

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123898.1 - phase: 0 /pseudo
(177 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
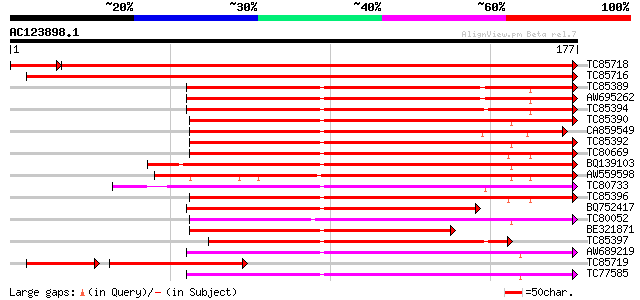
Score E
Sequences producing significant alignments: (bits) Value
TC85718 homologue to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa prot... 322 4e-92
TC85716 homologue to PIR|S19140|S19140 dnaK-type molecular chape... 291 7e-80
TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protei... 112 1e-25
AW695262 homologue to GP|170386|gb|A glucose-regulated protein 7... 112 1e-25
TC85394 homologue to GP|2642238|gb|AAB86942.1| endoplasmic retic... 111 2e-25
TC85390 homologue to SP|P27322|HS72_LYCES Heat shock cognate 70 ... 110 2e-25
CA859549 similar to GP|20260807|gb Hsp70 protein 1 {Rhizopus sto... 109 6e-25
TC85392 homologue to PIR|JC4786|JC4786 dnaK-type molecular chape... 108 8e-25
TC80669 homologue to PIR|S53498|S53498 dnaK-type molecular chape... 108 1e-24
BQ139103 homologue to GP|2655420|gb| heat shock cognate protein ... 108 1e-24
AW559598 weakly similar to SP|P09189|HS7C Heat shock cognate 70 ... 107 2e-24
TC80733 homologue to SP|Q01233|HS70_NEUCR Heat shock 70 kDa prot... 107 2e-24
TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protei... 105 1e-23
BQ752417 similar to GP|5597020|dbj ER chaperone BiP {Aspergillus... 97 3e-21
TC80052 similar to PIR|JC4786|JC4786 dnaK-type molecular chapero... 94 3e-20
BE321871 GP|20563125|db heat shock cognate protein {Bombyx mori}... 89 1e-18
TC85397 homologue to GP|170386|gb|AAA99920.1|| glucose-regulated... 87 3e-18
AW689219 similar to PIR|D96830|D96 probable heat-shock protein ... 86 6e-18
TC85719 similar to PIR|T49940|T49940 hypothetical protein F17I14... 73 2e-17
TC77585 similar to GP|17473863|gb|AAL38353.1 putative heat-shock... 85 2e-17
>TC85718 homologue to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa protein
mitochondrial precursor. [Kidney bean French bean]
{Phaseolus vulgaris}, complete
Length = 2371
Score = 322 bits (826), Expect(2) = 4e-92
Identities = 161/161 (100%), Positives = 161/161 (100%)
Frame = +2
Query: 17 ASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNP 76
ASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNP
Sbjct: 143 ASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNP 322
Query: 77 KVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQK 136
KVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQK
Sbjct: 323 KVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQK 502
Query: 137 EMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
EMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE
Sbjct: 503 EMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 625
Score = 32.3 bits (72), Expect(2) = 4e-92
Identities = 16/16 (100%), Positives = 16/16 (100%)
Frame = +1
Query: 1 MAAATALLRSLRRRDA 16
MAAATALLRSLRRRDA
Sbjct: 94 MAAATALLRSLRRRDA 141
>TC85716 homologue to PIR|S19140|S19140 dnaK-type molecular chaperone PHSP1
precursor mitochondrial - garden pea, partial (98%)
Length = 2422
Score = 291 bits (746), Expect = 7e-80
