

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
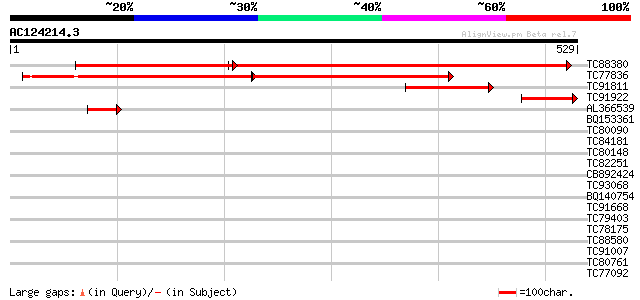
Query= AC124214.3 - phase: 0
(529 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88380 similar to GP|4324495|gb|AAD16897.1| glutamyl-tRNA reduc... 410 e-179
TC77836 similar to GP|4324495|gb|AAD16897.1| glutamyl-tRNA reduc... 266 e-139
TC91811 similar to GP|13676394|dbj|BAB41186. glutamyl-tRNA reduc... 162 4e-40
TC91922 similar to SP|P93111|HMA1_CUCSA Glutamyl-tRNA reductase ... 71 1e-12
AL366539 homologue to SP|P93111|HMA1_ Glutamyl-tRNA reductase 1 ... 61 1e-09
BQ153361 GP|146330|gb|A delta-aminolevulinate synthase (EC 2.3.1... 38 0.011
TC80090 similar to GP|15215794|gb|AAK91442.1 AT5g19180/T24G5_80 ... 33 0.21
TC84181 similar to SP|Q9SJA7|SOX_ARATH Potential sarcosine oxida... 31 1.0
TC80148 weakly similar to SP|Q39172|P1_ARATH Probable NADP-depen... 31 1.0
TC82251 31 1.3
CB892424 similar to PIR|T03964|T0 probable ubiquitin--protein li... 31 1.3
TC93068 similar to GP|3170523|gb|AAC18088.1| coatomer alpha subu... 30 2.3
BQ140754 similar to GP|22535581|dbj contains EST AU182189(C63973... 30 3.0
TC91668 similar to GP|11121445|emb|CAC14866. DNA Helicase {Arabi... 30 3.0
TC79403 homologue to GP|10190525|emb|CAC09322. unnamed protein p... 29 3.9
TC78175 similar to GP|1527001|gb|AAB07724.1| Ipomoea nil Pn47p, ... 29 3.9
TC88580 similar to GP|9758308|dbj|BAB08782.1 gene_id:MUA2.4~unkn... 29 3.9
TC91007 29 5.1
TC80761 homologue to GP|1370174|emb|CAA98164.1 RAB1Y {Lotus japo... 29 5.1
TC77092 squalene monooxygenase 2 [Medicago truncatula] 28 6.7
>TC88380 similar to GP|4324495|gb|AAD16897.1| glutamyl-tRNA reductase
precursor {Glycine max}, partial (92%)
Length = 1689
Score = 410 bits (1054), Expect(2) = e-179
Identities = 213/322 (66%), Positives = 264/322 (81%), Gaps = 2/322 (0%)
Frame = +3
Query: 205 KSGQGVPGFDRKISGLFKQAISVGKRVRTETNISSGSVSVSSAAVELALMKRLE-SSFGD 263
K + + F R ISGLFK AI+VGKRVRTETNI++G+VSVSSAAVELALMK + +S G+
Sbjct: 723 KQDKELMAFGRNISGLFKHAITVGKRVRTETNIAAGAVSVSSAAVELALMKLPDQASHGN 902
Query: 264 AKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMEC 323
A++LVIGAGKMGKLVIKHLVAKGC+KMVVVNR+E +V AIR+E+ DV+I+Y+PLSEM+EC
Sbjct: 903 ARMLVIGAGKMGKLVIKHLVAKGCKKMVVVNRTEGRVAAIREEINDVEIIYKPLSEMLEC 1082
Query: 324 AAEADVIFTGTASESLLFSKENVEILPPVGQGVR-RRLFVDISIPRNVDPGVSELENALV 382
EADV+FT TASE+ L K+NV+ LP + + +RLFVDIS+PRNV V +LE+ V
Sbjct: 1083 IGEADVVFTSTASENPLIFKQNVKELPLASEEMGGKRLFVDISVPRNVGSCVDDLESVKV 1262
Query: 383 YNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPTIKKFRAYVERIRA 442
YNVDDL+E V ANKEDR RKAMEA+ II EE FEAW+DSLETVPTIKK RAY ER+R
Sbjct: 1263 YNVDDLKEGVAANKEDRLRKAMEAQVIIGEESGQFEAWRDSLETVPTIKKLRAYAERLRV 1442
Query: 443 SELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDEVLENMRA 502
+ELEKCL KM D++K+ ++A+ LS GIVNK+LHGPMQHLRCDG+D++ L E LENM A
