

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124217.4 + phase: 0
(583 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
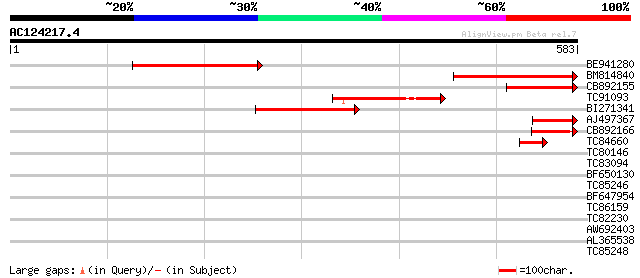
Score E
Sequences producing significant alignments: (bits) Value
BE941280 weakly similar to GP|20197614|g unknown protein {Arabid... 218 7e-57
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 156 2e-38
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 110 1e-24
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 102 5e-22
BI271341 87 2e-17
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 69 5e-12
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 65 9e-11
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 42 9e-04
TC80146 37 0.028
TC83094 similar to GP|20975622|emb|CAD31716. putative ripening r... 30 2.6
BF650130 similar to GP|20260594|gb| unknown protein {Arabidopsis... 29 4.4
TC85246 similar to GP|20975622|emb|CAD31716. putative ripening r... 29 5.7
BF647954 homologue to GP|11094192|db 26S proteasome regulatory p... 28 7.5
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 28 7.5
TC82230 similar to GP|9758915|dbj|BAB09452.1 contains similarity... 28 7.5
AW692403 28 9.8
AL365538 weakly similar to GP|12320927|g ABC transporter putati... 28 9.8
TC85248 similar to GP|20975622|emb|CAD31716. putative ripening r... 28 9.8
>BE941280 weakly similar to GP|20197614|g unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 403
Score = 218 bits (554), Expect = 7e-57
Identities = 105/134 (78%), Positives = 117/134 (86%)
Frame = -1
Query: 127 LYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGIAALLIPGGRTAHSRFGIPFTVDECS 186
LY YGGT KTFIW+A+SAA+RS GEIVL ASSGI ALL+PGGRTAHSRFGIPF +DE S
Sbjct: 403 LYDYGGTEKTFIWRALSAALRSEGEIVLACASSGIDALLMPGGRTAHSRFGIPFIIDETS 224
Query: 187 TCGVKPNTPLAQLVVQARLIIWDEAPMMHKFCFEALDRSCRDVMKEVDKRNKNIPFGGKV 246
CGV PN PLA LV++A+LIIWDEAPMMHK CFEALDRS RDV+K VD+RNK+IPFGGKV
Sbjct: 223 MCGVTPNIPLASLVIKAKLIIWDEAPMMHKHCFEALDRSLRDVLKTVDERNKDIPFGGKV 44
Query: 247 VVFGGDFRQILPVI 260
VV GGDFRQIL V+
Sbjct: 43 VVLGGDFRQILLVM 2
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 156 bits (394), Expect = 2e-38
Identities = 76/127 (59%), Positives = 100/127 (77%)
Frame = +3
Query: 457 NGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPSDVRIPFKFQRKQFPLAVSFAM 516
+GTRLII +G+ V+ +VI G++ G+ +IPR++L PS + F+R QFPL +SFAM
Sbjct: 27 HGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAM 206
Query: 517 TINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVY 576
TINKSQGQ+L +VG+YLP PVF+HGQLYVAVSRV SR GLKILITDE+G ++ T NVVY
Sbjct: 207 TINKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVVY 386
Query: 577 EEVFRNV 583
+EVF+ +
Sbjct: 387 QEVFQKI 407
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 110 bits (276), Expect = 1e-24
Identities = 54/72 (75%), Positives = 65/72 (90%)
Frame = -2
Query: 512 VSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLT 571
V FAMTINKSQGQSL+++GVYLP+ VFSHGQLYVA+SRVTSR GLKILI+++DG+D +T
Sbjct: 364 VYFAMTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSREGLKILISNDDGEDDCVT 185
Query: 572 SNVVYEEVFRNV 583
SNVVY EVF N+
Sbjct: 184 SNVVYREVFHNL 149
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 102 bits (253), Expect = 5e-22
Identities = 54/119 (45%), Positives = 81/119 (67%), Gaps = 3/119 (2%)
Frame = +3
