

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.5 - phase: 0
(340 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
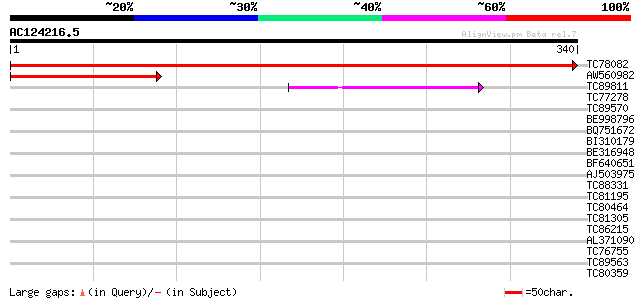
TC78082 similar to GP|21618292|gb|AAM67342.1 unknown {Arabidopsi... 702 0.0
AW560982 homologue to GP|21618292|gb| unknown {Arabidopsis thali... 182 2e-46
TC89811 similar to GP|16604350|gb|AAL24181.1 At1g14680/F10B6_22 ... 83 1e-16
TC77278 similar to GP|11493822|gb|AAG35658.1 transcription facto... 33 0.21
TC89570 similar to GP|13936308|gb|AAK40307.1 putative methyl-bin... 31 0.60
BE998796 similar to GP|5441893|dbj| ESTs C99174(E10437) D22295(C... 30 1.3
BQ751672 weakly similar to GP|16945435|em conserved hypothetical... 30 1.7
BI310179 similar to PIR|T52381|T523 zinc finger protein Zat1 C2... 30 1.7
BE316948 similar to GP|14423554|gb| Unknown protein {Arabidopsis... 28 3.9
BF640651 28 5.1
AJ503975 similar to PIR|H96731|H96 hypothetical protein F5A18.8 ... 28 6.6
TC88331 weakly similar to GP|21593279|gb|AAM65228.1 unknown {Ara... 28 6.6
TC81195 similar to GP|15810597|gb|AAL07186.1 unknown protein {Ar... 28 6.6
TC80464 weakly similar to GP|13548328|emb|CAC35875. putative pro... 28 6.6
TC81305 similar to PIR|C83591|C83591 N-carbamoyl-beta-alanine am... 28 6.6
TC86215 similar to GP|19698867|gb|AAL91169.1 unknown protein {Ar... 28 6.6
AL371090 28 6.6
TC76755 similar to PIR|S70586|S70586 proline-rich protein 14K -... 27 8.7
TC89563 similar to GP|15865943|gb|AAL10121.1 NADH dehydrogenase ... 27 8.7
TC80359 GP|1435157|emb|CAA58445.1 histone H3 variant H3.3 {Lycop... 27 8.7
>TC78082 similar to GP|21618292|gb|AAM67342.1 unknown {Arabidopsis
thaliana}, partial (60%)
Length = 1626
Score = 702 bits (1812), Expect = 0.0
Identities = 340/340 (100%), Positives = 340/340 (100%)
Frame = +2
Query: 1 MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA 60
MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA
Sbjct: 272 MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA 451
Query: 61 LAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVKHI 120
LAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVKHI
Sbjct: 452 LAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVKHI 631
Query: 121 HHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKT 180
HHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKT
Sbjct: 632 HHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKT 811
Query: 181 PKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQV 240
PKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQV
Sbjct: 812 PKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQV 991
Query: 241 AYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMS 300
AYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMS
Sbjct: 992 AYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMS 1171
Query: 301 GSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
GSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK
Sbjct: 1172GSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 1291
>AW560982 homologue to GP|21618292|gb| unknown {Arabidopsis thaliana},
partial (19%)
Length = 703