Sbjct: 1443 AELEKCLGKMGDDINKKTQKAVDDLSKGIVNKMLHGPMQHLRCDGSDSRTLSETLENMHA 1622
Query: 503 LNRMYDLETEISLMEEKIRVKM 524
LNRM+ LETE+S++E+KIR K+
Sbjct: 1623 LNRMFSLETEVSVLEQKIRAKV 1688
Score = 237 bits (604), Expect(2) = e-179
Identities = 115/151 (76%), Positives = 136/151 (89%)
Frame = +2
Query: 62 SPLEILKTSSADRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNHIE 121
S L+ LK S+ADRYTKE+SSI+VIGL++HT PV++REKLAIPEA+WP+ I ELC LNHIE
Sbjct: 293 SALQQLKASAADRYTKERSSIVVIGLSIHTTPVDVREKLAIPEAEWPRAIGELCNLNHIE 472
Query: 122 EAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQHLFE 181
EAAVLSTCNR+EIY+VALSQHRGV+EVT+W+S SG+ E+SK+ LLYNKDATQHLFE
Sbjct: 473 EAAVLSTCNRMEIYVVALSQHRGVKEVTEWMSNTSGIPASELSKYLFLLYNKDATQHLFE 652
Query: 182 VAAGLDSLVLGEGQILSQVKQVVKSGQGVPG 212
V+AGLDSLV+GEGQILSQVKQVVK+GQGV G
Sbjct: 653 VSAGLDSLVMGEGQILSQVKQVVKAGQGVNG 745
>TC77836 similar to GP|4324495|gb|AAD16897.1| glutamyl-tRNA reductase
precursor {Glycine max}, partial (64%)
Length = 1404
Score = 266 bits (681), Expect(2) = e-139
Identities = 142/219 (64%), Positives = 171/219 (77%), Gaps = 1/219 (0%)
Frame = +3
Query: 13 LRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLR-SDNPLPQNFIVSKPSPLEILKTSS 71
L+SS + P S F R +F I +C + SD + +S LE+LKTS+
Sbjct: 204 LKSSSRL-PTPTHLSVFGKPRTTFIQKGIIRCDAQNSDQTIASTTTLSA---LELLKTSA 371
Query: 72 ADRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNHIEEAAVLSTCNR 131
ADRYTKE+SSI+VIGL+VHTAPVEMREKLAIPEA+WP+ I ELC+LNHIEEAAVLSTCNR
Sbjct: 372 ADRYTKERSSIVVIGLSVHTAPVEMREKLAIPEAEWPRAIAELCSLNHIEEAAVLSTCNR 551
Query: 132 IEIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQHLFEVAAGLDSLVL 191
+EIY+VALSQHRGV+EV +W+SK S V V E+ +H+ LLYNKDATQHLF+V+AGL+SLVL
Sbjct: 552 MEIYVVALSQHRGVKEVMEWMSKTSSVPVSELCEHRFLLYNKDATQHLFQVSAGLESLVL 731
Query: 192 GEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISVGKR 230
GEGQIL+QVKQVVK GQGV GF R ISGLFK AI+ K+
Sbjct: 732 GEGQILAQVKQVVKVGQGVNGFGRNISGLFKHAITCRKK 848
Score = 246 bits (627), Expect(2) = e-139
Identities = 127/189 (67%), Positives = 159/189 (83%), Gaps = 1/189 (0%)
Frame = +1
Query: 227 VGKRVRTETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHLVAKG 286
VGKRVR ETNI++G+VSVSSAAVELA MK E+S +A++L+IGAGKMGKLVIKHLV+KG
Sbjct: 838 VGKRVRAETNIAAGAVSVSSAAVELAYMKLPEASHDNARMLIIGAGKMGKLVIKHLVSKG 1017
Query: 287 CRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENV 346
C +MVVVNR+EE+V AIR+ELKD++I+Y+PLSEM+ C EADV+FT TASE+ LF K++V
Sbjct: 1018 CTRMVVVNRTEERVAAIREELKDIEIIYKPLSEMLSCVCEADVVFTSTASETPLFMKDDV 1197
Query: 347 EILPPVGQGV-RRRLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQRKAME 405
+ LPP Q V RLF+DIS+PRNV VS++E+ VYNVDDL+EVV ANKEDR +KAME
Sbjct: 1198 KDLPPASQDVGGLRLFIDISVPRNVGSCVSDIESVRVYNVDDLKEVVAANKEDRLKKAME 1377
Query: 406 ARGIIKEEL 414
A+ II E+L
Sbjct: 1378 AQAIIGEKL 1404
>TC91811 similar to GP|13676394|dbj|BAB41186. glutamyl-tRNA reductase
{Amaranthus tricolor}, partial (31%)
