

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
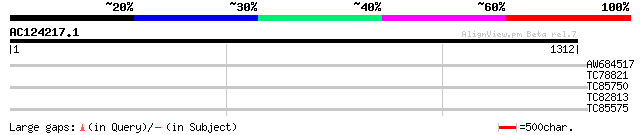
Query= AC124217.1 - phase: 0 /pseudo
(1312 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 38 0.029
TC78821 similar to GP|15293173|gb|AAK93697.1 unknown protein {Ar... 31 3.6
TC85750 nodulin 25 31 3.6
TC82813 30 4.7
TC85575 similar to PIR|D86410|D86410 protein F3M18.16 [imported]... 30 4.7
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 488
Score = 37.7 bits (86), Expect = 0.029
Identities = 17/33 (51%), Positives = 22/33 (66%)
Frame = +1
Query: 780 FLVVASKANYNLLLGTEWIHDVGDVPSTMH*RL 812
F V+ A+Y+ LLG WIHD G V ST+H +L
Sbjct: 7 FQVMDINASYSCLLGRPWIHDAGAVTSTLHQKL 105
>TC78821 similar to GP|15293173|gb|AAK93697.1 unknown protein {Arabidopsis
thaliana}, partial (81%)
Length = 1572
Score = 30.8 bits (68), Expect = 3.6
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Frame = +2
Query: 630 FTDFEADFDVICVVLILPIE*DVTSEVTEDEEDFIKEMVVHK-TMCYYVMNNGCVEGQHA 688
F + +D++C ++ P +V E+ K VV K ++C+Y +GC++G HA
Sbjct: 1394 FPELTLSYDMLCTIIENPC--NVRGEIRRH-----KIYVVAKFSLCFYFFLSGCIKGMHA 1552
>TC85750 nodulin 25
Length = 1132
Score = 30.8 bits (68), Expect = 3.6
Identities = 26/95 (27%), Positives = 42/95 (43%), Gaps = 4/95 (4%)
Frame = +3
Query: 481 NKRGVPYKDTPPTRHQNRGQVRTFTPSSKS---PTEKWV-FSSGKKSSYTSPPAKWVKRF 536
N++G+ YKDT RH N G + T + K P++ + SS + P + ++
Sbjct: 384 NQKGLEYKDT---RH-NDGSIEISTRTEKDNFIPSDGPIEISSRTEKKNVIPSDESIEIS 551
Query: 537 TGTDNQNKAPVSKKFAYNNNYKGKNPMTKT*WRRF 571
TGT+ +N P + KN + WR F
Sbjct: 552 TGTEKENFIPSVGPIDLTTGAQKKNFIPSVDWRTF 656
>TC82813
Length = 788
Score = 30.4 bits (67), Expect = 4.7
Identities = 28/100 (28%), Positives = 41/100 (41%), Gaps = 3/100 (3%)
Frame = -3
Query: 505 TPSSKSPTEKWVFSSGKKSSYTSPPAKWVKRFTGTDNQNKAPVSKKFAYNNNYKGKNPMT 564
T SPTEKW+ + G+K S++ + T N+ P +GK P
Sbjct: 321 TQKQASPTEKWIINVGQKLSWSKNTFSETRVITPESNRIIHP-----------EGKQP-- 181
Query: 565 KT*WRRFQRQKKVDGLKYVTNN---DKGKNKQMETFKMIR 601
+++K+ D LKY N K N + F MIR
Sbjct: 180 -------RKEKEEDPLKYSFMNIAEYKILNVILRKFAMIR 82
>TC85575 similar to PIR|D86410|D86410 protein F3M18.16 [imported] -
Arabidopsis thaliana, partial (15%)
Length = 1225
Score = 30.4 bits (67), Expect = 4.7
Identities = 26/98 (26%), Positives = 40/98 (40%), Gaps = 5/98 (5%)
Frame = +3
Query: 473 DNKPKFIFNKRGVPYKDTPPTRHQNRGQVRTFTPSSKSPTEKWVFSSGKKSSYTSPPAKW 532
+NK FN Y+ ++ N TP+S S + + + K SS + +
Sbjct: 369 NNKVNNYFN-----YQQDSDNKYPNEVSDTKLTPTSYSSSSNNNYDNNKYSSSSDDFSN- 530
Query: 533 VKRF-----TGTDNQNKAPVSKKFAYNNNYKGKNPMTK 565
K+F +NQN K+ YNNN K + TK
Sbjct: 531 -KKFPEEGYNSNENQNNNNNDNKYFYNNNNKDSSSNTK 641
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.160 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,876,290
Number of Sequences: 36976
Number of extensions: 546194
Number of successful extensions: 5783
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1733
Number of HSP's successfully gapped in prelim test: 311
Number of HSP's that attempted gapping in prelim test: 3981
Number of HSP's gapped (non-prelim): 2233
length of query: 1312
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1204
effective length of database: 5,021,319
effective search space: 6045668076
effective search space used: 6045668076
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC124217.1


