

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.11 + phase: 0 /pseudo
(240 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
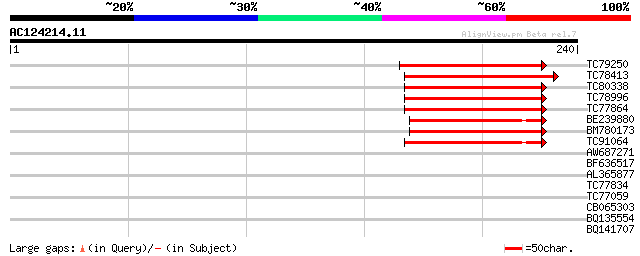
TC79250 similar to PIR|T06329|T06329 symbiotic ammonium transpor... 74 5e-14
TC78413 similar to PIR|T06329|T06329 symbiotic ammonium transpor... 68 3e-12
TC80338 weakly similar to GP|20127018|gb|AAM10936.1 putative bHL... 60 6e-10
TC78996 similar to GP|20127022|gb|AAM10937.1 putative bHLH trans... 58 4e-09
TC77864 weakly similar to PIR|T06329|T06329 symbiotic ammonium t... 58 4e-09
BE239880 weakly similar to PIR|T06329|T063 symbiotic ammonium tr... 54 7e-08
BM780173 similar to PIR|T06329|T063 symbiotic ammonium transport... 50 8e-07
TC91064 weakly similar to GP|20127014|gb|AAM10934.1 putative bHL... 48 4e-06
AW687271 weakly similar to PIR|T06329|T063 symbiotic ammonium tr... 33 0.098
BF636517 SP|Q06366|BXE Botulinum neurotoxin type E nontoxic com... 32 0.22
AL365877 29 1.4
TC77834 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 28 4.1
TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 28 4.1
CB065303 weakly similar to PIR|E83162|E83 respiratory nitrate re... 27 7.0
BQ135554 similar to GP|15487981|gb| Cf50 {Dictyostelium discoide... 27 9.2
BQ141707 homologue to PIR|T23451|T234 hypothetical protein K08E3... 27 9.2
>TC79250 similar to PIR|T06329|T06329 symbiotic ammonium transport protein
SAT1 - soybean, partial (34%)
Length = 1145
Score = 73.9 bits (180), Expect = 5e-14
Identities = 36/62 (58%), Positives = 47/62 (75%)
Frame = +2
Query: 166 KRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITII 225
+ LPE+EAR ++NVLIRVHCEK KG + I +IE L+L VT+SS + FG+ LDITII
Sbjct: 818 RSLPEVEARVSEKNVLIRVHCEKHKGALMNIIQEIENLHLSVTSSSALLFGTTKLDITII 997
Query: 226 AQ 227
A+
Sbjct: 998 AE 1003
>TC78413 similar to PIR|T06329|T06329 symbiotic ammonium transport protein
SAT1 - soybean, partial (30%)
Length = 1397
Score = 68.2 bits (165), Expect = 3e-12
Identities = 31/65 (47%), Positives = 48/65 (73%)
Frame = +2
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LP +EA+ D++VLIR+HC+K KG++ K + +I+KL+L V N+S + FG LDITI+AQ
Sbjct: 806 LPHVEAKVLDKDVLIRIHCQKQKGLLLKILVEIQKLHLFVVNNSVLPFGDSILDITIVAQ 985
Query: 228 PTLSF 232
+ +
Sbjct: 986 MGIGY 1000
>TC80338 weakly similar to GP|20127018|gb|AAM10936.1 putative bHLH
transcription factor {Arabidopsis thaliana}, partial
(15%)
Length = 1128
Score = 60.5 bits (145), Expect = 6e-10
Identities = 28/60 (46%), Positives = 41/60 (67%)
Frame = +1
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPE++A+ +VL+ VHCEK G++ K + +E L+L V NSS + FG LDITI+A+