Query: 333 PTDLLVEVD---GCPIASIIRSTYPNLLASMGDIEYFQNRAILTSKNTIVEKINEYMLDM 389
P D EVD G PI +I++STYPNL++ + ++ Q+RAILTS + +V++IN+Y+L +
Sbjct: 351 PNDSYAEVDTPPGDPIDAIVQSTYPNLVSQYNNEQFLQSRAILTSTDEVVDQINDYVLKL 530
Query: 390 VPGEEKVYLSYDSPDERNVCGDAMDDVHTPKFLNTIVASGLPNHKLRLKEGVPVMLLRN 448
+PGEE+V S + + +V A D + P+FL ++ S LPNHKL LK G P+MLLR+
Sbjct: 531 IPGEERVIYSANRSEVNDV--QAFDAI-PPEFLQSLKTSDLPNHKLTLKVGTPIMLLRD 698
>BI271341
Length = 468
Score = 86.7 bits (213), Expect = 2e-17
Identities = 48/107 (44%), Positives = 66/107 (60%)
Frame = +1
Query: 253 FRQILPVIPRGIRPEIVHATINSSHLWRHCEVLRLTKNMRLLNGASEADIEDRRKFSEWV 312
FR ILPVIPRG R +I+HATINSS + HC+V+RL KNM L ++ + +FS+ +
Sbjct: 1 FR*ILPVIPRGSRSDIIHATINSSCI*DHCQVVRLKKNMWLQQNGQSSNDPEFEQFSK*I 180
Query: 313 LSIGDGTIGEDNLVDKTITIPTDLLVEVDGCPIASIIRSTYPNLLAS 359
L +GDG I E N I IP +LL+ + +I++STY N S
Sbjct: 181 LKVGDGKIYEPNDSYADIDIPPELLISNYDDSLQTIVQSTYQNFWMS 321
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 543
Score = 68.9 bits (167), Expect = 5e-12
Identities = 34/46 (73%), Positives = 40/46 (86%)
Frame = +1
Query: 538 FSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVYEEVFRNV 583
FS+G+LYVAVSRVTSR GLKIL+ EDG+ N TSNVVY+EVFRN+
Sbjct: 19 FSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRNL 156
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 64.7 bits (156), Expect = 9e-11
Identities = 33/47 (70%), Positives = 41/47 (87%)
Frame = -1
Query: 537 VFSHGQLYVAVSRVTSRGGLKILITDEDGDDTNLTSNVVYEEVFRNV 583
VFSHGQLYVA+SRV+SR GLKIL+ DE+GD + T+NVVY +VF+NV
Sbjct: 274 VFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVY-KVFQNV 137
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 41.6 bits (96), Expect = 9e-04
Identities = 19/29 (65%), Positives = 23/29 (78%)
Frame = +2
Query: 525 SLQNVGVYLPAPVFSHGQLYVAVSRVTSR 553
S+ G+YLP P+F HG LYVA+SRVTSR
Sbjct: 68 SMATKGMYLPQPIF*HG*LYVALSRVTSR 154
>TC80146
Length = 476
Score = 36.6 bits (83), Expect = 0.028
Identities = 15/35 (42%), Positives = 24/35 (67%)
Frame = -3
Query: 510 LAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLY 544
+ + + TINKS+ QSL + +YL P+FSH ++Y
Sbjct: 471 IQIPYFKTINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>TC83094 similar to GP|20975622|emb|CAD31716. putative ripening related
protein {Cicer arietinum}, partial (58%)
Length = 707
Score = 30.0 bits (66), Expect = 2.6
Identities = 17/62 (27%), Positives = 31/62 (49%), Gaps = 3/62 (4%)
Frame = +2
Query: 215 HKFCFEALDRSCRDVMKEVDKRNKNIP---FGGKVVVFGGDFRQILPVIPRGIRPEIVHA 271
H+ + +C + ++E+D++NK FGG + DF+ I+ VI +G V
Sbjct: 53 HEITTDGKVHTCHEKVEEIDEKNKKGTYKLFGGDIDEHYKDFKLIIEVIDKGDGTSAVKW 232
Query: 272 TI 273
T+
Sbjct: 233 TV 238
>BF650130 similar to GP|20260594|gb| unknown protein {Arabidopsis thaliana},
partial (14%)
Length = 635
Score = 29.3 bits (64), Expect = 4.4
Identities = 29/103 (28%), Positives = 52/103 (50%), Gaps = 9/103 (8%)
Frame = +1
Query: 27 VFDADLTLTDE-QLQIYALADVEKLLRSFGKSLSD-YPPMPTADPSL-MPDLQNHFIHEE 83
+FD T DE L + A+ ++++ + S SLSD +P P++ + M L++ F +
Sbjct: 181 IFDQTTTNRDEIDLYLQAVDEIQRSISS--TSLSDNHPSKPSSTIQIAMARLEDEFRNIL 354
Query: 84 INYNRRELQ---EEHNQLMS---KMTDEQHKVYDTIMARVDEK 120
I++ ++ E+ L S K+ DE+ YD VD+K
Sbjct: 355 ISHTNNQIDPSLEDDTYLSSSSSKLQDEEDNSYDDDGVDVDDK 483
>TC85246 similar to GP|20975622|emb|CAD31716. putative ripening related
protein {Cicer arietinum}, complete
Length = 724
Score = 28.9 bits (63), Expect = 5.7
