 bits (85), Expect = 0.025
Identities = 34/109 (31%), Positives = 49/109 (44%), Gaps = 16/109 (14%)
Frame = +3
Query: 245 SSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRD---------RDRDRDRDRDAPVSC 295
SS K+E DE EETKD+ +EA E D+D + D+ DA V
Sbjct: 426 SSASKKEASE----DEAEETKDVEMQEAGETKDDKDVELEKKVVNENEDKKGAEDADVKE 593
Query: 296 EVQCAD-------NSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDE 337
+Q + N+ D DL ++ NI D G+EG G +K++E
Sbjct: 594 SIQPGETTDEGKSNTADGDLQASKE-----NIDDKGAEGSGVVQNKDEE 725
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 35.8 bits (81), Expect = 0.073
Identities = 23/74 (31%), Positives = 42/74 (56%), Gaps = 1/74 (1%)
Frame = +3
Query: 80 SNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGERE-IETE 138
+++K+ + G+ G + G+E D+ G D E ++ E+ D E GGE E E ++ E
Sbjct: 543 NSKKAPEGGAGGADENGEEEDDED--GDDQDE--DDDEDEDDDDEEEGGEEDEEEGVDEE 710
Query: 139 EEEEEEFSEGDDSM 152
+ EEEE E ++++
Sbjct: 711 DNEEEEEDEDEEAL 752
Score = 28.9 bits (63), Expect = 9.0
Identities = 26/91 (28%), Positives = 39/91 (42%), Gaps = 2/91 (2%)
Frame = +3
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNE--NGGEVGEREI 135
+D N + G +G+ DELS + GNN +S+ + GG G E
Sbjct: 420 EDGNDDEDEEDDDGDGAFGEGEDELSS---EDGGGYGNNSNNKSNSKKAPEGGAGGADEN 590
Query: 136 ETEEEEEEEFSEGDDSMFNYGSGCDGDNENE 166
EE++E +GDD + D D+E E
Sbjct: 591 GEEEDDE----DGDDQDEDDDEDEDDDDEEE 671
>TC82935 similar to GP|15150635|gb|AAK85442.1 Hypothetical protein C07H6.4
{Caenorhabditis elegans}, partial (2%)
Length = 807
Score = 35.4 bits (80), Expect = 0.096
Identities = 30/98 (30%), Positives = 43/98 (43%), Gaps = 1/98 (1%)
Frame = +3
Query: 269 DREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGK 328
D +A E V DRDR RDRDRDR V ++ D D + ++S+ DG +
Sbjct: 21 DDQARESVLDRDRSRDRDRDRGIKV-------EHLQDYD----DQARESMLDRDGSRDRD 167
Query: 329 GNRYSKNDEAGASGDAQREN-LDLDNFEFKSDQFCDNR 365
+R + + D RE+ LD D + D R
Sbjct: 168 RDRSRRVEHLQGYDDQARESMLDRDGSRDRDIMVRDRR 281
Score = 33.1 bits (74), Expect = 0.48
Identities = 45/202 (22%), Positives = 81/202 (39%), Gaps = 25/202 (12%)
Frame = +3
Query: 248 RKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDR-----------------DRD 290
R ++D I + +++ D L+ RDRDRDR R DRD
Sbjct: 63 RDRDRDRGIKVEHLQDYDDQARESMLDRDGSRDRDRDRSRRVEHLQGYDDQARESMLDRD 242
Query: 291 APVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRE-NL 349
++ D + L +D ++S+ D +G + GD ++ L
Sbjct: 243 GSRDRDIMVRDRRVE-HLQDYDDHRESMLDDDRNRDGDRAENELENSRVQGGDVDKDRTL 419
Query: 350 DLDNFEFKSDQ-------FCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKI 402
D+D + K+D+ F +++ + N+ ++V N ++ S N N
Sbjct: 420 DVDP-DKKNDRISDPDKSFDEHKKEQPRRNIDLNVTNQNSPSDSN-----GDHNDEVEDE 581
Query: 403 LERSSTSTNVLEKSHVISKIED 424
LERS + L+K +SK+E+
Sbjct: 582 LERSIQLLDQLKKE--VSKLEE 641
>TC78046 similar to GP|21952848|dbj|BAC06263. putative calreticulin {Oryza
sativa (japonica cultivar-group)}, partial (72%)
Length = 1268
Score = 35.0 bits (79), Expect = 0.13
Identities = 26/85 (30%), Positives = 39/85 (45%), Gaps = 3/85 (3%)
Frame = +3
Query: 208 GSHDLDDFLLQNGPV---SVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEET 264
G D+ L+ + P VV ++F N EE VRK +++ +E +
Sbjct: 834 GGSVFDNILICDDPEYAKQVVDEVFANREIEKEAFEE--AEKVRKAQEE-----EEAQRA 992
Query: 265 KDIGDREALEEVRDRDRDRDRDRDR 289
++ G+R E DR RDR RDR R
Sbjct: 993 REDGERRRKERGYDRHRDRHRDRYR 1067
>TC78045 similar to GP|15810563|gb|AAL07169.1 putative calreticulin protein
{Arabidopsis thaliana}, partial (91%)
Length = 1528
Score = 35.0 bits (79), Expect = 0.13
Identities = 26/85 (30%), Positives = 39/85 (45%), Gaps = 3/85 (3%)
Frame = +1
