

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.10 - phase: 0
(543 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
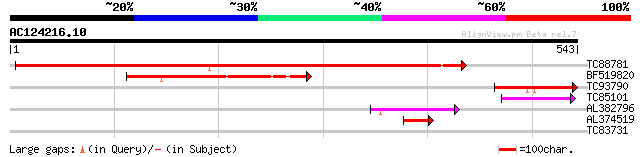
Score E
Sequences producing significant alignments: (bits) Value
TC88781 similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Arabidops... 664 0.0
BF519820 weakly similar to GP|15809876|gb AT3g53950/F5K20_250 {A... 174 8e-44
TC93790 weakly similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Ar... 110 1e-24
TC85101 similar to GP|15809876|gb|AAL06866.1 AT3g53950/F5K20_250... 57 2e-08
AL382796 weakly similar to GP|20161084|dbj putative glyoxal oxid... 54 2e-07
AL374519 similar to GP|15809876|gb| AT3g53950/F5K20_250 {Arabido... 47 2e-05
TC83731 similar to PIR|T46155|T46155 hypothetical protein T4D2.2... 28 9.0
>TC88781 similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Arabidopsis
thaliana}, partial (67%)
Length = 1391
Score = 664 bits (1712), Expect = 0.0
Identities = 311/435 (71%), Positives = 362/435 (82%), Gaps = 3/435 (0%)
Frame = +1
Query: 6 YLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLS 65
+LL L I+I+T T P A GQWQ+LQ +IGIVAMHMQLLHNDR++IFDRTDFG S
Sbjct: 88 HLLIPLIIIISTTFTIPSPAIAAGGQWQLLQNNIGIVAMHMQLLHNDRVIIFDRTDFGFS 267
Query: 66 KLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQ 125
L LPNG+CR +P+E VK DCTAHS+EY++ SNTFRPLF+QT++WCSSGSVNP GTL+Q
Sbjct: 268 NLSLPNGRCRVNPQEQAVKNDCTAHSLEYDVFSNTFRPLFIQTNIWCSSGSVNPNGTLIQ 447
Query: 126 TGGYNDGDRTIRMFDTCNNCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFY 185
TGG+NDGDR++R F+ C NCDWQEF+ L+ARRWYATNH LPDGRQIIIGGR+QFNYEFY
Sbjct: 448 TGGFNDGDRSVRTFNPCPNCDWQEFNASLSARRWYATNHKLPDGRQIIIGGRRQFNYEFY 627
Query: 186 PKNNI---GVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVR 242
PK + Y LPFL QTNDA AENNLYPFV LNVDGNLFIFANNRAILFDY N VV+
Sbjct: 628 PKKDATAKNTYSLPFLVQTNDASAENNLYPFVFLNVDGNLFIFANNRAILFDYNSNKVVK 807
Query: 243 TFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGALN 302
T+P IPGGDPRSYPSSGS LLPL+NLQ+ +EAEV+ICGGAPKGSY+K + F+GALN
Sbjct: 808 TYPTIPGGDPRSYPSSGSAALLPLRNLQNPLVEAEVMICGGAPKGSYEKTLQGSFIGALN 987
Query: 303 TCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPV 362
TC R+KITDPNP+WV+ETMP RVM DM++LPNGDVL+INGA G+AGWE GR+PVLNP
Sbjct: 988 TCGRLKITDPNPSWVMETMPGGRVMSDMILLPNGDVLLINGAAVGSAGWESGRNPVLNPF 1167
Query: 363 LYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSI 422
LYK N+ +G+RF+LQ PS PRMYHSTA+L+RDGRVLV GSNPH+ YNF LFPTEL +
Sbjct: 1168LYKPNSLVGSRFQLQKPSDIPRMYHSTAVLLRDGRVLVAGSNPHVNYNFTR-LFPTELRL 1344
Query: 423 EAFSPSYLEPRFANV 437
EAFSPSYLE F +V
Sbjct: 1345EAFSPSYLELGFNDV 1389
>BF519820 weakly similar to GP|15809876|gb AT3g53950/F5K20_250 {Arabidopsis
thaliana}, partial (34%)
Length = 574
Score = 174 bits (441), Expect = 8e-44
Identities = 91/181 (50%), Positives = 125/181 (68%), Gaps = 4/181 (2%)
Frame = +1
Query: 113 SSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNN---CDWQEF-DGGLAARRWYATNHILPD 168
SSGS P G+L+QTGG DG + IR+F+ C++ CDW+E D L+ RWYATN ILPD
Sbjct: 4 SSGSFLPNGSLLQTGGDLDGFKKIRIFNPCDSTGSCDWEELNDVELSEGRWYATNQILPD 183
Query: 169 GRQIIIGGRKQFNYEFYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANN 228
G IIIGGR EFYPK GV PFL++T D + NLYP+V L +G LF+FAN
Sbjct: 184 GSIIIIGGRGSNTVEFYPKRENGVVLFPFLQETEDTQMD-NLYPYVHLLPNGKLFVFANT 360
