

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.12 + phase: 2 /pseudo
(313 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
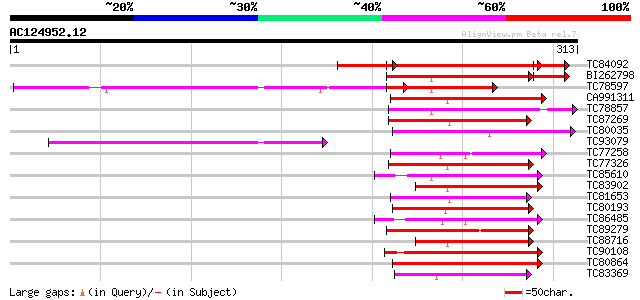
Score E
Sequences producing significant alignments: (bits) Value
TC84092 weakly similar to GP|17644123|gb|AAL38986.1 cytochrome P... 127 1e-46
BI262798 weakly similar to PIR|T05735|T05 cytochrome P450 71A10 ... 86 8e-19
TC78597 similar to GP|166949|gb|AAA32913.1|| cytochrome P-450LXX... 76 2e-14
CA991311 weakly similar to SP|O81973|C933 Cytochrome P450 93A3 (... 71 5e-13
TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 7... 65 4e-11
TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soy... 64 6e-11
TC80035 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P... 61 6e-10
TC93079 weakly similar to PIR|T05735|T05735 cytochrome P450 71A1... 60 8e-10
TC77258 similar to GP|5059126|gb|AAD38930.1| cytochrome P450 mon... 60 8e-10
TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pi... 60 1e-09
TC85610 similar to GP|21068674|emb|CAD31843. putative cytochrome... 59 2e-09
TC83902 similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC... 59 3e-09
TC81653 similar to SP|P93149|C9B1_GLYEC Cytochrome P450 93B1 (EC... 57 1e-08
TC80193 weakly similar to SP|O22307|C7DB_LOTJA Cytochrome P450 7... 56 2e-08
TC86485 similar to GP|23315211|gb|AAN20392.1 Sequence 405 from p... 56 2e-08
TC89279 weakly similar to SP|O81972|C822_SOYBN Cytochrome P450 8... 54 8e-08
TC88716 weakly similar to GP|20258812|gb|AAM14017.1 putative cyt... 53 1e-07
TC90108 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (E... 51 5e-07
TC80864 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (E... 50 9e-07
TC83369 similar to SP|P48419|C753_PETHY Flavonoid 3' 5'-hydroxyl... 50 1e-06
>TC84092 weakly similar to GP|17644123|gb|AAL38986.1 cytochrome P450-3 {Musa
acuminata}, partial (27%)
Length = 679
Score = 127 bits (318), Expect(3) = 1e-46
Identities = 62/86 (72%), Positives = 72/86 (83%)
Frame = +3
Query: 209 KLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKAL 268
++K K + EEM+DF D VIVEHKM+ RDP KKDFLDILLQLQDDG +E +LTQNDLKAL
Sbjct: 81 QIKNFKATFEEMNDFFDSVIVEHKMATRDPNKKDFLDILLQLQDDGRSELDLTQNDLKAL 260
Query: 269 LMNMFLGGSDTTSTTLEWAMAACKKS 294
LM+MFL GSDTTSTT+EWAMA K+
Sbjct: 261 LMDMFLAGSDTTSTTVEWAMAELVKN 338
Score = 50.4 bits (119), Expect(3) = 1e-46
Identities = 27/33 (81%), Positives = 29/33 (87%)
Frame = +1
Query: 182 GG*RLKF*ILVWEICFLC*VGLMFSLAKLKKIK 214
GG* LKF*ILV EICFLC VGLMFSLAKLK ++
Sbjct: 1 GG*WLKF*ILV*EICFLCWVGLMFSLAKLKILR 99
Score = 47.8 bits (112), Expect(3) = 1e-46