Query: 208 GSHDLDDFLLQNGPV---SVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEET 264
G D+ L+ + P VV ++F N EE VRK +++ +E +
Sbjct: 1009 GGSVFDNILICDDPEYAKQVVDEVFANREIEKEAFEE--AEKVRKAQEE-----EEAQRA 1167
Query: 265 KDIGDREALEEVRDRDRDRDRDRDR 289
++ G+R E DR RDR RDR R
Sbjct: 1168 REDGERRRKERGYDRHRDRHRDRYR 1242
>BE325350 weakly similar to GP|13702821|gb| putative snRNP protein {Oryza
sativa}, partial (6%)
Length = 473
Score = 35.0 bits (79), Expect = 0.13
Identities = 18/34 (52%), Positives = 25/34 (72%), Gaps = 1/34 (2%)
Frame = +2
Query: 258 NDEVEETKDIG-DREALEEVRDRDRDRDRDRDRD 290
+D + +D G DR++ E RDRDRDR+R+RDRD
Sbjct: 233 SDRRDRDRDRGHDRDSYRE-RDRDRDRERERDRD 331
Score = 29.3 bits (64), Expect = 6.9
Identities = 18/62 (29%), Positives = 26/62 (41%), Gaps = 4/62 (6%)
Frame = +2
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRD----RDRD 288
RE H + K +++ L++ D + + RDR DRD RDRD
Sbjct: 122 REQQALERRHNQVAGHKRDQEELLLLSTNNNINRSSDSDRRDRDRDRGHDRDSYRERDRD 301
Query: 289 RD 290
RD
Sbjct: 302 RD 307
>AJ388952 similar to PIR|T04526|T045 hypothetical protein F16A16.160 -
Arabidopsis thaliana, partial (24%)
Length = 591
Score = 34.7 bits (78), Expect = 0.16
Identities = 26/72 (36%), Positives = 32/72 (44%)
Frame = -3
Query: 129 EVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLY 188
E E E E EEEEEEE EGD+ + G G +E M GE V + +
Sbjct: 283 EPKEEEEEEEEEEEEEEGEGDEEE-DMARGKVGARRDEALVMMGEEVEMGV*KVNILEIE 107
Query: 189 EEGEVRNENPLL 200
+E E E LL
Sbjct: 106 DEEEEEEERFLL 71
>BQ149938 similar to GP|21166135|gb| hypothetical protein {Dictyostelium
discoideum}, partial (5%)
Length = 812
Score = 34.3 bits (77), Expect = 0.21
Identities = 15/29 (51%), Positives = 19/29 (64%)
Frame = +1
Query: 274 EEVRDRDRDRDRDRDRDAPVSCEVQCADN 302
E R+R+RDRDR+RDRD E + DN
Sbjct: 55 ERERERERDRDRERDRDTERDIEREIEDN 141
Score = 30.0 bits (66), Expect = 4.0
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +1
Query: 274 EEVRDRDRDRDRDRDRDAPVSCEVQ 298
E R+R+R+RDRDR+RD +++
Sbjct: 49 ERERERERERDRDRERDRDTERDIE 123
>BG449867 similar to GP|4127878|emb NDX1 homeobox protein {Glycine max},
partial (29%)
Length = 670
Score = 34.3 bits (77), Expect = 0.21
Identities = 30/108 (27%), Positives = 50/108 (45%), Gaps = 4/108 (3%)
Frame = +2
Query: 326 EGKG-NRYSKNDEAGASGD-AQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVD-A 382
+G G +R S+ + G SG A D+D K + C + NV VHV+N + +
Sbjct: 8 QGNGPSRQSQVENKGISGKTASGGARDIDKDAHKIETSCSDASSAKGKNVVVHVDNGELS 187
Query: 383 KSFKNLKPIVLPSNGGTRKI-LERSSTSTNVLEKSHVISKIEDSELSE 429
KS + LK + + N KI L + + + ++ IE++ L E
Sbjct: 188 KSNERLKRVGVEENPEDEKIELAQRKKRKRTIMNAEQVTMIENALLDE 331
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 33.9 bits (76), Expect = 0.28
Identities = 18/49 (36%), Positives = 27/49 (54%)
Frame = +3
Query: 118 IEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENE 166
+EE + E+ GEV E E E +EEEE+E E ++ + D+E E
Sbjct: 42 VEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEE 188
Score = 31.2 bits (69), Expect = 1.8
Identities = 20/54 (37%), Positives = 27/54 (49%)
Frame = +3
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDD 150
DE DE D E EE + +E+ E E E E EEEEE++ EG++
Sbjct: 93 DEHDEEEEDEHDEEE---EEEEEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGEE 245
>TC87951 similar to PIR|C86418|C86418 hypothetical protein AAG12629.1
[imported] - Arabidopsis thaliana, partial (13%)
Length = 922
Score = 33.9 bits (76), Expect = 0.28
Identities = 46/184 (25%), Positives = 69/184 (37%), Gaps = 6/184 (3%)
Frame = +3
Query: 75 RTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGERE 134
R D F GS +E +E+ + S+ + EE G GG+ G E
Sbjct: 210 RNHGDDQNNQFNEGS-------EEVEEIEVK--PSAAAVVAKSSEEKVGEAEGGKEGVVE 362