Score = 182 bits (462), Expect = 2e-46
Identities = 91/91 (100%), Positives = 91/91 (100%)
Frame = +2
Query: 1 MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA 60
MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA
Sbjct: 407 MDDRENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEKKAA 586
Query: 61 LAERDMAFLQRDTAIAERNNALMERDNAIAT 91
LAERDMAFLQRDTAIAERNNALMERDNAIAT
Sbjct: 587 LAERDMAFLQRDTAIAERNNALMERDNAIAT 679
>TC89811 similar to GP|16604350|gb|AAL24181.1 At1g14680/F10B6_22
{Arabidopsis thaliana}, partial (28%)
Length = 786
Score = 83.2 bits (204), Expect = 1e-16
Identities = 44/117 (37%), Positives = 61/117 (51%)
Frame = +2
Query: 168 PKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISK 227
P+ S + N + + VKKEG K + + +++ DD + K
Sbjct: 434 PQPDASRNENVENAEELSVKKEGGKSKKRK--SKGVVTTPKAKKPRKPKDDSSVSAQGVK 607
Query: 228 ADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLP 284
K L +N + D S +P P CSCTG +QCY+WG GGWQSACCTT +S+YPLP
Sbjct: 608 PPKKTLALEINGIEMDISGLPVPVCSCTGNPQQCYRWGVGGWQSACCTTNVSIYPLP 778
>TC77278 similar to GP|11493822|gb|AAG35658.1 transcription factor WRKY4
{Petroselinum crispum}, partial (40%)
Length = 1604
Score = 32.7 bits (73), Expect = 0.21
Identities = 27/96 (28%), Positives = 40/96 (41%), Gaps = 13/96 (13%)
Frame = +2
Query: 106 SCPPGCQISRGV------------KHIHHLPQQVNHLPNMGDS-SYGTRELHTTDALPAA 152
SCP ++ R V +H H P Q+ G S ++ + + PAA
Sbjct: 779 SCPVKKKVQRSVDDQSMLVATYEGEHNHPQPPQIESTSGSGRSVNHSSVPCSASLTSPAA 958
Query: 153 PVSLEVGKPPRRAKRPKESKSDSPNKKTPKSRKVKK 188
P + + +K K+SKS P K +PK KV K
Sbjct: 959 PKVVTLDSTT--SKNSKDSKSIEPRKDSPKEAKVPK 1060
>TC89570 similar to GP|13936308|gb|AAK40307.1 putative methyl-binding domain
protein MBD105 {Zea mays}, partial (9%)
Length = 728
Score = 31.2 bits (69), Expect = 0.60
Identities = 31/118 (26%), Positives = 50/118 (42%), Gaps = 12/118 (10%)
Frame = +3
Query: 126 QVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKR---------PKESKSDSP 176
+ N PN+ + +GT G+ PRR+ R P E K+D P
Sbjct: 282 KTNGGPNISEFDWGT------------------GETPRRSSRIVEKVKAAPPAEPKTDPP 407
Query: 177 NKKTPKSRKVKKEG--DDLNKTMFANNKELEWKSSEEIINGDD-DLNKQLAISKADWK 231
K++ KS KKE D+ +T K++E + + E + D +L K++ D K
Sbjct: 408 KKRSRKSSASKKEASEDEAEET-----KDVEMQEAGETKDDKDVELEKKVVNENEDKK 566
>BE998796 similar to GP|5441893|dbj| ESTs C99174(E10437) D22295(C10709)
correspond to a region of the predicted gene.~Similar
to, partial (28%)
Length = 609
Score = 30.0 bits (66), Expect = 1.3
Identities = 25/115 (21%), Positives = 42/115 (35%)
Frame = +1
Query: 104 MSSCPPGCQISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPR 163
+ +C GC +HH+P N + M ++ T A P AP ++ PR
Sbjct: 157 IQTCTSGCPFGESCHFLHHVPGGYNAVSQM---------MNLTPAAPPAPRNVPA---PR 300
Query: 164 RAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDD 218
A P S + + + +K FA+ EW+ + + DD
Sbjct: 301 NAHAPNGSAPSAVKSRICSKFNTAEGCKFGDKCHFAHG---EWELGKPVAPSFDD 456
>BQ751672 weakly similar to GP|16945435|em conserved hypothetical protein
{Neurospora crassa}, partial (16%)
Length = 756
Score = 29.6 bits (65), Expect = 1.7
Identities = 17/64 (26%), Positives = 29/64 (44%), Gaps = 3/64 (4%)