Identities = 20/20 (100%), Positives = 20/20 (100%)
Frame = +2
Query: 290 ACKKSSHNEESTGGGKENCR 309
ACKKSSHNEESTGGGKENCR
Sbjct: 326 ACKKSSHNEESTGGGKENCR 385
>BI262798 weakly similar to PIR|T05735|T05 cytochrome P450 71A10 - soybean,
partial (20%)
Length = 548
Score = 86.3 bits (212), Expect(2) = 8e-19
Identities = 43/86 (50%), Positives = 59/86 (68%), Gaps = 5/86 (5%)
Frame = +1
Query: 209 KLKKIKDSSEEMDDFLDRVIVEH-----KMSRRDPKKKDFLDILLQLQDDGLTEFELTQN 263
K+++ K++ E+D D+VI E KM KKK F+DILLQLQ+ G+ FEL+ N
Sbjct: 13 KIREFKETFRELDSLFDQVIEERLALKKKMENDQFKKKGFVDILLQLQEGGMLGFELSNN 192
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMA 289
D+KALL +MF+GG+DT S TLEW M+
Sbjct: 193 DIKALLTDMFVGGTDTASATLEWGMS 270
Score = 24.6 bits (52), Expect(2) = 8e-19
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +3
Query: 290 ACKKSSHNEESTGGGKENCR 309
A + S+HNEES K++CR
Sbjct: 273 AYETSNHNEESARRSKKSCR 332
>TC78597 similar to GP|166949|gb|AAA32913.1|| cytochrome P-450LXXIA1
(cyp71A1) {Persea americana}, partial (18%)
Length = 1170
Score = 75.9 bits (185), Expect = 2e-14
Identities = 79/228 (34%), Positives = 101/228 (43%), Gaps = 10/228 (4%)
Frame = +3
Query: 3 PFHVYFHPHLDCQ*LATTFN*EHFLIALFNPSHKSMVL**CCILVSFLCW--------WF 54
P + FH H + + +* HF L H +M * CC W WF
Sbjct: 177 PIQINFHLHRNYRSSVIFIS*AHFHTVLLETFHSNMGT*CCC------SWDKNKIQL*WF 338
Query: 55 PQYIWPRKSCKHMA*FLRTDQARH*QKPYSMVVRTLRFHLMAIHGDRKRNCV*MNC*VRK 114
Q + P KS K F +TD QK Y M V TL M G++K N V N * K
Sbjct: 339 HQQM*PWKS*KITIEFFQTDLNI*LQKSYCMDVMTLVLDFMEKIGNKKENYVFPNF*T*K 518
Query: 115 GCNQYSLLEKSNLLHWLIRYVKL*V*VVVVAV*TLVRC*LKQQTR*SVGVYLEGNM--TM 172
CN LEK L +WLI Y KL ++ V* VR + Q V V LE +M M
Sbjct: 519 VCNHIISLEKKKLTNWLISYAKL---A*MMNV*I*VR*LFQHQIILFVNVLLEESMKEIM 689
Query: 173 KVTDWESLAGG*RLKF*ILVWEICFLC*VGLMFSLAKLKKIKDSSEEM 220
K T ++ G F +L EI FL VGLMF + KL+ ++ E +
Sbjct: 690 KAT-*KN*QGRL*FIFKLLQLEITFLLWVGLMFLVEKLENLRKLFESL 830
Score = 58.9 bits (141), Expect = 2e-09
Identities = 31/66 (46%), Positives = 43/66 (64%), Gaps = 5/66 (7%)
Frame = +2
Query: 209 KLKKIKDSSEEMDDFLDRVIVEH-----KMSRRDPKKKDFLDILLQLQDDGLTEFELTQN 263
K+++ K++ E+D D+VI E KM KKK F+DILLQLQ+ G+ FEL+ N
Sbjct: 794 KIREFKETFRELDSLFDQVIEERLALKKKMENDQFKKKGFVDILLQLQEGGMLGFELSNN 973
Query: 264 DLKALL 269
D+KALL
Sbjct: 974 DIKALL 991
>CA991311 weakly similar to SP|O81973|C933 Cytochrome P450 93A3 (EC 1.14.-.-)
(P450 CP5). [Soybean] {Glycine max}, partial (33%)
Length = 768
Score = 71.2 bits (173), Expect = 5e-13
Identities = 35/93 (37%), Positives = 58/93 (61%), Gaps = 7/93 (7%)
Frame = +3
