

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
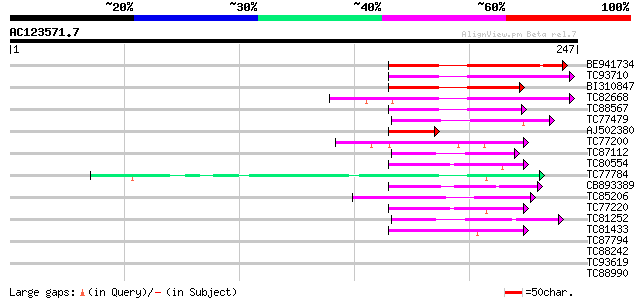
Query= AC123571.7 - phase: 0
(247 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE941734 similar to GP|7406677|emb| putative ripening-related bZ... 64 5e-11
TC93710 similar to PIR|T51273|T51273 promoter-binding factor-lik... 64 7e-11
BI310847 similar to GP|13346155|gb| bZIP protein DPBF3 {Arabidop... 63 9e-11
TC82668 weakly similar to GP|13346157|gb|AAK19602.1 bZIP protein... 55 2e-08
TC88567 similar to SP|P42777|GBF4_ARATH G-box binding factor 4. ... 47 5e-06
TC77479 similar to GP|19423874|gb|AAL87314.1 unknown protein {Ar... 44 4e-05
AJ502380 similar to GP|13346159|gb| bZIP protein DPBF5 {Arabidop... 43 1e-04
TC77200 similar to GP|22597162|gb|AAN03468.1 bZIP transcription ... 43 1e-04
TC87112 similar to GP|2921823|gb|AAC04862.1| shoot-forming PKSF1... 42 2e-04
TC80554 similar to GP|13430400|gb|AAK25822.1 bZip transcription ... 42 2e-04
TC77784 similar to GP|14272369|emb|CAC40022. bZIP transcription ... 42 2e-04
CB893389 similar to PIR|T48034|T480 bZIP transcription factor-li... 42 3e-04
TC85206 similar to GP|16580134|gb|AAK92215.1 bZIP transcription ... 42 3e-04
TC77220 similar to GP|22597162|gb|AAN03468.1 bZIP transcription ... 41 5e-04
TC81252 similar to PIR|T52624|T52624 b-Zip DNA binding protein [... 41 5e-04
TC81433 similar to GP|13430400|gb|AAK25822.1 bZip transcription ... 40 6e-04
TC87794 similar to GP|13775107|gb|AAK39130.1 bZIP transcription ... 39 0.002
TC88242 similar to GP|15100049|gb|AAK84220.1 transcription facto... 38 0.003
TC93619 similar to PIR|T52624|T52624 b-Zip DNA binding protein [... 38 0.004
TC88990 similar to PIR|T11751|T11751 transcription repressor ROM... 38 0.004
>BE941734 similar to GP|7406677|emb| putative ripening-related bZIP protein
{Vitis vinifera}, partial (18%)
Length = 607
Score = 63.9 bits (154), Expect = 5e-11
Identities = 39/78 (50%), Positives = 50/78 (64%)
Frame = +2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELE +V L+EEN L+++Q+E
Sbjct: 8 ERRQRRMIKNRESAARSRARKQ------------AYTMELEAEVAKLKEENEELQKKQEE 151
Query: 226 LWEAESGGQQKKKSSLYR 243
+ E + Q K+ +L R
Sbjct: 152 IMELQK-NQVKEMMNLQR 202
>TC93710 similar to PIR|T51273|T51273 promoter-binding factor-like protein -
Arabidopsis thaliana, partial (30%)
Length = 598
Score = 63.5 bits (153), Expect = 7e-11
Identities = 43/81 (53%), Positives = 46/81 (56%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT ELE KV L+EEN RL+R +
Sbjct: 61 ERRQKRMIKNRESAARSRARKQ------------AYTQELEIKVSHLEEENERLKRLHEI 204
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RTSS
Sbjct: 205 ERVLPSMPPPDPKHQLRRTSS 267
>BI310847 similar to GP|13346155|gb| bZIP protein DPBF3 {Arabidopsis
thaliana}, partial (30%)
Length = 670
Score = 63.2 bits (152), Expect = 9e-11
Identities = 36/59 (61%), Positives = 41/59 (69%)
Frame = +2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
ERR KRMIKNRESAARSRARKQ AYTNELE KV L+EEN LR++++
Sbjct: 305 ERRQKRMIKNRESAARSRARKQ------------AYTNELEIKVSRLEEENEMLRKRKE 445
