

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122729.6 + phase: 0 /pseudo
(247 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
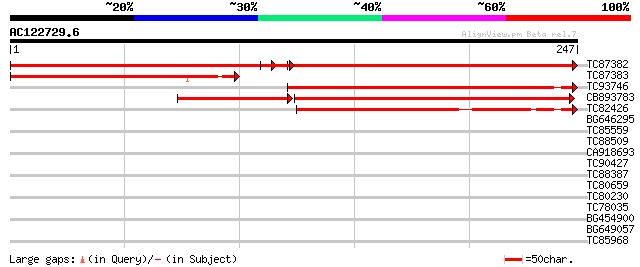
Score E
Sequences producing significant alignments: (bits) Value
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 240 e-126
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 166 6e-42
TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical... 147 3e-36
CB893783 weakly similar to GP|22830935|dbj hypothetical protein~... 103 3e-30
TC82426 107 6e-24
BG646295 similar to GP|13992688|gb Putative retroelement {Oryza ... 39 0.002
TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC ... 33 0.13
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 32 0.23
CA918693 similar to SP|P34066|PS11 Proteasome subunit alpha type... 30 0.86
TC90427 weakly similar to GP|16648830|gb|AAL25605.1 AT5g39670/MI... 30 1.1
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 27 7.3
TC80659 similar to PIR|T48607|T48607 probable transcription fact... 27 9.5
TC80230 similar to PIR|T05160|T05160 hypothetical protein F18E5.... 27 9.5
TC78035 similar to GP|8778540|gb|AAF79548.1| F22G5.9 {Arabidopsi... 27 9.5
BG454900 weakly similar to PIR|T39903|T399 serine-rich protein -... 27 9.5
BG649057 similar to PIR|H86201|H86 hypothetical protein [importe... 27 9.5
TC85968 homologue to SP|P80269|NUIM_SOLTU NADH-ubiquinone oxidor... 27 9.5
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 240 bits (612), Expect(3) = e-126
Identities = 123/126 (97%), Positives = 123/126 (97%)
Frame = +1
Query: 122 KDMFLMSSSITKVVKMLHQTI*KKS*SSE*LITKIHTSCNGSTRITR*KYLNTPLSHFPL 181
K FLMSSSITKVVKMLHQTI*KKS*SSE*LITKIHTSCNGSTRITR*KYLNTPLSHFPL
Sbjct: 1411 KTWFLMSSSITKVVKMLHQTI*KKS*SSE*LITKIHTSCNGSTRITR*KYLNTPLSHFPL 1590
Query: 182 TRIIKKMYGVILFQWILATLILGCHGNMIVVHYMMVMPTSTLL*NMVLKSSWHVYHLMNS 241
TRIIKK YGVILFQWILATLILGCHGNMIVVHYMMVMPTSTLL*NMVLKSSWHVYHLMNS
Sbjct: 1591 TRIIKKRYGVILFQWILATLILGCHGNMIVVHYMMVMPTSTLL*NMVLKSSWHVYHLMNS 1770
Query: 242 LKEKRN 247
LKEKRN
Sbjct: 1771 LKEKRN 1788
Score = 218 bits (554), Expect(3) = e-126
Identities = 101/116 (87%), Positives = 105/116 (90%)
Frame = +2
Query: 1 CVMCQGYGHIAIDCVNYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGAD 60
C +CQGYGHIA+DCVN K TIVN EINNIFEEERED+YESFEDETL +PIYDEEYVGAD
Sbjct: 1046 CFICQGYGHIALDCVNQKVFTIVNEEINNIFEEEREDVYESFEDETLGEPIYDEEYVGAD 1225
Query: 61 ICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRF 116
ICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEY K F
Sbjct: 1226 ICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYFGD*KTF 1393
Score = 33.9 bits (76), Expect(3) = e-126
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +3
Query: 110 LETKKRFQTSTNKDM 124
LETKKRFQTSTNKDM
Sbjct: 1374 LETKKRFQTSTNKDM 1418
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 166 bits (421), Expect = 6e-42
Identities = 82/101 (81%), Positives = 88/101 (86%), Gaps = 1/101 (0%)
Frame = -2
Query: 1 CVMCQGYGHIAIDCVNYKAVTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGAD 60
CVMCQGYGHIAIDCVNYKAV IVNREINNIFEEEREDI+ESFE ET+ +PIYDEEYVG D