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 3/52 (5%)
Frame = +1
Query: 225 SCRDVMKEVDKRNKNIP---FGGKVVVFGGDFRQILPVIPRGIRPEIVHATI 273
+C + ++E+D++NK FGG + DF+ I+ VI +G V T+
Sbjct: 268 TCHEKVEEIDEKNKKGTYKLFGGDIDEHYKDFKLIIEVIDKGDGTSAVKWTV 423
>BF647954 homologue to GP|11094192|db 26S proteasome regulatory particle
triple-A ATPase subunit4 {Oryza sativa (japonica
cultivar-group)}, partial (53%)
Length = 649
Score = 28.5 bits (62), Expect = 7.5
Identities = 19/49 (38%), Positives = 26/49 (52%)
Frame = +2
Query: 113 IMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGI 161
+ RV K P LYG GTGKT + +A+++ I + L V SS I
Sbjct: 14 LFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNIDAN---FLKVVSSAI 151
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 28.5 bits (62), Expect = 7.5
Identities = 19/49 (38%), Positives = 26/49 (52%)
Frame = +2
Query: 113 IMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGI 161
+ RV K P LYG GTGKT + +A+++ I + L V SS I
Sbjct: 560 LFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNIDAN---FLKVVSSAI 697
>TC82230 similar to GP|9758915|dbj|BAB09452.1 contains similarity to ABC
transporter~gene_id:MXI22.4 {Arabidopsis thaliana},
partial (69%)
Length = 1166
Score = 28.5 bits (62), Expect = 7.5
Identities = 25/86 (29%), Positives = 39/86 (45%)
Frame = -2
Query: 27 VFDADLTLTDEQLQIYALADVEKLLRSFGKSLSDYPPMPTADPSLMPDLQNHFIHEEINY 86
VF L+L +Q+ A ++ L S+G S P + A+ P Q+HF H +
Sbjct: 817 VFPFPLSLNIDQI*YLFFACTKQKLASYGYLSSSDPELQDAE----P*QQHHFCHLSVLL 650
Query: 87 NRRELQEEHNQLMSKMTDEQHKVYDT 112
E Q + NQ + QH++Y T
Sbjct: 649 YEPE-QLKQNQ------EAQHQIYQT 593
>AW692403
Length = 425
Score = 28.1 bits (61), Expect = 9.8
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +3
Query: 3 ICFICVSFLHQCYTLPLTYVNKCYVFDADLTL 34
I FIC F+ + YTL +V+ C +F +L L
Sbjct: 162 IFFICFVFIKKIYTLLSFFVHVCKIFSYNLGL 257
>AL365538 weakly similar to GP|12320927|g ABC transporter putative
{Arabidopsis thaliana}, partial (10%)
Length = 441
Score = 28.1 bits (61), Expect = 9.8
Identities = 25/86 (29%), Positives = 39/86 (45%), Gaps = 8/86 (9%)
Frame = +3
Query: 145 AIRSRGEIVLTVASSG--IAALLIPGGRTAH----SRFGIPFTVDECSTC--GVKPNTPL 196
A SR +IVL + ++G +A + P TA RF V C ++PN
Sbjct: 42 AALSR*KIVLAMETTGCTVAEFVAPRKSTAEYMAPKRFSKSCCVLVFLACFQNIRPNNNP 221
Query: 197 AQLVVQARLIIWDEAPMMHKFCFEAL 222
+V++ R ++ E KFCF+ L
Sbjct: 222 MIMVIKNRTALY*ECLQYDKFCFQKL 299
>TC85248 similar to GP|20975622|emb|CAD31716. putative ripening related
protein {Cicer arietinum}, complete
Length = 919
Score = 28.1 bits (61), Expect = 9.8
Identities = 15/52 (28%), Positives = 28/52 (53%), Gaps = 3/52 (5%)
Frame = +1
Query: 225 SCRDVMKEVDKRNKNIP---FGGKVVVFGGDFRQILPVIPRGIRPEIVHATI 273
+C + ++EVD++NK + FGG + DF+ I+ +I + + TI
Sbjct: 232 TCHESVEEVDEKNKKLSYKLFGGDIGENYKDFKLIIEIIDKSDSSAAIKWTI 387
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,554,775
Number of Sequences: 36976
Number of extensions: 237220
Number of successful extensions: 1199
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1191
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1197
length of query: 583
length of database: 9,014,727
effective HSP length: 101
effective length of query: 482
effective length of database: 5,280,151
effective search space: 2545032782
effective search space used: 2545032782
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124217.4