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-------KDFLDILLQLQDDGLTEFELTQN 263
K++K+ D ++ VI EH+ + K+ KD LDILL++ +D +E +LT+
Sbjct: 441 KRLKELMNMFDTMIESVIREHQEEMKKRKENGEGAHVKDLLDILLEIHEDESSEIKLTRK 620
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMAACKKSSH 296
++KA + +MFL G+DT+STT+EWA+A + H
Sbjct: 621 NVKAFIFDMFLSGTDTSSTTIEWALAELINNPH 719
>TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC
1.14.-.-) (P450 CP7). [Soybean] {Glycine max}, partial
(38%)
Length = 1724
Score = 64.7 bits (156), Expect = 4e-11
Identities = 31/107 (28%), Positives = 62/107 (56%), Gaps = 3/107 (2%)
Frame = +3
Query: 210 LKKIKDSSEEMDDFLDRVIVEH---KMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLK 266
+ ++ ++ + +D+F ++V+ EH ++D ++KD +D+LL+L++ G +LT N +K
Sbjct: 792 IARVDNTFKALDEFFEQVLKEHLNPNNRKKDDEEKDIVDVLLELKNQGRLSIDLTNNHIK 971
Query: 267 ALLMNMFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
A++MN+ + +DT++ T W M K N + +E R I K
Sbjct: 972 AVVMNLLVAATDTSAATSVWVMTGLIK---NPRAMKKAQEEIRNIKK 1103
>TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soybean
(fragment), partial (79%)
Length = 1878
Score = 64.3 bits (155), Expect = 6e-11
Identities = 34/84 (40%), Positives = 52/84 (61%), Gaps = 5/84 (5%)
Frame = +1
Query: 210 LKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKK-----DFLDILLQLQDDGLTEFELTQND 264
L K+ + +E+D RVI +H + PK K D +DILLQ+ +D F+LT +
Sbjct: 955 LGKLDKTFKELDLIYQRVIDDHMDNSARPKTKEQEVDDIIDILLQMMNDHSLSFDLTLDH 1134
Query: 265 LKALLMNMFLGGSDTTSTTLEWAM 288
+KA+LMN+F+ G+DT+S + WAM
Sbjct: 1135 IKAVLMNIFIAGTDTSSAIVVWAM 1206
>TC80035 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P450 {Pyrus
communis}, partial (49%)
Length = 1342
Score = 60.8 bits (146), Expect = 6e-10
Identities = 32/108 (29%), Positives = 60/108 (54%), Gaps = 7/108 (6%)
Frame = +1
Query: 212 KIKDSSEEMDDFLDRVIVEHKMS-RRDPKKKDFLDILLQLQDDGLTEFELTQN------D 264
++K +S+ D ++++I +H+ ++DP KDF+DILL L + + Q +
Sbjct: 706 RMKKTSKAFDQIVEKIIKDHEQDMKKDPHHKDFIDILLSLMHKSMDPHDEEQKQVIDITN 885
Query: 265 LKALLMNMFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGIN 312
+KA++++M G D+ STT+EWAM+ + + + EN G+N
Sbjct: 886 IKAIILDMIGGALDSASTTIEWAMSELLRHPNVMKKLQNELENVVGLN 1029
>TC93079 weakly similar to PIR|T05735|T05735 cytochrome P450 71A10 -
soybean, partial (24%)
Length = 671
Score = 60.5 bits (145), Expect = 8e-10
Identities = 60/154 (38%), Positives = 70/154 (44%)
Frame = +3
Query: 22 N*EHFLIALFNPSHKSMVL**CCILVSFLCWWFPQYIWPRKSCKHMA*FLRTDQARH*QK 81
+* HF LF H +M * C F Q I P KS M * RT+ QK
Sbjct: 189 S*AHFHTVLFETYHTNMET*CYCN*DKDKL**FHQQI*PWKS*ITMI*LFRTEPKL*LQK 368