Sbjct: 307 CVMCQGYGHIAIDCVNYKAVIIVNREINNIFEEEREDIHESFEAETMGEPIYDEEYVGTD 128
Query: 61 ICEVFDEEGNIDPIYD-ENGLDDIHEAFEKEEHNEPIYDEE 100
ICEVF EEGN DP+ D E+ DDIHE FEKEE NEP+YD E
Sbjct: 127 ICEVFKEEGNNDPMNDVEHDPDDIHEVFEKEE-NEPLYDGE 8
>TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical
protein~similar to gag-pol polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 1019
Score = 147 bits (372), Expect = 3e-36
Identities = 84/126 (66%), Positives = 92/126 (72%)
Frame = +3
Query: 122 KDMFLMSSSITKVVKMLHQTI*KKS*SSE*LITKIHTSCNGSTRITR*KYLNTPLSHFPL 181
K F+MSSSITK V+ML+ I K+* T IHTSCNGSTRI R* YLNT L HFPL
Sbjct: 36 KTWFVMSSSITKSVRMLNPIIWWKN*RCRQRSTHIHTSCNGSTRIMR*NYLNTQLFHFPL 215
Query: 182 TRIIKKMYGVILFQWILATLILGCHGNMIVVHYMMVMPTSTLL*NMVLKSSWHVYHLMNS 241
T II KMYGV+LFQWILAT ILG GNMIV+HY MVMPT L * M LKSS H+ H S
Sbjct: 216 TTIITKMYGVMLFQWILATFILGGLGNMIVMHYSMVMPTPALS*RMRLKSSLHLCH---S 386
Query: 242 LKEKRN 247
+KEKRN
Sbjct: 387 MKEKRN 404
>CB893783 weakly similar to GP|22830935|dbj hypothetical protein~similar to
gag-pol polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 853
Score = 103 bits (257), Expect(2) = 3e-30
Identities = 62/122 (50%), Positives = 77/122 (62%)
Frame = +3
Query: 125 FLMSSSITKVVKMLHQTI*KKS*SSE*LITKIHTSCNGSTRITR*KYLNTPLSHFPLTRI 184
F+MSS + V+ML+ TI K+ S IT I TS NG ++TR + L+ L FPL +
Sbjct: 258 FVMSSLTVEAVRMLYPTIWWKNWSCRQKITHIDTSYNG*KKVTRSESLSVALYLFPLGKN 437
Query: 185 IKKMYGVILFQWILATLILGCHGNMIVVHYMMVMPTSTLL*NMVLKSSWHVYHLMNSLKE 244
K MYGV+ W+L T LG GNMIV+ Y VMPT T L*NMVLKSSW YH M+ +K
Sbjct: 438 TKTMYGVMSSPWMLVTCFLGDLGNMIVMPYTTVMPTLTHL*NMVLKSSWCPYHPMHLMKA 617
Query: 245 KR 246
KR
Sbjct: 618 KR 623
Score = 45.4 bits (106), Expect(2) = 3e-30
Identities = 23/50 (46%), Positives = 32/50 (64%)
Frame = +2
Query: 74 IYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRFQTSTNKD 123
I++ + +DIHEA E EE E YDEE VLA+ GE L ++ T+T K+
Sbjct: 53 IHEASEEEDIHEALELEEMGEQTYDEEPVLADSGESLIVRRSLHTTTVKE 202
Score = 33.1 bits (74), Expect = 0.10
Identities = 19/41 (46%), Positives = 26/41 (63%)
Frame = +2
Query: 20 VTIVNREINNIFEEEREDIYESFEDETLEKPIYDEEYVGAD 60
+TI+ EI+ EEE DI+E+ E E + + YDEE V AD
Sbjct: 32 ITIIKGEIHEASEEE--DIHEALELEEMGEQTYDEEPVLAD 148
>TC82426
Length = 1017
Score = 107 bits (266), Expect = 6e-24
Identities = 69/122 (56%), Positives = 79/122 (64%)
Frame = +1
Query: 126 LMSSSITKVVKMLHQTI*KKS*SSE*LITKIHTSCNGSTRITR*KYLNTPLSHFPLTRII 185
+MS S +VVKML+ I K+* + TS NGSTRI R*KYLNT LSHFPL II
Sbjct: 145 VMSLSTVEVVKMLYPIIW*KN*IYQQRSAYTLTSFNGSTRIMR*KYLNTLLSHFPLAIII 324
Query: 186 KKMYGVILFQWILATLILGCHGNMIVVHYMMVMPTSTLL*NMVLKSSWHVYHLMNSLKEK 245
KK+YGV+LF W ILG H NMIVVHY M+M L * M SSWH+YH S+ EK
Sbjct: 325 KKIYGVMLFLW-----ILGGHVNMIVVHYTMIMSIHALS*KM--*SSWHLYH---SMMEK 474
Query: 246 RN 247
RN
Sbjct: 475 RN 480
>BG646295 similar to GP|13992688|gb Putative retroelement {Oryza sativa}
[Oryza sativa (japonica cultivar-group)], partial (0%)
Length = 640
Score = 38.9 bits (89), Expect = 0.002
Identities = 16/21 (76%), Positives = 18/21 (85%)
Frame = +1
Query: 227 MVLKSSWHVYHLMNSLKEKRN 247
M +KS WH+YH MNSLKEKRN
Sbjct: 160 MRIKSFWHLYHSMNSLKEKRN 222
>TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC 6.4.1.2) -