Query: 229 RAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGS 288
+++++D+ +N VVR +P++ GG PR+YPS+GS V+L L+ +F EA V++CGGA G+
Sbjct: 361 KSVMYDFNENRVVRVYPELTGG-PRNYPSAGSSVMLA---LEGEFSEAVVVVCGGAQYGA 528
Query: 289 Y 289
+
Sbjct: 529 F 531
>TC93790 weakly similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Arabidopsis
thaliana}, partial (11%)
Length = 574
Score = 110 bits (275), Expect = 1e-24
Identities = 54/83 (65%), Positives = 68/83 (81%), Gaps = 4/83 (4%)
Frame = +1
Query: 465 LDKNLVYVTMLAPPFNTHSFSMNQRLLVLE--SNKVNIV--EGTTYDVQVTMPGSPILAP 520
L KN+V VTM AP FNTHSFSMNQRLLVL+ S+ N+V +TY V+V +P SPILAP
Sbjct: 4 LMKNMVSVTMYAPSFNTHSFSMNQRLLVLDELSSSKNVVGESTSTYQVEVAIPNSPILAP 183
Query: 521 PGFYLLFVVHKEIPSEGIWIQIL 543
PG+YLLFVVH+E+PS+G+W+Q+L
Sbjct: 184 PGYYLLFVVHQEVPSQGVWVQLL 252
>TC85101 similar to GP|15809876|gb|AAL06866.1 AT3g53950/F5K20_250
{Arabidopsis thaliana}, partial (12%)
Length = 519
Score = 57.0 bits (136), Expect = 2e-08
Identities = 27/71 (38%), Positives = 42/71 (59%)
Frame = +2
Query: 472 VTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHK 531
V + + PF THSFS QRL+ L + + ++ T P S ++APPG+Y+LF V++
Sbjct: 5 VNLGSAPFATHSFSQGQRLIKLGVASAMVDGDQRWRIRCTAPPSGMVAPPGYYMLFAVNQ 184
Query: 532 EIPSEGIWIQI 542
+PS WI +
Sbjct: 185 GVPSVARWIHM 217
>AL382796 weakly similar to GP|20161084|dbj putative glyoxal oxidase {Oryza
sativa (japonica cultivar-group)}, partial (4%)
Length = 465
Score = 53.9 bits (128), Expect = 2e-07
Identities = 34/97 (35%), Positives = 45/97 (46%), Gaps = 12/97 (12%)
Frame = +3
Query: 346 SGTAGWEY-----------GRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVR 394
+G AGW+ P NP+LY + G+R+ R+YHS+A L+
Sbjct: 3 TGMAGWDKKATNKTPRLHTAGKPTRNPLLYNPHEKKGSRWSKFTEDPIIRVYHSSATLIP 182
Query: 395 DGRVLVGGSNPHIGY-NFNNVLFPTELSIEAFSPSYL 430
DG V V GSNP+ Y + PTE E F P YL
Sbjct: 183 DGTVFVAGSNPNRYYCGMDQCEHPTEHRAEKFYPPYL 293
>AL374519 similar to GP|15809876|gb| AT3g53950/F5K20_250 {Arabidopsis
thaliana}, partial (17%)
Length = 532
Score = 47.0 bits (110), Expect = 2e-05
Identities = 20/29 (68%), Positives = 23/29 (78%)
Frame = +1
Query: 378 NPSHTPRMYHSTAILVRDGRVLVGGSNPH 406
NP PR+YHSTA L+ DGRVL+ GSNPH
Sbjct: 10 NPGTVPRLYHSTANLLPDGRVLLAGSNPH 96
Score = 28.1 bits (61), Expect = 9.0
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +3
Query: 469 LVYVTMLAPPFNTHSFSMNQRLLVL 493
++ V + + PF THSFS QRL+ L
Sbjct: 294 IIEVNLGSAPFATHSFSQGQRLIKL 368
>TC83731 similar to PIR|T46155|T46155 hypothetical protein T4D2.20 -
Arabidopsis thaliana, partial (27%)
Length = 954
Score = 28.1 bits (61), Expect = 9.0
Identities = 10/14 (71%), Positives = 13/14 (92%)
Frame = +2
Query: 126 TGGYNDGDRTIRMF 139
TGGYN+G RTI++F
Sbjct: 611 TGGYNEGSRTIKIF 652
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,578,384
Number of Sequences: 36976
Number of extensions: 258998
Number of successful extensions: 1143
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1123
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1131
length of query: 543
length of database: 9,014,727
effective HSP length: 101
effective length of query: 442
effective length of database: 5,280,151
effective search space: 2333826742
effective search space used: 2333826742
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124216.10