Frame = -2
Query: 130 LPNMGDSSYGTRELHTTDALPAAPVSLEVGK---PPRRAKRPKESKSDSPNKKTPKSRKV 186
LP + ++ TR P P S +G+ PP +RPK + + PN T +
Sbjct: 587 LPRLATATRATRP----SPRPTVPCSSCIGRAAEPPSNRRRPKRTAAQRPNPSTCSAPPA 420
Query: 187 KKEG 190
+++G
Sbjct: 419 RRKG 408
>BI310179 similar to PIR|T52381|T523 zinc finger protein Zat1 C2H2-type
[imported] - Arabidopsis thaliana, partial (16%)
Length = 704
Score = 29.6 bits (65), Expect = 1.7
Identities = 31/113 (27%), Positives = 49/113 (42%), Gaps = 4/113 (3%)
Frame = +2
Query: 136 SSYGTRELHTTDALPAAPVS-LEVGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLN 194
SSYG RE P ++ G + R E++S S N P+S++VKK+ D +
Sbjct: 245 SSYGLRENPKRSFRFTDPNQFIDAGSIILQEDRGSETES-SKNPTGPRSKRVKKDNDPV- 418
Query: 195 KTMFANNKELEWKSSEEIINGDDDLNKQLAISKAD---WKPQDLALNQVAYDD 244
K N SS +I ++ +++S+ D WK +Q DD
Sbjct: 419 KNSVQNESFSVLSSSSDITTEEEVAFFLISLSREDNNIWKTHTQRHDQQHQDD 577
>BE316948 similar to GP|14423554|gb| Unknown protein {Arabidopsis thaliana},
partial (17%)
Length = 234
Score = 28.5 bits (62), Expect = 3.9
Identities = 19/61 (31%), Positives = 28/61 (45%)
Frame = -2
Query: 120 IHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKK 179
+H LP++ D YGT LH+ LP AP L + R+ R +E P K+
Sbjct: 233 LHFLPRR--------DLRYGTSFLHSLPVLPLAP*GLTM---KTRSLRREEEDESMPRKR 87
Query: 180 T 180
+
Sbjct: 86 S 84
>BF640651
Length = 468
Score = 28.1 bits (61), Expect = 5.1
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = +3
Query: 316 HDLSHPVDLKDHWAKHGTNRY 336
H++SHP LKD K G NRY
Sbjct: 126 HNISHPFRLKDSPKKCGDNRY 188
>AJ503975 similar to PIR|H96731|H96 hypothetical protein F5A18.8 [imported] -
Arabidopsis thaliana, partial (33%)
Length = 381
Score = 27.7 bits (60), Expect = 6.6
Identities = 13/27 (48%), Positives = 19/27 (70%), Gaps = 2/27 (7%)
Frame = +3
Query: 191 DDLNKTMFANNK--ELEWKSSEEIING 215
+ L+K +F +NK EL+WK +IING
Sbjct: 213 ESLDKFLFRSNKKQELDWKRRFDIING 293
>TC88331 weakly similar to GP|21593279|gb|AAM65228.1 unknown {Arabidopsis
thaliana}, partial (43%)
Length = 1085
Score = 27.7 bits (60), Expect = 6.6
Identities = 20/83 (24%), Positives = 30/83 (36%), Gaps = 14/83 (16%)
Frame = +1
Query: 21 WLMQQHQHQHPSMKQIMSIM--------AERDAAIQERNLALSEKKAALAERDMAF---- 68
WL+ H H HP+ S + + +I R+ +LS + L D
Sbjct: 40 WLLHFHHHHHPTSSTKFSALYHHLLLPTSSPQESINIRSFSLSASRGKLRRADSPLPTAP 219
Query: 69 --LQRDTAIAERNNALMERDNAI 89
+ D AI RN E A+
Sbjct: 220 EEEEEDAAIKSRNQLKREAKRAV 288
>TC81195 similar to GP|15810597|gb|AAL07186.1 unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 788
Score = 27.7 bits (60), Expect = 6.6
Identities = 16/60 (26%), Positives = 25/60 (41%), Gaps = 6/60 (10%)
Frame = +3
Query: 125 QQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGK------PPRRAKRPKESKSDSPNK 178
+ V+H MG+ E + P P+ E K P K+P+ESK +P +
Sbjct: 102 ESVSHTKPMGEEEKKPEETKVEEKKPEQPLKKEEEKKETEKPPTEEEKKPEESKEVTPRE 281
>TC80464 weakly similar to GP|13548328|emb|CAC35875. putative protein
{Arabidopsis thaliana}, partial (19%)
Length = 1437
Score = 27.7 bits (60), Expect = 6.6
Identities = 15/42 (35%), Positives = 21/42 (49%)
Frame = +3