Length = 246
Score = 162 bits (409), Expect = 4e-40
Identities = 81/82 (98%), Positives = 82/82 (99%)
Frame = +1
Query: 370 VDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPT 429
VDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPT
Sbjct: 1 VDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPT 180
Query: 430 IKKFRAYVERIRASELEKCLSK 451
IKKFRAYVERI+ASELEKCLSK
Sbjct: 181 IKKFRAYVERIKASELEKCLSK 246
>TC91922 similar to SP|P93111|HMA1_CUCSA Glutamyl-tRNA reductase 1
chloroplast precursor (EC 1.2.1.-) (GluTR). [Cucumber]
{Cucumis sativus}, partial (9%)
Length = 513
Score = 70.9 bits (172), Expect = 1e-12
Identities = 32/52 (61%), Positives = 41/52 (78%)
Frame = +2
Query: 478 GPMQHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEKAKK 529
GPMQHLRCDG+D + L E LE M ALN M+ LETEIS++E+K+R K+E +K
Sbjct: 2 GPMQHLRCDGSDDRTLTETLEYMHALNSMFSLETEISVLEQKVRAKVELNQK 157
>AL366539 homologue to SP|P93111|HMA1_ Glutamyl-tRNA reductase 1 chloroplast
precursor (EC 1.2.1.-) (GluTR). [Cucumber] {Cucumis
sativus}, partial (7%)
Length = 476
Score = 60.8 bits (146), Expect = 1e-09
Identities = 28/32 (87%), Positives = 32/32 (99%)
Frame = -1
Query: 73 DRYTKEKSSIIVIGLNVHTAPVEMREKLAIPE 104
+RYTKE+SSI+VIGL+VHTAPVEMREKLAIPE
Sbjct: 98 ERYTKERSSIVVIGLSVHTAPVEMREKLAIPE 3
>BQ153361 GP|146330|gb|A delta-aminolevulinate synthase (EC 2.3.1.37)
{Escherichia coli}, partial (12%)
Length = 161
Score = 37.7 bits (86), Expect = 0.011
Identities = 18/37 (48%), Positives = 28/37 (75%)
Frame = +1
Query: 216 KISGLFKQAISVGKRVRTETNISSGSVSVSSAAVELA 252
++ +F+++ SV KRVRTET+I + +VSV+ AA LA
Sbjct: 40 ELERMFQKSFSVAKRVRTETDIGASAVSVAFAACTLA 150
>TC80090 similar to GP|15215794|gb|AAK91442.1 AT5g19180/T24G5_80
{Arabidopsis thaliana}, partial (59%)
Length = 903
Score = 33.5 bits (75), Expect = 0.21
Identities = 21/58 (36%), Positives = 34/58 (58%), Gaps = 1/58 (1%)
Frame = +1
Query: 264 AKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSE-EKVNAIRKELKDVDIVYRPLSEM 320
AKVLV+GAG +G ++K L G R + V++ E N R+ L ++ V +P +E+
Sbjct: 184 AKVLVVGAGGLGCELLKDLALSGFRNLDVIDMDRIEVTNLNRQFLFRLEDVGKPKAEV 357
>TC84181 similar to SP|Q9SJA7|SOX_ARATH Potential sarcosine oxidase (EC
1.5.3.1). [Mouse-ear cress] {Arabidopsis thaliana},
partial (20%)
Length = 761
Score = 31.2 bits (69), Expect = 1.0
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = +3
Query: 255 KRLESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKMVV 292
K++ESS D V+++GAG MG + +G + +V+
Sbjct: 27 KKMESSGSDYDVIIVGAGVMGSSTAYYAAKRGLKTLVL 140
>TC80148 weakly similar to SP|Q39172|P1_ARATH Probable NADP-dependent
oxidoreductase P1 (EC 1.-.-.-). [Mouse-ear cress]
{Arabidopsis thaliana}, partial (33%)
Length = 1339
Score = 31.2 bits (69), Expect = 1.0
Identities = 28/117 (23%), Positives = 50/117 (41%), Gaps = 2/117 (1%)
Frame = +2
Query: 117 LNHIEEAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVS--VPEISKHQILLYNKD 174