Sbjct: 538 LPEVKAKVLQNDVLVIVHCEKQNGILLKILTCLENLHLSVVNSSVLNFGKSILDITIVAK 717
>TC78996 similar to GP|20127022|gb|AAM10937.1 putative bHLH transcription
factor {Arabidopsis thaliana}, partial (27%)
Length = 1081
Score = 57.8 bits (138), Expect = 4e-09
Identities = 26/60 (43%), Positives = 41/60 (68%)
Frame = +3
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE R ++VLI++ C+K G + ++E LNL V +S+F+ FG+ +D+TI+AQ
Sbjct: 333 LPEIEVRVSSKDVLIKIQCDKHSGRAATVLGQLENLNLTVHSSTFLPFGNNIVDVTIVAQ 512
>TC77864 weakly similar to PIR|T06329|T06329 symbiotic ammonium transport
protein SAT1 - soybean, partial (21%)
Length = 1411
Score = 57.8 bits (138), Expect = 4e-09
Identities = 29/60 (48%), Positives = 39/60 (64%)
Frame = +3
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
L E+EAR ++ VLIR+HCEK K +V K +EK N+ +T+SS + FG L I I AQ
Sbjct: 807 LLEVEARVKEKEVLIRIHCEKQKDIVLKIHELLEKFNITITSSSMLPFGDSILVINICAQ 986
>BE239880 weakly similar to PIR|T06329|T063 symbiotic ammonium transport
protein SAT1 - soybean, partial (10%)
Length = 659
Score = 53.5 bits (127), Expect = 7e-08
Identities = 25/58 (43%), Positives = 40/58 (68%)
Frame = +2
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
+++AR + +LIR+HCEK KG++ K + +I+ L NSS + FG ++DITIIA+
Sbjct: 215 QVQARVSGKEMLIRIHCEKHKGILVKVMAEIQSFQLFAVNSSVLPFGD-SIDITIIAE 385
>BM780173 similar to PIR|T06329|T063 symbiotic ammonium transport protein
SAT1 - soybean, partial (14%)
Length = 732
Score = 50.1 bits (118), Expect = 8e-07
Identities = 24/58 (41%), Positives = 37/58 (63%)
Frame = -1
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
+++AR + VLI++HCEK + K + +E L+L VT SS + FG+ L TI+AQ
Sbjct: 393 DVKARIMENEVLIQMHCEKENDIEIKIYNVLENLDLFVTASSVLAFGTSTLGFTIVAQ 220
>TC91064 weakly similar to GP|20127014|gb|AAM10934.1 putative bHLH
transcription factor {Arabidopsis thaliana}, partial
(19%)
Length = 751
Score = 47.8 bits (112), Expect = 4e-06
Identities = 23/60 (38%), Positives = 42/60 (69%)
Frame = +2
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
L E+EAR ++ VLI+++C +G+V + +++ L+L +T+ + + FG+ LDITIIA+
Sbjct: 464 LLEVEARILEKEVLIKIYCGMQEGIVVNIMSQLQLLHLSITSINVLPFGN-TLDITIIAK 640
>AW687271 weakly similar to PIR|T06329|T063 symbiotic ammonium transport
protein SAT1 - soybean, partial (10%)
Length = 469
Score = 33.1 bits (74), Expect = 0.098
Identities = 15/31 (48%), Positives = 24/31 (77%)
Frame = +1
Query: 197 IHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
++++EK +L V +SS + FG+ LDITI+AQ
Sbjct: 7 LNELEKHHLSVQSSSILPFGNNYLDITIVAQ 99
>BF636517 SP|Q06366|BXE Botulinum neurotoxin type E nontoxic component.