potato chloroplast, partial (66%)
Length = 5011
Score = 32.7 bits (73), Expect = 0.13
Identities = 25/75 (33%), Positives = 34/75 (45%), Gaps = 2/75 (2%)
Frame = +3
Query: 29 NIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFE 88
N E+E E S EDE EK + DE+ E DE E GL+D E E
Sbjct: 3546 NSLEDEDEPSENSLEDEPSEKGLEDEDEPSEKGLEDEDEP-------SEKGLEDEDEPSE 3704
Query: 89 K--EEHNEPIYDEEY 101
E+ +EP +++Y
Sbjct: 3705 NSLEDEDEPSENDDY 3749
Score = 28.5 bits (62), Expect = 2.5
Identities = 21/66 (31%), Positives = 30/66 (44%), Gaps = 2/66 (3%)
Frame = +3
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEK-- 89
E+E E S EDE E + DE+ E +++ E GL+D E EK
Sbjct: 3495 EDEDEPSENSLEDEPSENSLEDEDE---------PSENSLEDEPSEKGLEDEDEPSEKGL 3647
Query: 90 EEHNEP 95
E+ +EP
Sbjct: 3648 EDEDEP 3665
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 32.0 bits (71), Expect = 0.23
Identities = 30/100 (30%), Positives = 44/100 (44%), Gaps = 8/100 (8%)
Frame = +1
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVG-ADICEVFDEEGNIDPIYDENGLDDIHEAFE-- 88
EEE E+ E E+E+ + DEEY G A + VF + D I + L+ EA E
Sbjct: 505 EEEEEEEEEEEEEESDYETDSDEEYTGVAMVKPVFVPKSERDTIAERERLEAEEEALEEA 684
Query: 89 -----KEEHNEPIYDEEYVLAEYGEYLETKKRFQTSTNKD 123
+E NE + V+ E + E +K + N D
Sbjct: 685 RKRRMEERRNE---TRQIVVEEIRKDQEIQKNIELEANID 795
>CA918693 similar to SP|P34066|PS11 Proteasome subunit alpha type 1-1 (EC
3.4.25.1) (20S proteasome alpha subunit F1), partial
(43%)
Length = 680
Score = 30.0 bits (66), Expect = 0.86
Identities = 29/103 (28%), Positives = 41/103 (39%), Gaps = 10/103 (9%)
Frame = -2
Query: 7 YGHIAIDCVNYKAVTIVNREINNIFEEEREDIY--------ESFEDETLEKPIYDEEYVG 58
Y +AID + A T + R+ NN RED+ ES + E L + VG
Sbjct: 592 YQAVAIDSRSQAAKTYLERKFNNFTASSREDLIKDALIATRESLQGEELRSSVCTIAVVG 413
Query: 59 ADICEVFDEEGNIDPIYDENGLDDIHEAFE--KEEHNEPIYDE 99
D E F I D+ + + + FE +EE P E
Sbjct: 412 VD--EPFH-------ILDQETVQQLIDTFEIVREEEAAPAEPE 311
>TC90427 weakly similar to GP|16648830|gb|AAL25605.1 AT5g39670/MIJ24_140
{Arabidopsis thaliana}, partial (46%)
Length = 730
Score = 29.6 bits (65), Expect = 1.1
Identities = 18/58 (31%), Positives = 31/58 (53%)
Frame = +1
Query: 37 DIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNE 94
D + S E E LE E+Y ++CEVF+E +E L+++ +AF+ + N+
Sbjct: 268 DFFCSSESEELE-----EKYGSKELCEVFEE--------NEPSLEELKQAFDVFDENK 402
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 26.9 bits (58), Expect = 7.3
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = +1
Query: 1 CVMCQGYGHIAIDCVNYKA 19
C C GHIA++C N KA
Sbjct: 673 CNNCYKQGHIAVECTNEKA 729
>TC80659 similar to PIR|T48607|T48607 probable transcription factor MYB40 -
Arabidopsis thaliana, partial (46%)
Length = 1040
Score = 26.6 bits (57), Expect = 9.5
Identities = 23/107 (21%), Positives = 44/107 (40%), Gaps = 9/107 (8%)
Frame = +2
Query: 44 DETLEKPIYDEEYVGADICEVFDEEGNIDPIYDEN---------GLDDIHEAFEKEEHNE 94
D KPI ++ D + ++E NI ++EN G ++ + K E N+
Sbjct: 359 DPVNHKPIEQKQQTLDDDKNIINQEPNISEEFEENMEMKSLVSDGTKEMTKTELKREENK 538
Query: 95 PIYDEEYVLAEYGEYLETKKRFQTSTNKDMFLMSSSITKVVKMLHQT 141
+D+ L E L + +T ++D ++S++ L T