LNH+ + A + C I Y ++ GVR + + + K+ + + E H+ + KD
Sbjct: 887 LNHVNKHARIPPCGMISQYNKVWTEREGVRNLLNMVGKQVRMEGFMLESYWHRFGDFAKD 1066
Query: 175 ATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISVGKRV 231
++L E +VK K G+ GF ++ LF + ++GK V
Sbjct: 1067MEKYLQE----------------GKVKSKNKINIGIEGFLESLNSLFSSS-NIGKVV 1186
>TC82251
Length = 597
Score = 30.8 bits (68), Expect = 1.3
Identities = 28/99 (28%), Positives = 48/99 (48%), Gaps = 9/99 (9%)
Frame = -1
Query: 70 SSADRYTKEKSSII---VIGLNVHTAPVEMREKLAIPEAQWPQVIQEL-----CALNHIE 121
SSA R + ++ ++ GLN + P + + A P +PQV +E+ L++ E
Sbjct: 546 SSAQRPSDDRQALADGYSTGLNESSNPFNLAQDGAQPREDYPQVSEEVGRAPSSPLSYAE 367
Query: 122 -EAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVS 159
+ ST R EIY L ++ V +++ S+ GVS
Sbjct: 366 SHRTIPSTAMRPEIYRQTLFPYQAVEQISG--SRPFGVS 256
>CB892424 similar to PIR|T03964|T0 probable ubiquitin--protein ligase (EC
6.3.2.19) - common tobacco, partial (13%)
Length = 425
Score = 30.8 bits (68), Expect = 1.3
Identities = 13/32 (40%), Positives = 21/32 (65%)
Frame = +1
Query: 255 KRLESSFGDAKVLVIGAGKMGKLVIKHLVAKG 286
++L+ F DAKV V+G+G +G +K+L G
Sbjct: 265 QKLQKKFEDAKVFVVGSGALGCEFLKNLALMG 360
>TC93068 similar to GP|3170523|gb|AAC18088.1| coatomer alpha subunit
{Aspergillus nidulans}, partial (8%)
Length = 700
Score = 30.0 bits (66), Expect = 2.3
Identities = 16/60 (26%), Positives = 26/60 (42%)
Frame = +3
Query: 282 LVAKGCRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLF 341
++A G + N + K R ++I + +E CAA I++GTA E F
Sbjct: 213 IIANGGSSKLTENAKKAKAQCERNPSDAIEIEFDQFAEFDVCAASHSPIYSGTAYEECAF 392
>BQ140754 similar to GP|22535581|dbj contains EST AU182189(C63973)~similar to
Arabidopsis thaliana chromosome5 At5g65910~unknown,
partial (4%)
Length = 460
Score = 29.6 bits (65), Expect = 3.0
Identities = 32/112 (28%), Positives = 49/112 (43%), Gaps = 1/112 (0%)
Frame = +3
Query: 44 CTLRSDNPLPQNFIVSKPSPLEILKTSSADRYTKEKSSIIVIGLNVHTAPVEMREKLAIP 103
CTL S +P + + K LE T + DR +KEK + + N+ + E + L++P
Sbjct: 21 CTLNSAKMMPLFYQLRK*WRLEQPLTQAIDRRSKEKKEVDL--SNIPSKVEEQEQHLSVP 194
Query: 104 -EAQWPQVIQELCALNHIEEAAVLSTCNRIEIYLVALSQHRGVREVTDWISK 154
AQ V + A +EEA + N V +H +VT I K
Sbjct: 195 SNAQLESVPLQTSA---VEEAPPMVVSN------VKTEEHSVKSDVTQPIEK 323
>TC91668 similar to GP|11121445|emb|CAC14866. DNA Helicase {Arabidopsis
thaliana}, partial (26%)
Length = 708
Score = 29.6 bits (65), Expect = 3.0
Identities = 12/33 (36%), Positives = 22/33 (66%)
Frame = +2
Query: 497 LENMRALNRMYDLETEISLMEEKIRVKMEKAKK 529
+EN L ++++E EI ++EKIR +EK ++
Sbjct: 77 MENYEVLEELHNVEAEIEDVQEKIRALIEKQER 175
>TC79403 homologue to GP|10190525|emb|CAC09322. unnamed protein product
{Pisum sativum}, partial (42%)
Length = 1027
Score = 29.3 bits (64), Expect = 3.9
Identities = 22/77 (28%), Positives = 40/77 (51%), Gaps = 11/77 (14%)
Frame = +3