Query: 82 PYSMVVRTLRFHLMAIHGDRKRNCV*MNC*VRKGCNQYSLLEKSNLLHWLIRYVKL*V*V 141
Y M R L H+M GD+K V N * K N + LEK L +W + YVKL
Sbjct: 369 SYFMEERMLVLHIMEKIGDKKEKYVS*NF*A*KVYNHFIKLEKK*LRNW*VNYVKL---A 539
Query: 142 VVVAV*TLVRC*LKQQTR*SVGVYLEGNMTMKVT 175
++ V* V C LK QT V LEGNM VT
Sbjct: 540 QMMHV*I*VSCLLKLQTILFVHALLEGNMEEIVT 641
>TC77258 similar to GP|5059126|gb|AAD38930.1| cytochrome P450
monooxygenaseCYP93D1 {Glycine max}, partial (95%)
Length = 1823
Score = 60.5 bits (145), Expect = 8e-10
Identities = 34/93 (36%), Positives = 56/93 (59%), Gaps = 7/93 (7%)
Frame = +1
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRR-----DPKKKDFLDILLQL--QDDGLTEFELTQN 263
K+ KD+ +MD +++V+ EH+ +R +KKD DILL L DDG E +LT+
Sbjct: 787 KRNKDTHHKMDVMMEKVLKEHEEARAKEGAGSDRKKDLFDILLNLIEADDG-AESKLTRQ 963
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMAACKKSSH 296
KA ++MF+ G++ ++ LEWA+A ++ H
Sbjct: 964 SAKAFALDMFIAGTNGPASVLEWALAELIRNPH 1062
>TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pisum
sativum}, complete
Length = 1864
Score = 59.7 bits (143), Expect = 1e-09
Identities = 28/83 (33%), Positives = 55/83 (65%), Gaps = 3/83 (3%)
Frame = +1
Query: 210 LKKIKDSSEEMDDFLDRVIVEHKMSRRDPKK---KDFLDILLQLQDDGLTEFELTQNDLK 266
+K++K S++ D F++ V+ EH R++ K KD +D+LLQL +D E +L ++ +K
Sbjct: 769 VKRMKTLSKKFDRFMEHVLEEHIERRKNVKDYVAKDMVDVLLQLAEDPNLEVKLERHGVK 948
Query: 267 ALLMNMFLGGSDTTSTTLEWAMA 289
A ++ GG+++++ T+EWA++
Sbjct: 949 AFTQDLIAGGTESSAVTVEWAIS 1017
>TC85610 similar to GP|21068674|emb|CAD31843. putative cytochrome P450
monooxygenase {Cicer arietinum}, complete
Length = 1578
Score = 58.9 bits (141), Expect = 2e-09
Identities = 34/96 (35%), Positives = 51/96 (52%), Gaps = 3/96 (3%)
Frame = +2
Query: 202 GLMFSLAKLKKIKDSSEEMDDFLDRVIVEH---KMSRRDPKKKDFLDILLQLQDDGLTEF 258
GLM L KL KI +D F V EH + P ++D +D L++L++D
Sbjct: 701 GLMSRLEKLFKI------LDGFFQSVFDEHIDPNRKKLPPHEEDVIDALIELKNDPYCSM 862
Query: 259 ELTQNDLKALLMNMFLGGSDTTSTTLEWAMAACKKS 294
+L+ +K L+MNM L G+DT + + WAM A K+
Sbjct: 863 DLSAEHIKPLIMNMLLAGTDTIAAAVVWAMTALMKN 970
>TC83902 similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC 1.14.-.-)
(P450 CP7). [Soybean] {Glycine max}, partial (41%)
Length = 900
Score = 58.5 bits (140), Expect = 3e-09
Identities = 27/78 (34%), Positives = 50/78 (63%), Gaps = 8/78 (10%)
Frame = +1
Query: 225 DRVIVEHKMSRRDPKK--------KDFLDILLQLQDDGLTEFELTQNDLKALLMNMFLGG 276
+ +I +H+ R K+ +D LD+LL++Q + ++T N++KA++ ++F+ G
Sbjct: 49 ENIIRQHQEKRERAKEDDNNEVDNEDLLDVLLRVQQSDNLDIKITTNNIKAVIWDVFVAG 228
Query: 277 SDTTSTTLEWAMAACKKS 294
+DTTSTT+EWAM+ K+