Sbjct: 539 VAWDDTSELLNNFEMLCSNLDLETWMSQDTNTSANSVSSSSVSLDNT 679
>TC80230 similar to PIR|T05160|T05160 hypothetical protein F18E5.140 -
Arabidopsis thaliana, partial (41%)
Length = 816
Score = 26.6 bits (57), Expect = 9.5
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = +3
Query: 183 RIIKKMYGVILFQWILATLILGCHGNMIVVHYMMVMPTST 222
+I KK V+ W+ LILGC +++ ++ +P T
Sbjct: 690 KIKKKAKQVLYLHWLFLHLILGC*LWVLIAKLLLFIPEET 809
>TC78035 similar to GP|8778540|gb|AAF79548.1| F22G5.9 {Arabidopsis
thaliana}, partial (24%)
Length = 2694
Score = 26.6 bits (57), Expect = 9.5
Identities = 22/93 (23%), Positives = 37/93 (39%), Gaps = 3/93 (3%)
Frame = +3
Query: 53 DEEYVGADICEVFDEEGNIDPIYDEN-GLDD--IHEAFEKEEHNEPIYDEEYVLAEYGEY 109
D +VG + FD+ GN+ + D N LDD I + ++EP+
Sbjct: 90 DPNFVGFETTYKFDDHGNLILLSDPNPNLDDFTITNPYVGPSNSEPLV------------ 233
Query: 110 LETKKRFQTSTNKDMFLMSSSITKVVKMLHQTI 142
F + T KD + ++ VK + Q +
Sbjct: 234 ------FSSDTTKDSTFEDADFSETVKYISQIL 314
>BG454900 weakly similar to PIR|T39903|T399 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (12%)
Length = 258
Score = 26.6 bits (57), Expect = 9.5
Identities = 22/79 (27%), Positives = 38/79 (47%), Gaps = 1/79 (1%)
Frame = +3
Query: 25 REINNIFEEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIH 84
+E + EEE E+ + E +T ++ + + EYV ++ EG+ D + DD H
Sbjct: 42 QEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEV------EGDDDDVAG----DDEH 191
Query: 85 EAFEK-EEHNEPIYDEEYV 102
+ E EH + +EE V
Sbjct: 192 ASEENVHEHLADVKEEEQV 248
>BG649057 similar to PIR|H86201|H86 hypothetical protein [imported] -
Arabidopsis thaliana, partial (8%)
Length = 644
Score = 26.6 bits (57), Expect = 9.5
Identities = 22/71 (30%), Positives = 32/71 (44%), Gaps = 2/71 (2%)
Frame = +2
Query: 32 EEEREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDD--IHEAFEK 89
E+E ED+Y FED LE E Y D + ++G +D +E L +H F
Sbjct: 173 EDEDEDVYGDFED--LETGENHENYKTDDAFAITTQKG-VDREAEERRLKKLALHAKFVS 343
Query: 90 EEHNEPIYDEE 100
++P EE
Sbjct: 344 RYDDDPETPEE 376
>TC85968 homologue to SP|P80269|NUIM_SOLTU NADH-ubiquinone oxidoreductase 23
kDa subunit mitochondrial precursor (EC 1.6.5.3) (EC
1.6.99.3), partial (79%)
Length = 1021
Score = 26.6 bits (57), Expect = 9.5
Identities = 21/74 (28%), Positives = 31/74 (41%)
Frame = +1
Query: 40 ESFEDETLEKPIYDEEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDE 99
E ED + YD + C E +D I + + F E H E +YD+
Sbjct: 481 EEREDGSRRTTRYDIDMTKCIYCGFCQEACPVDAIVEGPNFE-----FSTETHEELLYDK 645
Query: 100 EYVLAEYGEYLETK 113
E +L E G+ ET+
Sbjct: 646 EKLL-ENGDRWETE 684
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.144 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,582,947
Number of Sequences: 36976
Number of extensions: 100154
Number of successful extensions: 630
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 613
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 619
length of query: 247
length of database: 9,014,727
effective HSP length: 94
effective length of query: 153
effective length of database: 5,538,983
effective search space: 847464399
effective search space used: 847464399
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC122729.6