Query: 177 NKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDD 218
N K+ KSR +K +G + F + EW SE + DDD
Sbjct: 636 NPKSDKSRDIKHDGVISDMDYFKSKVTTEWSDSE---SSDDD 752
>TC81305 similar to PIR|C83591|C83591 N-carbamoyl-beta-alanine
amidohydrolase PA0444 [imported] - Pseudomonas
aeruginosa (strain PAO1), partial (16%)
Length = 308
Score = 27.7 bits (60), Expect = 6.6
Identities = 23/81 (28%), Positives = 30/81 (36%), Gaps = 2/81 (2%)
Frame = -3
Query: 103 GMSSCPPGCQISRGVKHI--HHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGK 160
G+ G Q + G +H HH PQQ H P + +R + L A S
Sbjct: 300 GVKPGDSGSQFAAGNRHRRRHHRPQQHRHTPPPDHPARHSRFRYD*CRLHTAQRSSPPAP 121
Query: 161 PPRRAKRPKESKSDSPNKKTP 181
PRR R +S P P
Sbjct: 120 APRRKSRHALPRSQCPCSTGP 58
>TC86215 similar to GP|19698867|gb|AAL91169.1 unknown protein {Arabidopsis
thaliana}, partial (91%)
Length = 2293
Score = 27.7 bits (60), Expect = 6.6
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -3
Query: 274 CTTTLSVYPLPAVPNKRHAR 293
CT S+YP+ PN RH R
Sbjct: 1991 CTLPCSIYPVSTDPNNRHCR 1932
>AL371090
Length = 425
Score = 27.7 bits (60), Expect = 6.6
Identities = 15/54 (27%), Positives = 27/54 (49%), Gaps = 1/54 (1%)
Frame = +3
Query: 176 PNKKTPKSRKVKKE-GDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKA 228
P KK K +++++E G K ++L +++ +DD KQL + KA
Sbjct: 129 PTKKMKKRKEMEQERGKKRGKVQSNKKQKLSKTQKKKLKKSEDDKEKQLLLEKA 290
>TC76755 similar to PIR|S70586|S70586 proline-rich protein 14K - kidney
bean, partial (85%)
Length = 789
Score = 27.3 bits (59), Expect = 8.7
Identities = 17/57 (29%), Positives = 23/57 (39%)
Frame = +1
Query: 107 CPPGCQISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPR 163
CPP G H HH P N +G D L +++E+GKPP+
Sbjct: 151 CPPPPHKDHGHSHPHHPPSSKNPTCPRDTIKFGV----CADVL--GLINVELGKPPK 303
>TC89563 similar to GP|15865943|gb|AAL10121.1 NADH dehydrogenase subunit 2
{Varanus salvator cumingi}, partial (6%)
Length = 1400
Score = 27.3 bits (59), Expect = 8.7
Identities = 14/30 (46%), Positives = 19/30 (62%), Gaps = 1/30 (3%)
Frame = -3
Query: 161 PPRRAKRPKE-SKSDSPNKKTPKSRKVKKE 189
PP A RPKE S+ + K TPK+++ K E
Sbjct: 1134 PPEEAPRPKEPSQKVNEQKGTPKAKEEKVE 1045
>TC80359 GP|1435157|emb|CAA58445.1 histone H3 variant H3.3 {Lycopersicon
esculentum}, complete
Length = 1440
Score = 27.3 bits (59), Expect = 8.7
Identities = 14/32 (43%), Positives = 16/32 (49%), Gaps = 1/32 (3%)
Frame = +1
Query: 243 DDSTMPAPACSCT-GVLRQCYKWGNGGWQSAC 273
D S +P P CSCT G R W G+Q C
Sbjct: 292 DGSQIPEPCCSCTSGSSRSLSGWIV*GYQFVC 387
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.128 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,524,567
Number of Sequences: 36976
Number of extensions: 147011
Number of successful extensions: 859
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 835
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 847
length of query: 340
length of database: 9,014,727
effective HSP length: 97
effective length of query: 243
effective length of database: 5,428,055
effective search space: 1319017365
effective search space used: 1319017365
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC124216.5