Sbjct: 229 TDTTSTTIEWAMSEMMKN 282
>TC81653 similar to SP|P93149|C9B1_GLYEC Cytochrome P450 93B1 (EC 1.14.-.-)
((2S)-flavanone 2-hydroxylase) (Licodione synthase),
partial (62%)
Length = 1193
Score = 56.6 bits (135), Expect = 1e-08
Identities = 30/89 (33%), Positives = 51/89 (56%), Gaps = 11/89 (12%)
Frame = +1
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKK-----------DFLDILLQLQDDGLTEFE 259
K+I+D D ++R+I + + R++ +K DFLDILL +D +E +
Sbjct: 121 KRIEDLFMRFDTLVERIITKREELRKNKGRKENKGEQGAEFRDFLDILLDCAEDQNSEIK 300
Query: 260 LTQNDLKALLMNMFLGGSDTTSTTLEWAM 288
+ + +KAL+M+ F G+DTTS + EWA+
Sbjct: 301 VQRVHIKALIMDFFTAGTDTTSISTEWAL 387
>TC80193 weakly similar to SP|O22307|C7DB_LOTJA Cytochrome P450 71D11 (EC
1.14.-.-) (Fragment). {Lotus japonicus}, partial (32%)
Length = 941
Score = 56.2 bits (134), Expect = 2e-08
Identities = 27/84 (32%), Positives = 53/84 (62%), Gaps = 6/84 (7%)
Frame = +2
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPK------KKDFLDILLQLQDDGLTEFELTQNDL 265
K++ +D ++ +I EHK ++ + +D +D+LL+ QD EF LT +++
Sbjct: 686 KLERVHRVVDQIMENIINEHKEAKSKGEFDQVESDEDLVDVLLKYQDGNNKEFFLTIDNI 865
Query: 266 KALLMNMFLGGSDTTSTTLEWAMA 289
KA++M++F G +T+++T++WAMA
Sbjct: 866 KAIIMDIFGAGGETSASTIDWAMA 937
>TC86485 similar to GP|23315211|gb|AAN20392.1 Sequence 405 from patent US
6410718, partial (90%)
Length = 1927
Score = 55.8 bits (133), Expect = 2e-08
Identities = 33/105 (31%), Positives = 57/105 (53%), Gaps = 12/105 (11%)
Frame = +3
Query: 202 GLMFSLAKLKKIKDSSEEMDDFLDRVIVEHKMSRRD--PKKKDFLDILLQL--------- 250
GL L K +K E+D F+D++I EH ++ ++ D +D LL
Sbjct: 843 GLNARLVKARK------ELDSFIDKIIDEHVQKKKSVVDEETDMVDELLAFYSEEAKLNN 1004
Query: 251 -QDDGLTEFELTQNDLKALLMNMFLGGSDTTSTTLEWAMAACKKS 294
DD +LT++++KA++M++ GG++T ++ +EWAMA KS
Sbjct: 1005ESDDLNNSIKLTKDNIKAIIMDVMFGGTETVASAIEWAMAELMKS 1139
>TC89279 weakly similar to SP|O81972|C822_SOYBN Cytochrome P450 82A2 (EC
1.14.-.-) (P450 CP4). [Soybean] {Glycine max}, partial
(32%)
Length = 1088
Score = 53.9 bits (128), Expect = 8e-08
Identities = 26/81 (32%), Positives = 49/81 (60%)
Frame = +1
Query: 209 KLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKAL 268
K K++K +++E+DDF+ + +HK + +++DF+D+LL D+ L + +KA
Sbjct: 136 KQKQMKKTAKELDDFVCVWLDQHKHKKNSGREQDFMDVLLSTVDEDLDGRD-ADTTIKAT 312
Query: 269 LMNMFLGGSDTTSTTLEWAMA 289
M + L +DTT+ TL W ++
Sbjct: 313 CMALILAATDTTAGTLTWTLS 375
>TC88716 weakly similar to GP|20258812|gb|AAM14017.1 putative cytochrome
P450 homolog {Arabidopsis thaliana}, partial (57%)
Length = 1228
Score = 53.1 bits (126), Expect = 1e-07
Identities = 28/69 (40%), Positives = 44/69 (63%), Gaps = 4/69 (5%)