Identities = 144/172 (83%), Positives = 155/172 (89%)
Frame = +1
Query: 6 ALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGIDLGTTN 65
A+LR RRD ASS+ +AFRSLTG+TKP+Y W+SL+RPFSSR AG+DVIGIDLGTTN
Sbjct: 94 AVLRRALRRDLASSSASAFRSLTGSTKPSYAAQKWASLARPFSSRPAGSDVIGIDLGTTN 273
Query: 66 SCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLI 125
SCVS+MEGKNPKV+ENSEGARTTPSVVAF QKGELLVGTPAKRQAVTNP NT+ G KRLI
Sbjct: 274 SCVSLMEGKNPKVIENSEGARTTPSVVAFNQKGELLVGTPAKRQAVTNPTNTLFGTKRLI 453
Query: 126 GRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
GRRFDDPQTQKEMKMVPYKIVKAPNGDAWVE QQYSPSQIGAFVLTKMKE
Sbjct: 454 GRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEINKQQYSPSQIGAFVLTKMKE 609
>TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protein 4
precursor (BiP 4) (78 kDa glucose-regulated protein
homolog 4) (GRP 78-4)., partial (94%)
Length = 2265
Score = 112 bits (279), Expect = 1e-25
Identities = 60/127 (47%), Positives = 83/127 (65%), Gaps = 5/127 (3%)
Frame = +2
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + + +++ N +G R TPS V+FT E L+G AK A NPE
Sbjct: 161 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSWVSFTDD-ERLIGEAAKNLAAVNPE 337
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQ-----YSPSQIGAF 170
TI KRLIGR+F D + Q++MK+VPYKIV +G +++ + + +SP ++ A
Sbjct: 338 RTIFDVKRLIGRKFADKEVQRDMKLVPYKIVN-KDGKPYIQVRVKDGETKVFSPEEVSAM 514
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 515 ILTKMKE 535
>AW695262 homologue to GP|170386|gb|A glucose-regulated protein 78
{Lycopersicon esculentum}, partial (39%)
Length = 607
Score = 112 bits (279), Expect = 1e-25
Identities = 60/127 (47%), Positives = 83/127 (65%), Gaps = 5/127 (3%)
Frame = +2
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + + +++ N +G R TPS V+FT E L+G AK A NPE
Sbjct: 134 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSWVSFTDD-ERLIGEAAKNLAAVNPE 310
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQ-----YSPSQIGAF 170
TI KRLIGR+F D + Q++MK+VPYKIV +G +++ + + +SP ++ A
Sbjct: 311 RTIFDVKRLIGRKFADKEVQRDMKLVPYKIVN-KDGKPYIQVRVKDGETKVFSPEEVSAM 487
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 488 ILTKMKE 508
>TC85394 homologue to GP|2642238|gb|AAB86942.1| endoplasmic reticulum
HSC70-cognate binding protein precursor {Glycine max},
partial (38%)
Length = 962
Score = 111 bits (277), Expect = 2e-25
Identities = 60/127 (47%), Positives = 83/127 (65%), Gaps = 5/127 (3%)
Frame = +1
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + + +++ N +G R TPS V+FT E L+G AK A NPE
Sbjct: 295 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSWVSFTDD-ERLIGEAAKNLAAVNPE 471
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQ-----YSPSQIGAF 170
TI KRLIGR+F+D + Q++MK+VPYKIV +G +++ + + +SP +I A
Sbjct: 472 RTIFDVKRLIGRKFEDKEVQRDMKLVPYKIVNR-DGKPYIQVRVKDGETKVFSPEEISAM 648
Query: 171 VLTKMKE 177
+L KMKE
Sbjct: 649 ILGKMKE 669
>TC85390 homologue to SP|P27322|HS72_LYCES Heat shock cognate 70 kDa protein
2. [Tomato] {Lycopersicon esculentum}, complete
Length = 2278
Score = 110 bits (276), Expect = 2e-25
Identities = 59/125 (47%), Positives = 78/125 (62%), Gaps = 4/125 (3%)