Query: 148 VTDWISKKSGVSVPEISKHQILL-------YNKDATQHLFEVAA--GLDSLVLGEGQILS 198
+ +WI PE K+ +L+ +N++ATQ + ++AA G+ +++G+ ILS
Sbjct: 435 LANWIQALFNSLPPEDYKNGVLVLGGDGRYFNREATQIIVKIAAGNGVGKILVGKEGILS 614
Query: 199 --QVKQVVKSGQGVPGF 213
V V++ Q GF
Sbjct: 615 TPAVSAVIRKRQANGGF 665
>TC78175 similar to GP|1527001|gb|AAB07724.1| Ipomoea nil Pn47p, partial
(32%)
Length = 1153
Score = 29.3 bits (64), Expect = 3.9
Identities = 16/57 (28%), Positives = 24/57 (42%)
Frame = -1
Query: 19 FSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSPLEILKTSSADRY 75
F P Q P+ + Q+L ++P +P P N + PL+ILK Y
Sbjct: 541 FEPTQHPELFLSKQQLCPSSPMLP*LHQWQQQRTPSNLFPNSKHPLKILKLRETLHY 371
>TC88580 similar to GP|9758308|dbj|BAB08782.1 gene_id:MUA2.4~unknown protein
{Arabidopsis thaliana}, partial (51%)
Length = 1388
Score = 29.3 bits (64), Expect = 3.9
Identities = 17/47 (36%), Positives = 29/47 (61%)
Frame = -3
Query: 182 VAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISVG 228
V G+D +VLG G+ ++V + KS +G G++ ++GL K S+G
Sbjct: 459 VFGGVDFVVLGCGECAARVTESGKSAEGA*GWE-PLAGLKKDIGSIG 322
>TC91007
Length = 757
Score = 28.9 bits (63), Expect = 5.1
Identities = 24/83 (28%), Positives = 39/83 (46%), Gaps = 2/83 (2%)
Frame = +1
Query: 335 ASESLLFSKENVEILPPVGQGVRRRLFVDISIPRNVDPGVSELENALVYNVD--DLREVV 392
AS S S ++V L Q + L +D+ R PG+S+ VYNV+ DL +
Sbjct: 361 ASSSYFSSPQSVAQLTE-NQYMDEELDIDLEYLRMTFPGISDESLVDVYNVNSGDLEAAI 537
Query: 393 DANKEDRQRKAMEARGIIKEELN 415
+ + A++A G + E L+
Sbjct: 538 EMLSQLEFDDAVDAHGSLPESLD 606
>TC80761 homologue to GP|1370174|emb|CAA98164.1 RAB1Y {Lotus japonicus},
partial (94%)
Length = 994
Score = 28.9 bits (63), Expect = 5.1
Identities = 17/77 (22%), Positives = 37/77 (47%)
Frame = +1
Query: 287 CRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENV 346
C KM+V N+ +++ + + + + +D +EC+A+ V E +L +
Sbjct: 469 CMKMLVGNKVDKEGDRVVTKKEGIDFAREYGCLFIECSAKTRVNVQQCFEELVLKVLDTP 648
Query: 347 EILPPVGQGVRRRLFVD 363
+L +GV++ +F D
Sbjct: 649 SLLAEGSKGVKKNIFKD 699
>TC77092 squalene monooxygenase 2 [Medicago truncatula]
Length = 1910
Score = 28.5 bits (62), Expect = 6.7
Identities = 15/46 (32%), Positives = 26/46 (55%)
Frame = +3
Query: 262 GDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKVNAIRKEL 307
GDA V+++GAG G + H + K R++ ++ R + + I EL
Sbjct: 297 GDADVIIVGAGIAG-AALAHTLGKDGRRVHIIERDLSEPDRIVGEL 431
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,738,950
Number of Sequences: 36976
Number of extensions: 155385
Number of successful extensions: 755
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 743
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 751
length of query: 529
length of database: 9,014,727
effective HSP length: 101
effective length of query: 428
effective length of database: 5,280,151
effective search space: 2259904628
effective search space used: 2259904628
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124214.3