Frame = +2
Query: 225 DRVIVEHKMSRRDPKK----KDFLDILLQLQDDGLTEFELTQNDLKALLMNMFLGGSDTT 280
+++I E D K+ +DFL LL L+++G ++ T +KALLM+M +GGSDT+
Sbjct: 641 EKMIGERLKKEEDGKENNGSRDFLQFLLNLKEEGDSKTPFTNTHVKALLMDMVVGGSDTS 820
Query: 281 STTLEWAMA 289
S T+E+ MA
Sbjct: 821 SNTIEFVMA 847
>TC90108 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (41%)
Length = 958
Score = 51.2 bits (121), Expect = 5e-07
Identities = 26/89 (29%), Positives = 59/89 (66%), Gaps = 2/89 (2%)
Frame = +1
Query: 208 AKLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFL-DILLQLQDDGL-TEFELTQNDL 265
AK++K+ +E+D L +I +HK ++ +++L D+LL++Q + ++ LT +++
Sbjct: 103 AKMEKLH---KEIDMILQDIIDDHKSIHKEDSNEEYLVDVLLKIQQENYHSQHPLTDDNI 273
Query: 266 KALLMNMFLGGSDTTSTTLEWAMAACKKS 294
K+++ ++F G++T+STT+ WA++ K+
Sbjct: 274 KSIIQDIFDAGTETSSTTVLWAISEMVKN 360
>TC80864 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (46%)
Length = 1083
Score = 50.4 bits (119), Expect = 9e-07
Identities = 25/85 (29%), Positives = 52/85 (60%), Gaps = 2/85 (2%)
Frame = +3
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKK-KDFLDILLQLQDDGL-TEFELTQNDLKALL 269
K++ E+D +I +H+ ++ +D +D+LL++Q + +E LT +++K+++
Sbjct: 777 KMEKLHTEIDMIAQDIIDDHRSIHKEASNDEDLVDVLLKIQQENYHSEHPLTDDNMKSII 956
Query: 270 MNMFLGGSDTTSTTLEWAMAACKKS 294
+MFL G++T+S L WAM+ K+
Sbjct: 957 QDMFLAGTETSSQVLLWAMSEMVKN 1031
>TC83369 similar to SP|P48419|C753_PETHY Flavonoid 3' 5'-hydroxylase 2 (EC
1.14.-.-) (F3'5'H) (Cytochrome P450 75A3) (CYPLXXVA3).
[Petunia], partial (52%)
Length = 1084
Score = 50.1 bits (118), Expect = 1e-06
Identities = 27/78 (34%), Positives = 46/78 (58%), Gaps = 2/78 (2%)
Frame = +1
Query: 213 IKDSSEEMDDFLDRVIVEHKMS--RRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
+K ++ D L ++I EH S + K DFLD L+ + +L+ ++KALL+
Sbjct: 793 MKSLHKKFDALLTKMIEEHVASSLKSHRVKPDFLDRLIAQSKEDSDGEKLSLTNIKALLL 972
Query: 271 NMFLGGSDTTSTTLEWAM 288
N+F G+DT+S+ +EWA+
Sbjct: 973 NLFTAGTDTSSSIIEWAL 1026
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.353 0.156 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,770,696
Number of Sequences: 36976
Number of extensions: 155818
Number of successful extensions: 2566
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 2515
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2556
length of query: 313
length of database: 9,014,727
effective HSP length: 96
effective length of query: 217
effective length of database: 5,465,031
effective search space: 1185911727
effective search space used: 1185911727
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC124952.12


