

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
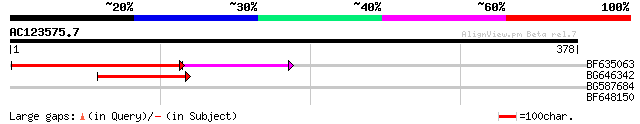
Query= AC123575.7 - phase: 0 /pseudo
(378 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF635063 weakly similar to PIR|F84486|F84 probable retroelement ... 162 2e-49
BG646342 weakly similar to PIR|F84486|F84 probable retroelement ... 81 8e-16
BG587684 similar to GP|12858190|dbj evidence:NAS~putative~unclas... 29 2.6
BF648150 similar to GP|14586969|gb| pol polyprotein {Citrus x pa... 28 4.4
>BF635063 weakly similar to PIR|F84486|F84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(4%)
Length = 677
Score = 162 bits (410), Expect(2) = 2e-49
Identities = 83/115 (72%), Positives = 92/115 (79%)
Frame = -2
Query: 2 MDSKWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAIS 61
M SK +IEKFTG NDFGLWKVKM +LIQQKC +ALKG+ +P ++ AEKTEM DKA S
Sbjct: 568 MGSKRDIEKFTGDNDFGLWKVKMEAVLIQQKCEKALKGEVSLPVTMSRAEKTEMVDKARS 389
Query: 62 AIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESK 116
AIVLCLGDKVLREVA+E SMW KL SLYMTKS HRQ LKQQLY +RMVESK
Sbjct: 388 AIVLCLGDKVLREVAKERTAASMWAKL*SLYMTKSLAHRQFLKQQLYSFRMVESK 224
Score = 51.2 bits (121), Expect(2) = 2e-49
Identities = 33/74 (44%), Positives = 42/74 (56%)
Frame = -1
Query: 116 KTNNEATDGIQ*DY**FGEHRCKYGRRG*SLTPVVCFT*VL*EFQEYKEFWQRGHYHFGG 175
K ++ AT G+Q + ** GE+ G S T VVC T + * Q Y WQR H + G
Sbjct: 227 KGHHGATYGVQQNP**LGEY*GAT*G*GKSYTLVVCSTEIF*IVQRYHALWQRRHCYLGR 48
Query: 176 RSSNFENQRVNQIQ 189
SS FENQ V+Q+Q
Sbjct: 47 SSSGFENQGVDQVQ 6
>BG646342 weakly similar to PIR|F84486|F84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(4%)
Length = 599
Score = 80.9 bits (198), Expect = 8e-16
Identities = 43/62 (69%), Positives = 47/62 (75%)
Frame = +2
Query: 59 AISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKTN 118
A SAIVLCLGDKVLREVA+E SM KL+ LYMTKS HRQ LKQQLY ++MVESK
Sbjct: 2 ARSAIVLCLGDKVLREVAKEPTATSMCAKLEYLYMTKSLAHRQFLKQQLYSFKMVESKAI 181
Query: 119 NE 120
E
Sbjct: 182 TE 187
>BG587684 similar to GP|12858190|dbj evidence:NAS~putative~unclassifiable
{Mus musculus}, partial (14%)
Length = 819
Score = 29.3 bits (64), Expect = 2.6
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = +3
Query: 2 MDSKWNIEKFTGSNDF 17
MD+KW+IE F G NDF
Sbjct: 735 MDTKWDIEIFIGENDF 782
>BF648150 similar to GP|14586969|gb| pol polyprotein {Citrus x paradisi},
partial (3%)
Length = 658
Score = 28.5 bits (62), Expect = 4.4
Identities = 11/34 (32%), Positives = 22/34 (64%)
Frame = +2
Query: 84 MWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKT 117
+W+KL++ YM + T ++ L Y+MV++K+
Sbjct: 275 LWDKLETRYMREDATSKKFLVSHFNNYKMVDNKS 376
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.356 0.157 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,773,358
Number of Sequences: 36976
Number of extensions: 133579
Number of successful extensions: 1235
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 938
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 294
Number of HSP's gapped (non-prelim): 983
length of query: 378
length of database: 9,014,727
effective HSP length: 98
effective length of query: 280
effective length of database: 5,391,079
effective search space: 1509502120
effective search space used: 1509502120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC123575.7