Frame = +1
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 163 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPTN 339
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK+ P+K++ P + +A+ +Q+S +I + VL
Sbjct: 340 TVFDAKRLIGRRFGDVSVQSDMKLWPFKVIPGPADKPMIVVNYKAEEKQFSAEEISSMVL 519
Query: 173 TKMKE 177
KMKE
Sbjct: 520 MKMKE 534
>CA859549 similar to GP|20260807|gb Hsp70 protein 1 {Rhizopus stolonifer},
partial (19%)
Length = 527
Score = 109 bits (272), Expect = 6e-25
Identities = 60/121 (49%), Positives = 78/121 (63%), Gaps = 3/121 (2%)
Frame = +3
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
+GIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 168 VGIDLGTTYSCVGVWQNDRVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPYN 344
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIV-KAPNGDAWVEAKGQ--QYSPSQIGAFVLT 173
T+ AKRLIGRRF D + Q +MK P+K++ KA VE KG+ Q++P +I + VLT
Sbjct: 345 TVFDAKRLIGRRFADQEVQSDMKHWPFKVIDKAAKPYIQVEYKGETKQFTPEEISSMVLT 524
Query: 174 K 174
K
Sbjct: 525 K 527
>TC85392 homologue to PIR|JC4786|JC4786 dnaK-type molecular chaperone
hsc70-3 - tomato, complete
Length = 2295
Score = 108 bits (271), Expect = 8e-25
Identities = 58/125 (46%), Positives = 77/125 (61%), Gaps = 4/125 (3%)
Frame = +1
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 157 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPTN 333
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
T+ AKRLIGRR D Q +MK+ P+K++ P + +A+ +Q+S +I + VL
Sbjct: 334 TVFDAKRLIGRRISDVSVQSDMKLWPFKVIPGPADKPMIVVNYKAEEKQFSAEEISSMVL 513
Query: 173 TKMKE 177
KMKE
Sbjct: 514 MKMKE 528
>TC80669 homologue to PIR|S53498|S53498 dnaK-type molecular chaperone
HSP71.2 - garden pea, partial (44%)
Length = 955
Score = 108 bits (270), Expect = 1e-24
Identities = 59/125 (47%), Positives = 78/125 (62%), Gaps = 4/125 (3%)
Frame = +2
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP+N
Sbjct: 122 IGIDLGTTYSCVGVWQNDRVEIIPNDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPQN 298
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAKGQQ--YSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK+ P+K+V P V KG++ ++ +I + VL
Sbjct: 299 TVFDAKRLIGRRFSDESVQNDMKLWPFKVVPGPAEKPMIVVNYKGEEKKFAAEEISSMVL 478
Query: 173 TKMKE 177
KM+E
Sbjct: 479 IKMRE 493
>BQ139103 homologue to GP|2655420|gb| heat shock cognate protein HSC70
{Brassica napus}, partial (29%)
Length = 628
Score = 108 bits (270), Expect = 1e-24
Identities = 58/138 (42%), Positives = 83/138 (60%), Gaps = 4/138 (2%)
Frame = +2
Query: 44 SRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVG 103
++P + + G IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G
Sbjct: 47 NKPMAGKGEG-PAIGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDS-ERLIG 220
Query: 104 TPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKG 159
AK Q NP NT+ AKRLIGRRF D Q +MK+ P+KI+ P + + +
Sbjct: 221 DAAKNQVAMNPINTVFDAKRLIGRRFSDASVQSDMKLWPFKIISGPAEKPLIGVNYKGED 400
Query: 160 QQYSPSQIGAFVLTKMKE 177
++++ +I + VL KM+E
Sbjct: 401 KEFAAEEISSMVLMKMRE 454
>AW559598 weakly similar to SP|P09189|HS7C Heat shock cognate 70 kDa protein.