>TC82668 weakly similar to GP|13346157|gb|AAK19602.1 bZIP protein DPBF4
{Arabidopsis thaliana}, partial (32%)
Length = 973
Score = 55.5 bits (132), Expect = 2e-08
Identities = 43/113 (38%), Positives = 59/113 (52%), Gaps = 6/113 (5%)
Frame = +1
Query: 140 NVGVPASSSFTCFGK----RFGEAPDISPG--ERRNKRMIKNRESAARSRARKQEKITSF 193
N + +SSS + R +APD ERR +R IKNRESAARSRARKQ
Sbjct: 190 NTSLVSSSSISSLSDAKPGRKRDAPDAYEKALERRLRRKIKNRESAARSRARKQ------ 351
Query: 194 LFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAESGGQQKKKSSLYRTSS 246
AY NEL KV LL+++N +L+++++ + + K L R SS
Sbjct: 352 ------AYHNELVTKVTLLEQQNMQLKKEKEFEQGLQPESSPEPKYRLRRISS 492
>TC88567 similar to SP|P42777|GBF4_ARATH G-box binding factor 4. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (32%)
Length = 1250
Score = 47.4 bits (111), Expect = 5e-06
Identities = 28/60 (46%), Positives = 36/60 (59%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
++R +RMIKNRESAARSR RKQ AY ELE L+EEN +L +++ E
Sbjct: 622 QQRQRRMIKNRESAARSRERKQ------------AYQVELESLAVKLEEENDKLMKEKAE 765
>TC77479 similar to GP|19423874|gb|AAL87314.1 unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 2241
Score = 44.3 bits (103), Expect = 4e-05
Identities = 30/75 (40%), Positives = 41/75 (54%), Gaps = 4/75 (5%)
Frame = +3
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQ---- 222
+R KR++ NR+SAARS+ RK + Y +ELEQKVQ LQ E L Q
Sbjct: 1413 KRAKRILANRQSAARSKERKMK------------YISELEQKVQTLQTETTTLSTQFTKL 1556
Query: 223 QQELWEAESGGQQKK 237
Q + EA+S ++ K
Sbjct: 1557 QMDHQEAKSENKEYK 1601
>AJ502380 similar to GP|13346159|gb| bZIP protein DPBF5 {Arabidopsis
thaliana}, partial (13%)
Length = 472
Score = 43.1 bits (100), Expect = 1e-04
Identities = 21/22 (95%), Positives = 21/22 (95%)
Frame = +2
Query: 166 ERRNKRMIKNRESAARSRARKQ 187
ERR KRMIKNRESAARSRARKQ
Sbjct: 395 ERRQKRMIKNRESAARSRARKQ 460
>TC77200 similar to GP|22597162|gb|AAN03468.1 bZIP transcription factor ATB2
{Glycine max}, partial (84%)
Length = 1481
Score = 42.7 bits (99), Expect = 1e-04
Identities = 36/99 (36%), Positives = 47/99 (47%), Gaps = 15/99 (15%)
Frame = +1
Query: 143 VPASSSFTCFGKRF----GEAPDISP--GERRNKRMIKNRESAARSRARKQEKITSFL-- 194
+ ASSS T G G D+ +R+ KRMI NRESA RSR RKQ+ + +
Sbjct: 529 IMASSSGTSSGSSLLQNSGSEEDLQALMDQRKRKRMISNRESARRSRMRKQKHLDDLVSQ 708
Query: 195 FSKFSAYTNEL-------EQKVQLLQEENARLRRQQQEL 226
SK E+ QK ++ EN+ LR Q EL
Sbjct: 709 VSKLRKENQEILTSVNITTQKYLSVEAENSVLRAQMGEL 825
>TC87112 similar to GP|2921823|gb|AAC04862.1| shoot-forming PKSF1 {Paulownia
kawakamii}, partial (30%)
Length = 1379
Score = 42.4 bits (98), Expect = 2e-04
Identities = 25/56 (44%), Positives = 33/56 (58%)
Frame = +1
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQ 222
+R KR++ NR+SAARS+ RK + YT+ELE+KVQ LQ E L Q
Sbjct: 571 KRAKRILANRQSAARSKERK------------TRYTSELERKVQTLQTEATNLSAQ 702
>TC80554 similar to GP|13430400|gb|AAK25822.1 bZip transcription factor
{Phaseolus vulgaris}, partial (52%)
Length = 839
Score = 42.4 bits (98), Expect = 2e-04
Identities = 27/71 (38%), Positives = 40/71 (56%), Gaps = 10/71 (14%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ----------EE 215
