

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.5 - phase: 0
(266 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
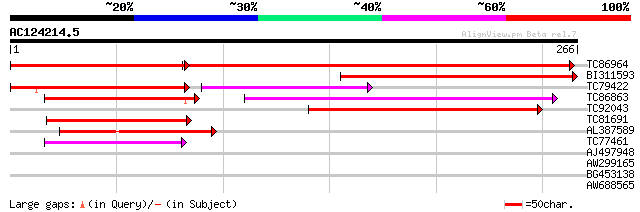
TC86964 similar to PIR|T47953|T47953 hypothetical protein F2A19.... 274 3e-74
BI311593 similar to PIR|T47953|T47 hypothetical protein F2A19.21... 213 8e-56
TC79422 similar to PIR|T47953|T47953 hypothetical protein F2A19.... 102 2e-22
TC86863 similar to GP|19387263|gb|AAL87174.1 putative apospory-a... 89 1e-18
TC92043 similar to GP|21554299|gb|AAM63374.1 apospory-associated... 86 1e-17
TC81691 weakly similar to GP|6016717|gb|AAF01543.1| unknown prot... 69 2e-12
AL387589 weakly similar to GP|19387263|gb| putative apospory-ass... 62 3e-10
TC77461 similar to GP|10177534|dbj|BAB10929. apospory-associated... 55 3e-08
AJ497948 similar to GP|8777366|dbj| apospory-associated protein ... 38 0.005
AW299165 35 0.023
BG453138 similar to GP|10177534|dbj apospory-associated protein ... 28 4.8
AW688565 similar to PIR|D96617|D96 probable xylan endohydrolase ... 27 8.2
>TC86964 similar to PIR|T47953|T47953 hypothetical protein F2A19.210 -
Arabidopsis thaliana, partial (98%)
Length = 1404
Score = 274 bits (700), Expect = 3e-74
Identities = 148/184 (80%), Positives = 159/184 (85%)
Frame = +3
Query: 82 AGHIVLSSAFECLLQQMET*P*YHE*GISMPSHLVSHLHTTHTCWFLT*VR*GLKVLRHL 141
AGHIVLSS FEC LQQ ET*P*YHE GISM SHLVSH HTTHT WFLT VR*GLKVLRHL
Sbjct: 606 AGHIVLSSGFECRLQQTET*P*YHESGISMASHLVSHSHTTHTYWFLTLVR*GLKVLRHL 785
Query: 142 ITWTTFPKKHELQNKEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCG 201
ITWTTFPKK++LQNKEM *HLNPRWIVFILA Q * QF*IMRGN+HIL*+RKVSLML CG
Sbjct: 786 ITWTTFPKKNDLQNKEMR*HLNPRWIVFILALQI*LQF*IMRGNKHIL*KRKVSLMLRCG 965
Query: 202 IHGRRNRSRWWTSVMKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQLCDQVFVVTV*V 261
I GRRN+SR T VM+STN+CFV*M Q *R+ *T* ER+GQGGF+S+LC QVFVVTV V
Sbjct: 966 IRGRRNQSR*RTLVMRSTNRCFV*MQQ**RHQ*T*GPERNGQGGFSSRLCHQVFVVTVSV 1145
Query: 262 STEV 265
TEV
Sbjct: 1146WTEV 1157
Score = 147 bits (371), Expect = 4e-36
Identities = 72/84 (85%), Positives = 75/84 (88%)
Frame = +2
Query: 1 MGHSAAVWDFRAATEFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTS 60
MGHSAAVWDFRAATE TKD NG+ QVVLRTPQGASARVSL+GAQVTSWRNEQGEELLFTS
Sbjct: 212 MGHSAAVWDFRAATEMTKDQNGVSQVVLRTPQGASARVSLNGAQVTSWRNEQGEELLFTS 391
Query: 61 CKTLSKAPKAIRGGIPICFPQAGH 84
KT KA KA RGGIPICFPQ G+
Sbjct: 392 SKTTPKAQKAARGGIPICFPQFGN 463
>BI311593 similar to PIR|T47953|T47 hypothetical protein F2A19.210 -
Arabidopsis thaliana, partial (33%)
Length = 423
Score = 213 bits (541), Expect = 8e-56
Identities = 110/111 (99%), Positives = 111/111 (99%)
Frame = +2
Query: 156 KEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSV 215
+EML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSV
Sbjct: 2 EEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSV 181
Query: 216 MKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQLCDQVFVVTV*VSTEVV 266
MKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQLCDQVFVVTV*VSTEVV
Sbjct: 182 MKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQLCDQVFVVTV*VSTEVV 334
>TC79422 similar to PIR|T47953|T47953 hypothetical protein F2A19.210 -
Arabidopsis thaliana, partial (60%)
Length = 1650
Score = 102 bits (254), Expect = 2e-22
Identities = 52/85 (61%), Positives = 61/85 (71%), Gaps = 1/85 (1%)
Frame = +2