{Clostridium butyricum}, partial (1%)
Length = 456
Score = 32.0 bits (71), Expect = 0.22
Identities = 17/47 (36%), Positives = 26/47 (55%)
Frame = +1
Query: 192 VVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQPTLSFYVFHLR 238
V+ K + KI+K L + ++ ++FG LDITII FY + R
Sbjct: 52 VLVKIMVKIQKFKLFIFKNNVLSFGD*ILDITIIITQVYVFYFWSCR 192
>AL365877
Length = 526
Score = 29.3 bits (64), Expect = 1.4
Identities = 16/31 (51%), Positives = 23/31 (73%), Gaps = 1/31 (3%)
Frame = +1
Query: 177 DRNVLIRVHCEKIKGVV-EKTIHKIEKLNLK 206
+R+VLIR+HC K +GV+ E K+EKL+ K
Sbjct: 1 ERSVLIRLHCLKSQGVM*ENY*VKLEKLHSK 93
>TC77834 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, complete
Length = 1604
Score = 27.7 bits (60), Expect = 4.1
Identities = 18/50 (36%), Positives = 28/50 (56%)
Frame = -3
Query: 175 FCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITI 224
F DRN+ R+H EK++ +V T KI LK+ S F + +C + I +
Sbjct: 288 FQDRNLAKRIHVEKLRCLVRDTHLKIGS-QLKLDRSIF-SCNACLVSIFV 145
>TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (96%)
Length = 1582
Score = 27.7 bits (60), Expect = 4.1
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 5/44 (11%)
Frame = -1
Query: 175 FCDRNVLIRVHCEKIKGVVEKTIHKIEK-----LNLKVTNSSFM 213
F DRN+ RVH ++++G V T KI N+ +N SF+
Sbjct: 259 FQDRNLSNRVHVDELRGFVRNTHLKIRSQLNLYTNIFSSNQSFV 128
>CB065303 weakly similar to PIR|E83162|E83 respiratory nitrate reductase
delta chain PA3873 [imported] - Pseudomonas aeruginosa
(strain PAO1), partial (41%)
Length = 633
Score = 26.9 bits (58), Expect = 7.0
Identities = 13/18 (72%), Positives = 15/18 (83%), Gaps = 1/18 (5%)
Frame = +2
Query: 158 YLPLFLDY-KRLPEIEAR 174
YLP+FL+Y RLPE EAR
Sbjct: 500 YLPVFLEYLSRLPEKEAR 553
>BQ135554 similar to GP|15487981|gb| Cf50 {Dictyostelium discoideum}, partial
(4%)
Length = 797
Score = 26.6 bits (57), Expect = 9.2
Identities = 21/85 (24%), Positives = 36/85 (41%), Gaps = 14/85 (16%)
Frame = +1
Query: 170 EIEARF-----CDRNVLIR-VHCEKIKGVVEKTIHK--------IEKLNLKVTNSSFMTF 215
EIE RF C +NV ++ +HCE ++ +H+ + K + +
Sbjct: 352 EIEYRFF*WAPCLKNVNVK*IHCEVKVNFIK*IVHRYVDTL*SRVHVWRNKYVECNIINM 531
Query: 216 GSCALDITIIAQPTLSFYVFHLRTF 240
D+ QP++S+YV H F
Sbjct: 532 SGTYTDMN---QPSMSYYVLHYLLF 597
>BQ141707 homologue to PIR|T23451|T234 hypothetical protein K08E3.2 -
Caenorhabditis elegans, partial (3%)
Length = 1181
Score = 26.6 bits (57), Expect = 9.2
Identities = 11/19 (57%), Positives = 16/19 (83%)
Frame = -1
Query: 150 SSASASLLYLPLFLDYKRL 168
SS+ ++++YLPLFL Y RL
Sbjct: 713 SSSISAVIYLPLFLPYLRL 657
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.364 0.163 0.551
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,732,804
Number of Sequences: 36976
Number of extensions: 110393
Number of successful extensions: 1559
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 1551
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1558
length of query: 240
length of database: 9,014,727
effective HSP length: 93
effective length of query: 147
effective length of database: 5,575,959
effective search space: 819665973
effective search space used: 819665973
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124214.11