ER+++RM+ NRESA RSR RKQ+++ L+S+ NE Q + L +E
Sbjct: 304 ERKHRRMVSNRESARRSRMRKQKQLDE-LWSQVVWLRNENHQLLDKLNNFCETHDKVVQE 480
Query: 216 NARLRRQQQEL 226
N +L+ Q EL
Sbjct: 481 NVQLKEQASEL 513
>TC77784 similar to GP|14272369|emb|CAC40022. bZIP transcription factor
{Arabidopsis thaliana}, partial (72%)
Length = 2052
Score = 42.4 bits (98), Expect = 2e-04
Identities = 53/202 (26%), Positives = 79/202 (38%), Gaps = 4/202 (1%)
Frame = +3
Query: 36 LQDFLTRPLTLDPTKSL---DYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLN 92
L+ F + P+ T ++ + S SS S S +N N P P LS
Sbjct: 537 LEMFPSWPMRFHQTSTVGGGNKSGGESSDSALSSKNEN------PFEPESP-----LSSK 683
Query: 93 SRPDFLYDPLIRHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCF 152
F D HN + + Q L QQQ I ++ S AS + A
Sbjct: 684 KASSFSSD----HNNNMDQQNLQLQQQKMIISNDASAAIRTASSSQNQISAAAKEK---- 839
Query: 153 GKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELE-QKVQL 211
K G D + +R+ +NRE+A +SR RK+ AY +LE +++L
Sbjct: 840 KKGAGSTSDKPLDAKTLRRLAQNREAARKSRLRKK------------AYVQQLESSRLKL 983
Query: 212 LQEENARLRRQQQELWEAESGG 233
Q E R + Q ++ SGG
Sbjct: 984 TQLEQDLQRARSQGMFMDWSGG 1049
>CB893389 similar to PIR|T48034|T480 bZIP transcription factor-like protein -
Arabidopsis thaliana, partial (35%)
Length = 724
Score = 41.6 bits (96), Expect = 3e-04
Identities = 28/67 (41%), Positives = 36/67 (52%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ER+ KRMI NRESA RSR RKQ+ L + N L + + L EN R+R +
Sbjct: 205 ERKRKRMISNRESARRSRERKQK-----LLEDYQDEANRLRNENRRL-SENIRVREEGFN 366
Query: 226 LWEAESG 232
EA +G
Sbjct: 367 ANEAANG 387
>TC85206 similar to GP|16580134|gb|AAK92215.1 bZIP transcription factor
BZI-4 {Nicotiana tabacum}, partial (31%)
Length = 1139
Score = 41.6 bits (96), Expect = 3e-04
Identities = 27/80 (33%), Positives = 39/80 (48%)
Frame = +2
Query: 150 TCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKV 209
+C + D+ ER+ KRM+ NRESA RSR RKQ+++ +L +
Sbjct: 491 SCSASGGSDGMDLQIDERKRKRMLSNRESARRSRLRKQQQV------------EDLTGEA 634
Query: 210 QLLQEENARLRRQQQELWEA 229
L+ EN RL R + EA
Sbjct: 635 GKLKIENDRLARSIKATEEA 694
>TC77220 similar to GP|22597162|gb|AAN03468.1 bZIP transcription factor ATB2
{Glycine max}, partial (65%)
Length = 1341
Score = 40.8 bits (94), Expect = 5e-04
Identities = 28/71 (39%), Positives = 38/71 (53%), Gaps = 10/71 (14%)
Frame = +3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELE----------QKVQLLQEE 215
+R+ KRMI NRESA RSR RKQ+ + L + S NE + Q+ ++ E
Sbjct: 645 QRKRKRMISNRESARRSRMRKQKHLDD-LAVQLSQLRNENQQILTSVNLTTQRFLAVESE 821
Query: 216 NARLRRQQQEL 226
N+ LR Q EL
Sbjct: 822 NSVLRAQLNEL 854
>TC81252 similar to PIR|T52624|T52624 b-Zip DNA binding protein [imported] -
Arabidopsis thaliana, partial (48%)
Length = 1221
Score = 40.8 bits (94), Expect = 5e-04
Identities = 27/75 (36%), Positives = 40/75 (53%)
Frame = +2
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R KR++ NR+SAARS+ RK + Y ELE+KVQ LQ E L Q L
Sbjct: 590 KRAKRILANRQSAARSKERK------------ARYIQELERKVQTLQTEATTL-SAQLTL 730
Query: 227 WEAESGGQQKKKSSL 241
++ ++ G + + L