Query: 1 MGHSAAVWDFR-AATEFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFT 59
M HS A D + + E KD NGI+QVVLR +GASA VSLHG QV SW+ E+GEELLFT
Sbjct: 1004 MNHSGAESDHKITSVEVIKDKNGIEQVVLRNVRGASATVSLHGGQVLSWKTERGEELLFT 1183
Query: 60 SCKTLSKAPKAIRGGIPICFPQAGH 84
S K + PK +RGGIPICFPQ G+
Sbjct: 1184 STKAVYTPPKPVRGGIPICFPQFGN 1258
Score = 53.9 bits (128), Expect = 6e-08
Identities = 35/80 (43%), Positives = 45/80 (55%)
Frame = +1
Query: 91 FECLLQQMET*P*YHE*GISMPSHLVSHLHTTHTCWFLT*VR*GLKVLRHLITWTTFPKK 150
+E + M * *Y G+SM S VS LH H + +*G K + L W T+ ++
Sbjct: 1390 YEDMAT*MAV*I*YLASGMSMASLSVSPLHFIHISRYPISTK*G*KGWKLLTIWITYTRR 1569
Query: 151 HELQNKEML*HLNPRWIVFI 170
+ L NKEML*HLN RWI FI
Sbjct: 1570 NALLNKEML*HLNLRWIEFI 1629
>TC86863 similar to GP|19387263|gb|AAL87174.1 putative apospory-associated
protein C {Oryza sativa (japonica cultivar-group)},
partial (88%)
Length = 1640
Score = 89.4 bits (220), Expect = 1e-18
Identities = 45/74 (60%), Positives = 54/74 (72%), Gaps = 1/74 (1%)
Frame = +1
Query: 17 TKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIP 76
TK NG+D+V+LR +GASA + L+G VTSW+N+ EELLF S K + K PKAIRGGIP
Sbjct: 238 TKGINGLDKVLLRESRGASAEIYLYGGHVTSWKNDHAEELLFLSSKAIFKPPKAIRGGIP 417
Query: 77 ICFPQ-AGHIVLSS 89
ICFPQ A H L S
Sbjct: 418 ICFPQFANHGNLES 459
Score = 68.6 bits (166), Expect = 2e-12
Identities = 56/147 (38%), Positives = 75/147 (50%)
Frame = +2
Query: 111 MPSHLVSHLHTTHTCWFLT*VR*GLKVLRHLITWTTFPKKHELQNKEML*HLNPRWIVFI 170
M SH HL C F T*V+ G + R I TT + L N+EML* LN + I +I
Sbjct: 674 MGSHFRLHLLIILICRFQT*VKFGSRD*RPWIILTTCRTRSALLNREML*LLNQKLIRYI 853
Query: 171 LAPQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL* 230
L Q IM+ H+ + LML CGI G R + W S M S + CFV* + +
Sbjct: 854 LVLLQRLQSLIMKKRGHLFCGKMAFLMLWCGILGIRKQRLWLISGMMSISICFV*RLLIL 1033
Query: 231 RNL*T*SLERSGQGGFNSQLCDQVFVV 257
+ L *+LE++G+ + QL V VV
Sbjct: 1034KRLSL*NLEKNGKED*SFQLFHLVTVV 1114
>TC92043 similar to GP|21554299|gb|AAM63374.1 apospory-associated protein C
{Arabidopsis thaliana}, partial (37%)
Length = 1163
Score = 85.9 bits (211), Expect = 1e-17
Identities = 53/110 (48%), Positives = 68/110 (61%)
Frame = +3
Query: 141 LITWTTFPKKHELQNKEML*HLNPRWIVFILAPQT*SQF*IMRGNEHIL*ERKVSLMLPC 200
L W T+ +++ NKEML*HLN RWI FI F*I RG+EH+ ERKV ML
Sbjct: 516 LTIWITYTRRNASLNKEML*HLNLRWIEFISILAVW*PF*ITRGSEHL*YERKVFQMLLF 695
Query: 201 GIHGRRNRSRWWTSVMKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQL 250
GI GR+N+S+ W MKST+KC MVQ + * *S E++G ++S L
Sbjct: 696 GIPGRKNQSQ*WILGMKSTSKCSAWMVQQLESQ*P*SREKNGPAVWSSPL 845
>TC81691 weakly similar to GP|6016717|gb|AAF01543.1| unknown protein
{Arabidopsis thaliana}, complete
Length = 1465
Score = 68.6 bits (166), Expect = 2e-12
Identities = 32/68 (47%), Positives = 45/68 (66%)
Frame = +3
Query: 18 KDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPI 77
+D +G+ ++VL P G+SA V L+G + SW+N++ EELLF S K K KAIRGGI
Sbjct: 78 QDKDGLPRIVLTDPNGSSAEVLLYGGHIVSWKNQRKEELLFMSSKAKWKQHKAIRGGISA 257
Query: 78 CFPQAGHI 85
CF + G +
Sbjct: 258 CFARFGDV 281
Score = 34.7 bits (78), Expect = 0.039
Identities = 41/145 (28%), Positives = 60/145 (41%)
Frame = +1
Query: 113 SHLVSHLHTTHTCWFLT*VR*GLKVLRHLITWTTFPKKHELQNKEML*HLNPRWIVFILA 172
+H + H C + V+ + RHLI + QN+ M HL R I +
Sbjct: 502 NHFLLHFRYATICLYQISVKCVSRD*RHLIMLII**IDQDSQNRLMQLHLMERRIEYTCT 681
Query: 173 PQT*SQF*IMRGNEHIL*ERKVSLMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL*RN 232
