

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
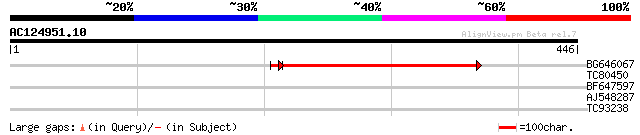
Query= AC124951.10 - phase: 0
(446 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG646067 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsi... 310 2e-85
TC80450 similar to GP|15982860|gb|AAL09777.1 AT4g11790/T5C23_220... 35 0.076
BF647597 weakly similar to GP|19071648|gb| putative protein tyro... 29 4.2
AJ548287 28 5.5
TC93238 SP|P03756|VE22_LAMBD EA22 gene protein. {Bacteriophage l... 28 7.1
>BG646067 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsis thaliana},
partial (32%)
Length = 516
Score = 310 bits (793), Expect(2) = 2e-85
Identities = 153/157 (97%), Positives = 154/157 (97%)
Frame = +2
Query: 215 KIGRVTAGVQYEPHHENAKLSNLMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASF 274
+IGRVTAGVQYEPHHENAKLSNLMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASF
Sbjct: 29 RIGRVTAGVQYEPHHENAKLSNLMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASF 208
Query: 275 YQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDDAIAANNISDSTFQIAASWQANK 334
YQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDDAIAANNISDSTFQIAASWQANK
Sbjct: 209 YQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDDAIAANNISDSTFQIAASWQANK 388
Query: 335 NFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDR 371
NFLVKAKAGPKSSTMALAFKSWWKPSFTFSIS T R
Sbjct: 389 NFLVKAKAGPKSSTMALAFKSWWKPSFTFSISGTSSR 499
Score = 24.6 bits (52), Expect(2) = 2e-85
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = +1
Query: 206 LPKSVWLVSK 215
LPKSVWLVSK
Sbjct: 1 LPKSVWLVSK 30
>TC80450 similar to GP|15982860|gb|AAL09777.1 AT4g11790/T5C23_220
{Arabidopsis thaliana}, partial (40%)
Length = 1192
Score = 34.7 bits (78), Expect = 0.076
Identities = 31/135 (22%), Positives = 56/135 (40%), Gaps = 18/135 (13%)
Frame = +3
Query: 280 VQRRVKNPLEENTVVGITNYIDFGFELQTSVDDAI------------------AANNISD 321
VQ ++KN +E G+ +Y+D + D + A + +
Sbjct: 204 VQTQLKNHPDELWEDGVRDYLDHATGIMEKFSDVVNWLKANATKAENSTPGAGAGVSFAG 383
Query: 322 STFQIAASWQANKNFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDRADGQVQYGFG 381
F + + NKNF K+ + P S+T +F S W P FS S +++ +G G
Sbjct: 384 KKFLPEVTNKENKNFGEKSGSPPVSTTTTTSFASPWSPGI-FSNSQNQNQNQNLFTFG-G 557
Query: 382 LQSESLREASYQRAD 396
QS + +++ +D
Sbjct: 558 NQSSAPSNSNHNASD 602
>BF647597 weakly similar to GP|19071648|gb| putative protein
tyrosine-serine-threonine kinase {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 520
Score = 28.9 bits (63), Expect = 4.2
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Frame = +3
Query: 266 KSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYIDFGFELQT----------SVDDAIA 315
KSS F+ FY H + RVK ++ VVG + G + + D+ +
Sbjct: 15 KSSLFLNYFYVHGLACFRVKVETSKHNVVGREEEVSSGLSYKAASSFVPLTSRTEDEVLR 194
Query: 316 ANNISDSTFQIAASWQANKNFLVKAKAGPKSSTMALAFKSW 356
+NN+ + F I A +NF + G FK W
Sbjct: 195 SNNL--TCFTINEVRAATRNFHPDSMIG--KGGFGCVFKGW 305
>AJ548287
Length = 328
Score = 28.5 bits (62), Expect = 5.5
Identities = 19/58 (32%), Positives = 26/58 (44%)
Frame = +2
Query: 15 VESPPPVVLVPPLFDFPPIAARNRMLESSYDVVFGKLALRCLFNDYFQQPKHFVTRIM 72
V+ PP+++ PPL FPP L S+ F L L L+ P F TR +
Sbjct: 131 VDPTPPIIITPPLSFFPP--THRSTLPPSFLAFFCFLFLSHLY------PSSFYTRFI 280
>TC93238 SP|P03756|VE22_LAMBD EA22 gene protein. {Bacteriophage lambda},
partial (93%)
Length = 887
Score = 28.1 bits (61), Expect = 7.1
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = -1
Query: 252 SPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYID 301
S S + +F+ + S AS HM + + NPL N V+ + N +D
Sbjct: 557 SAFSQALSFTKPISLSFDANASVRHHMQILTCILNPLTSNPVIAMRNDVD 408
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.135 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,320,545
Number of Sequences: 36976
Number of extensions: 195027
Number of successful extensions: 1011
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 980
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1010
length of query: 446
length of database: 9,014,727
effective HSP length: 99
effective length of query: 347
effective length of database: 5,354,103
effective search space: 1857873741
effective search space used: 1857873741
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124951.10