Query: 135 IETEEEEEEEFSEGDDSM-FNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEV 193
IE + + EE + D NYGS + N+ + E G
Sbjct: 363 IEWDMKSEESLEKIDSPKDLNYGSRKNNGGSND--------------------VIETGTA 482
Query: 194 RNENPLLMNSSVAFGSHDLDDFL-----LQNGPVSVVSDLFYNPRESNNRVEEHGVSSVR 248
+N N+SVA S DL + +Q+G V+D+ P S R E ++ R
Sbjct: 483 KNSKDESYNNSVAEVSDDLVKAVTSVTEMQSG--GTVNDILEKPLGS--RAAETDLAVKR 650
Query: 249 KEEK 252
E+K
Sbjct: 651 NEDK 662
>TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A3.4~unknown
protein {Arabidopsis thaliana}, partial (80%)
Length = 1253
Score = 33.5 bits (75), Expect = 0.36
Identities = 20/52 (38%), Positives = 25/52 (47%)
Frame = +1
Query: 101 ELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSM 152
E G+ SE IGN G++E + E EEEEEEE E DD +
Sbjct: 529 EQERIGVPRSEWIGNEGLQEEE-------------EEEEEEEEEEEEDDDDV 645
>TC80224 similar to GP|12324597|gb|AAG52258.1 putative coatomer protein
complex subunit beta 2 (beta prime); 18270-12231
{Arabidopsis thaliana}, partial (13%)
Length = 971
Score = 33.5 bits (75), Expect = 0.36
Identities = 27/93 (29%), Positives = 42/93 (45%), Gaps = 2/93 (2%)
Frame = +1
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEG--DDSMFN 154
+EGD+ G + EL +NG E D E E E TEE+EE+ EG DD++
Sbjct: 301 EEGDQPLENGDSNHELTEHNG--EEDYTEGHEEQNGEEDYTEEQEEQNGEEGSQDDAVV- 471
Query: 155 YGSGCDGDNENEFYSMKGENVSSEFYSSRSVSL 187
D D+ + + G E +++ S+
Sbjct: 472 ----VDADSSDGTVLINGNEADEELSTNKEGSV 558
>TC78611 similar to GP|15450599|gb|AAK96571.1 AT5g17440/K3M16_10
{Arabidopsis thaliana}, partial (91%)
Length = 1636
Score = 33.5 bits (75), Expect = 0.36
Identities = 26/128 (20%), Positives = 53/128 (41%), Gaps = 9/128 (7%)
Frame = +1
Query: 171 KGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSV--VSDL 228
K E++ + + EE E + P + + D + + + V +
Sbjct: 709 KAEDLGEQGLIDEAQKALEEAEALKKLPSRQEPPLDSSKYTAADVRITDQKLRVCDICGA 888
Query: 229 FYNPRESNNRVEEH-------GVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDR 281
F + +S+ R+ +H G +R++ + ++ + + DR + E RDRDR
Sbjct: 889 FLSVYDSDRRLADHFGGKLHLGYMQIREKLAELQEERNKSRKIDRLDDRRSKERSRDRDR 1068
Query: 282 DRDRDRDR 289
+ RDR+R
Sbjct: 1069EPSRDRER 1092
>TC80578 similar to GP|9759287|dbj|BAB09752.1 arginine-aspartate-rich RNA
binding protein-like {Arabidopsis thaliana}, partial
(74%)
Length = 987
Score = 33.5 bits (75), Expect = 0.36
Identities = 17/55 (30%), Positives = 31/55 (55%), Gaps = 4/55 (7%)
Frame = +2
Query: 239 VEEHGVSSVRKEEKDSLIVNDEVEETK----DIGDREALEEVRDRDRDRDRDRDR 289
+ EH ++ + +E++ L EVEE + +R+ + DR++ RD+DRDR
Sbjct: 812 INEHKIAKEKAKEEERLTREKEVEERRKQKEQDPERKRRSDPSDREKYRDKDRDR 976
>BI307955 weakly similar to GP|17064958|gb Unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 768
Score = 32.7 bits (73), Expect = 0.62
Identities = 17/47 (36%), Positives = 30/47 (63%)
Frame = +1
Query: 244 VSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRD 290
+S RK+++ S + E+ E + DR +RD+DRDR+RD++R+
Sbjct: 514 LSHDRKDDRASYLREFELREEERRRDR-----LRDKDRDRERDKERE 639
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.133 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,684,509
Number of Sequences: 36976
Number of extensions: 428112
Number of successful extensions: 2514
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 2106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2314
length of query: 880
length of database: 9,014,727
effective HSP length: 105
effective length of query: 775
effective length of database: 5,132,247
effective search space: 3977491425
effective search space used: 3977491425
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC124952.2