Q *IMR E + R M GIHG + R + + T C+V*+ QL
Sbjct: 682 AQIKLPS*IMRRKELLYSRRIHFQMQLYGIHGIKRRKLYLIWELMITK*CYV*IQQLLIL 861
Query: 233 L*T*SLERSGQGGFNSQLCDQVFVV 257
L +*+ ++G+ QL Q V
Sbjct: 862 LLS*NHVKNGKVIKKFQLFHQAIAV 936
>AL387589 weakly similar to GP|19387263|gb| putative apospory-associated
protein C {Oryza sativa (japonica cultivar-group)},
partial (7%)
Length = 472
Score = 61.6 bits (148), Expect = 3e-10
Identities = 35/74 (47%), Positives = 45/74 (60%)
Frame = +3
Query: 24 DQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPICFPQAG 83
D+V+L+ +G+SA + GA VTSW+ +G+ELLF S K+ K IRGGIPI FPQ
Sbjct: 6 DKVILKDSKGSSAEIYYFGATVTSWKY-RGKELLFLSKKSFLDGTKGIRGGIPIIFPQFA 182
Query: 84 HIVLSSAFECLLQQ 97
SSA L Q
Sbjct: 183KATDSSAETAALPQ 224
>TC77461 similar to GP|10177534|dbj|BAB10929. apospory-associated protein
C-like {Arabidopsis thaliana}, partial (89%)
Length = 1323
Score = 55.1 bits (131), Expect = 3e-08
Identities = 26/67 (38%), Positives = 38/67 (55%)
Frame = +3
Query: 17 TKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIP 76
T+ + ++VL +P G+ A + L G +TSW+ G++LLF + K I GGIP
Sbjct: 156 TEGEGNLPKLVLTSPAGSEAEIYLFGGCITSWKVPNGKDLLFVRPDAVFNKKKPISGGIP 335
Query: 77 ICFPQAG 83
CFPQ G
Sbjct: 336 HCFPQFG 356
>AJ497948 similar to GP|8777366|dbj| apospory-associated protein C
{Arabidopsis thaliana}, partial (25%)
Length = 520
Score = 37.7 bits (86), Expect = 0.005
Identities = 23/56 (41%), Positives = 32/56 (57%)
Frame = +1
Query: 196 LMLPCGIHGRRNRSRWWTSVMKSTNKCFV*MVQL*RNL*T*SLERSGQGGFNSQLC 251
LML CGI G R + W S M S + CFV* + + + L *+LE++G+ LC
Sbjct: 115 LMLWCGILGIRKQRLWLISGMMSISICFV*RLLILKRLSL*NLEKNGKKDHCCVLC 282
>AW299165
Length = 456
Score = 35.4 bits (80), Expect = 0.023
Identities = 20/40 (50%), Positives = 25/40 (62%), Gaps = 1/40 (2%)
Frame = +2
Query: 1 MGHSAAVWDFR-AATEFTKDWNGIDQVVLRTPQGASARVS 39
M HS A D + + E KD NGI+QVVLR +GAS V+
Sbjct: 302 MNHSGAESDHKITSVEVIKDKNGIEQVVLRNVRGASTTVT 421
>BG453138 similar to GP|10177534|dbj apospory-associated protein C-like
{Arabidopsis thaliana}, partial (28%)
Length = 538
Score = 27.7 bits (60), Expect = 4.8
Identities = 13/27 (48%), Positives = 14/27 (51%)
Frame = +1
Query: 57 LFTSCKTLSKAPKAIRGGIPICFPQAG 83
LF + K I GGIP CFPQ G
Sbjct: 1 LFVRPDAVFNKKKPISGGIPHCFPQFG 81
>AW688565 similar to PIR|D96617|D96 probable xylan endohydrolase isoenzyme
F9K23.10 [imported] - Arabidopsis thaliana, partial
(11%)
Length = 614
Score = 26.9 bits (58), Expect = 8.2
Identities = 16/43 (37%), Positives = 22/43 (50%), Gaps = 5/43 (11%)
Frame = +2
Query: 10 FRAATEFTKDWNGIDQVVLRTPQ-----GASARVSLHGAQVTS 47
F +ATE T+ WNGI Q + Q +A V + G VT+
Sbjct: 326 FASATERTQSWNGIQQEITGRVQRKLAYEVTALVRIFGNNVTT 454
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.351 0.152 0.549
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,128,477
Number of Sequences: 36976
Number of extensions: 134127
Number of successful extensions: 1277
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1268
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1276
length of query: 266
length of database: 9,014,727
effective HSP length: 94
effective length of query: 172
effective length of database: 5,538,983
effective search space: 952705076
effective search space used: 952705076
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124214.5