[Petunia] {Petunia hybrida}, partial (24%)
Length = 532
Score = 107 bits (268), Expect = 2e-24
Identities = 65/141 (46%), Positives = 88/141 (62%), Gaps = 9/141 (6%)
Frame = +1
Query: 46 PFSSRAAGND---VIGIDLGTTNSCVSV-MEGKNP-KVVENSEGARTTPSVVAFTQKGEL 100
P R A N +GIDLGTT SCV+V +E KN +++ N EG++ TPS VAFT G+
Sbjct: 16 PLKDRMANNCECYAVGIDLGTTYSCVAVWLEEKNRVEIIHNDEGSKITPSFVAFTD-GQR 192
Query: 101 LVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV--EAK 158
LVG AK QA NP+NT+ AKRLIGR+F DP QK++ + P+K+V N + + K
Sbjct: 193 LVGAAAKDQAAINPQNTVFDAKRLIGRKFSDPVVQKDILLWPFKVVSGVNDKPMISLKFK 372
Query: 159 GQQ--YSPSQIGAFVLTKMKE 177
GQ+ +I + VLT M+E
Sbjct: 373 GQEKLLCAEEISSIVLTNMRE 435
>TC80733 homologue to SP|Q01233|HS70_NEUCR Heat shock 70 kDa protein
(HSP70). {Neurospora crassa}, partial (54%)
Length = 1258
Score = 107 bits (268), Expect = 2e-24
Identities = 60/148 (40%), Positives = 85/148 (56%), Gaps = 3/148 (2%)
Frame = +2
Query: 33 PAYVGHNWSSLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVV 92
P YV + +S+ + A + IGIDLGTT SCV +++ N +G RTTPS V
Sbjct: 137 PRYVTYTHTSI------KMATSFAIGIDLGTTYSCVGRYANDKIEIIANDQGNRTTPSYV 298
Query: 93 AFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVK---AP 149
AF E L+G AK Q NP NT+ AKRLIGR+F D + Q +MK P+K++ P
Sbjct: 299 AFNDT-ERLIGDAAKNQVAMNPHNTVFDAKRLIGRKFSDSEVQADMKHFPFKVIDKGGKP 475
Query: 150 NGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
N + + + + ++P +I A VL KM+E
Sbjct: 476 NIEVEFKGENKTFTPEEISAMVLVKMRE 559
>TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protein 70
{Cucumis sativus}, complete
Length = 2320
Score = 105 bits (261), Expect = 1e-23
Identities = 57/125 (45%), Positives = 76/125 (60%), Gaps = 4/125 (3%)
Frame = +2
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 134 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDS-ERLIGDAAKNQVAMNPTN 310
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAKGQQ--YSPSQIGAFVL 172
T+ AKRLIGRR D Q +MK+ P+K++ P V KG++ ++ +I + VL
Sbjct: 311 TVFDAKRLIGRRISDASVQSDMKLWPFKVIAGPGEKPMIGVNYKGEEKLFASEEISSMVL 490
Query: 173 TKMKE 177
KM+E
Sbjct: 491 IKMRE 505
>BQ752417 similar to GP|5597020|dbj ER chaperone BiP {Aspergillus oryzae},
partial (18%)
Length = 690
Score = 97.1 bits (240), Expect = 3e-21
Identities = 49/92 (53%), Positives = 64/92 (69%)
Frame = +1
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV VM+G +++ N +G R TPS VAFT++ E LVG AK QA NP
Sbjct: 328 VIGIDLGTTYSCVGVMQGGKVEILVNDQGNRITPSYVAFTEE-ERLVGDAAKNQAAANPY 504
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVK 147
NTI KR+IG++F D Q ++K PYK+++
Sbjct: 505 NTIFDIKRMIGQKFADKAVQSDVKHFPYKVIE 600
>TC80052 similar to PIR|JC4786|JC4786 dnaK-type molecular chaperone hsc70-3
- tomato, partial (27%)
Length = 667
Score = 94.0 bits (232), Expect = 3e-20
Identities = 52/125 (41%), Positives = 75/125 (59%), Gaps = 4/125 (3%)
Frame = +3
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV+V + ++ N G RTTPS VAF + E ++G A A +NP N
Sbjct: 108 IGIDLGTTYSCVAVCKNGEIDIIVNDLGKRTTPSFVAF-KDSERMIGDAAFNIAASNPTN 284
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
TI AKRLIGR+F DP Q ++K+ P+K++ N + + + ++ +I + VL
Sbjct: 285 TIFDAKRLIGRKFSDPIVQSDVKLWPFKVIGDLNDKPMIVVNYNDEEKHFAAEEISSMVL 464
Query: 173 TKMKE 177
KM+E
Sbjct: 465 VKMRE 479
>BE321871 GP|20563125|db heat shock cognate protein {Bombyx mori}, partial
(13%)
Length = 346
Score = 88.6 bits (218), Expect = 1e-18
Identities = 45/83 (54%), Positives = 55/83 (66%)
Frame = +1
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
+GIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 97 VGIDLGTTYSCVGVFQHGKVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPNN 273
Query: 117 TISGAKRLIGRRFDDPQTQKEMK 139
TI AKRLIGR+F+D Q +MK
Sbjct: 274 TIFDAKRLIGRKFEDATVQADMK 342
>TC85397 homologue to GP|170386|gb|AAA99920.1|| glucose-regulated protein 78
{Lycopersicon esculentum}, partial (57%)
Length = 648
Score = 87.0 bits (214), Expect = 3e-18
Identities = 45/95 (47%), Positives = 63/95 (65%)
Frame = +1
Query: 63 TTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAK 122
TT SCV V + + +++ N +G R TPS V+FT E L+G AK A NPE TI K
Sbjct: 1 TTYSCVGVYKNGHVEIIANDQGNRITPSWVSFTDD-ERLIGEAAKNLAAVNPERTIFDVK 177
Query: 123 RLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEA 157
RLIGR+F+D + Q++MK+VPYKIV +G +++A
Sbjct: 178 RLIGRKFEDKEVQRDMKLVPYKIVNR-DGKPYIQA 279
>AW689219 similar to PIR|D96830|D96 probable heat-shock protein 41956-44878
[imported] - Arabidopsis thaliana, partial (20%)
Length = 534
Score = 86.3 bits (212), Expect = 6e-18
Identities = 43/126 (34%), Positives = 74/126 (58%), Gaps = 4/126 (3%)
Frame = +3
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
V+G D G + V+V + VV N E R TP++V F +K + +GT + NP+
Sbjct: 84 VVGFDFGNESCIVAVARQRGIDVVLNDESKRETPAIVCFGEK-QRFIGTAGAASTMMNPK 260
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK----GQQYSPSQIGAFV 171
N+IS KRLIG+ + DP Q+++K +P+ + + P+G ++A+ + ++P+Q+ A +
Sbjct: 261 NSISQIKRLIGKLYSDPDLQRDLKSLPFSVAEGPDGYPLIQARYLGEVKMFTPTQVFAMM 440
Query: 172 LTKMKE 177
L MKE
Sbjct: 441 LGNMKE 458
>TC85719 similar to PIR|T49940|T49940 hypothetical protein F17I14.230 -
Arabidopsis thaliana, partial (62%)
Length = 1970
Score = 73.2 bits (178), Expect(2) = 2e-17
Identities = 33/43 (76%), Positives = 37/43 (85%)
Frame = +3
Query: 32 KPAYVGHNWSSLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGK 74
KP+Y W+SL RPFSSR AG+DVIGIDLGTTNSCVS+MEGK
Sbjct: 105 KPSYAAQKWASLVRPFSSRPAGSDVIGIDLGTTNSCVSLMEGK 233
Score = 32.0 bits (71), Expect(2) = 2e-17
Identities = 16/23 (69%), Positives = 19/23 (82%)
Frame = +2
Query: 6 ALLRSLRRRDAASSTVTAFRSLT 28
A+LR RRD ASS+V+AFRSLT
Sbjct: 35 AVLRRALRRDLASSSVSAFRSLT 103
>TC77585 similar to GP|17473863|gb|AAL38353.1 putative heat-shock protein
{Arabidopsis thaliana}, partial (92%)
Length = 3027
Score = 84.7 bits (208), Expect = 2e-17
Identities = 41/126 (32%), Positives = 74/126 (58%), Gaps = 4/126 (3%)
Frame = +1
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
V+G D G + V+V + VV N E R TP++V F K + +GT + NP+
Sbjct: 244 VVGFDFGNESCIVAVARQRGIDVVLNDESKRETPAIVCFGDK-QRFIGTAGAASTMMNPK 420
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK----GQQYSPSQIGAFV 171
N+IS KRLIG++F DP+ Q+++K +P+ + + P+G + A+ ++++ +Q+ +
Sbjct: 421 NSISQIKRLIGKKFADPELQRDLKSLPFNVTEGPDGYPLIHARYLGESREFTATQVFGMM 600
Query: 172 LTKMKE 177
L+ +KE
Sbjct: 601 LSNLKE 618
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.127 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,265,678
Number of Sequences: 36976
Number of extensions: 48723
Number of successful extensions: 230
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 202
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 203
length of query: 177
length of database: 9,014,727
effective HSP length: 90
effective length of query: 87
effective length of database: 5,686,887
effective search space: 494759169
effective search space used: 494759169
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC123898.1