Sbjct: 731 YQRDTNGLSTENTEL 775
>TC81433 similar to GP|13430400|gb|AAK25822.1 bZip transcription factor
{Phaseolus vulgaris}, partial (68%)
Length = 774
Score = 40.4 bits (93), Expect = 6e-04
Identities = 24/70 (34%), Positives = 36/70 (51%), Gaps = 9/70 (12%)
Frame = +2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYT---------NELEQKVQLLQEEN 216
ER+++RMI NRESA RSR RKQ+ + T N + + + +EN
Sbjct: 272 ERKHRRMISNRESARRSRMRKQKHLDELWSQVMKLRTENHNLVDKLNHVSESHDTVVQEN 451
Query: 217 ARLRRQQQEL 226
ARL+ + +L
Sbjct: 452 ARLKEETFDL 481
>TC87794 similar to GP|13775107|gb|AAK39130.1 bZIP transcription factor 2
{Phaseolus vulgaris}, partial (50%)
Length = 1264
Score = 38.5 bits (88), Expect = 0.002
Identities = 23/56 (41%), Positives = 31/56 (55%)
Frame = +2
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQ 222
+R +R NRESA RSR RKQ A +EL Q+ ++L +ENA LR +
Sbjct: 389 KRQRRKQSNRESARRSRLRKQ------------AECDELAQRAEVLNQENASLRAE 520
>TC88242 similar to GP|15100049|gb|AAK84220.1 transcription factor bZIP29
{Arabidopsis thaliana}, partial (39%)
Length = 1930
Score = 38.1 bits (87), Expect = 0.003
Identities = 23/53 (43%), Positives = 29/53 (54%)
Frame = +2
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARL 219
+R KR++ NR+SAARS+ RK Y +ELE KVQ LQ E L
Sbjct: 1205 KRAKRILANRQSAARSKERKMR------------YISELEHKVQTLQTEATTL 1327
>TC93619 similar to PIR|T52624|T52624 b-Zip DNA binding protein [imported] -
Arabidopsis thaliana, partial (32%)
Length = 709
Score = 37.7 bits (86), Expect = 0.004
Identities = 25/75 (33%), Positives = 39/75 (51%)
Frame = +2
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R KR++ NR+SAARS+ RK + Y ELE+K+ LQ E L Q L
Sbjct: 125 KRAKRILANRQSAARSKERK------------ACYVVELERKIHTLQTEATTL-SAQLNL 265
Query: 227 WEAESGGQQKKKSSL 241
++ ++ G + + L
Sbjct: 266 FQRDTTGLSSENTEL 310
>TC88990 similar to PIR|T11751|T11751 transcription repressor ROM1 - kidney
bean, partial (91%)
Length = 1528
Score = 37.7 bits (86), Expect = 0.004
Identities = 34/128 (26%), Positives = 53/128 (40%)
Frame = +1
Query: 101 PLIRHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAP 160
P+ + + N + + NIG+ + A P + G+++ +
Sbjct: 799 PISQSSVPGNPVVSIPATNLNIGMDLWNASSAGAEAAKMRHNQPGAPGAGALGEQWMQQD 978
Query: 161 DISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLR 220
D +R KR NRESA +SR RKQ A EL ++V+ L EN LR
Sbjct: 979 DREL--KRQKRKQSNRESARKSRLRKQ------------AECEELLKRVEALGGENRTLR 1116
Query: 221 RQQQELWE 228
+ Q+L E
Sbjct: 1117EELQKLSE 1140
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.127 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,010,387
Number of Sequences: 36976
Number of extensions: 120303
Number of successful extensions: 855
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 814
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 840
length of query: 247
length of database: 9,014,727
effective HSP length: 94
effective length of query: 153
effective length of database: 5,538,983
effective search space: 847464399
effective search space used: 847464399
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC123571.7


