

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
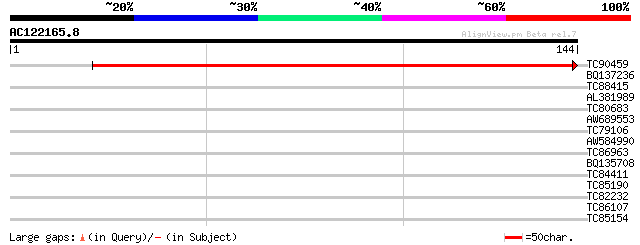
Query= AC122165.8 - phase: 1 /pseudo
(144 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC90459 similar to GP|9758598|dbj|BAB09231.1 gene_id:MDF20.8~unk... 252 3e-68
BQ137236 weakly similar to GP|19070096|emb UL49 tegument protein... 33 0.040
TC88415 similar to GP|18057164|gb|AAL58187.1 putative retinoblas... 31 0.15
AL381989 30 0.44
TC80683 homologue to GP|8777424|dbj|BAA97014.1 gb|AAF56406.1~gen... 30 0.44
AW689553 29 0.57
TC79106 weakly similar to PIR|A86426|A86426 hypothetical protein... 27 2.8
AW584990 weakly similar to PIR|T40618|T406 probable cell wall pr... 26 4.8
TC86963 similar to GP|15148918|gb|AAK84886.1 homeodomain leucine... 26 6.3
BQ135708 25 8.2
TC84411 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersico... 25 8.2
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 25 8.2
TC82232 similar to GP|10177316|dbj|BAB10642. kinesin-like protei... 25 8.2
TC86107 similar to SP|P09444|LE34_GOSHI Late embryogenesis abund... 25 8.2
TC85154 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 25 8.2
>TC90459 similar to GP|9758598|dbj|BAB09231.1 gene_id:MDF20.8~unknown
protein {Arabidopsis thaliana}, partial (58%)
Length = 953
Score = 252 bits (644), Expect = 3e-68
Identities = 123/123 (100%), Positives = 123/123 (100%)
Frame = +1
Query: 22 DTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPS 81
DTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPS
Sbjct: 235 DTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPS 414
Query: 82 NRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAE 141
NRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAE
Sbjct: 415 NRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAE 594
Query: 142 FER 144
FER
Sbjct: 595 FER 603
>BQ137236 weakly similar to GP|19070096|emb UL49 tegument protein
{Pseudorabies virus}, partial (10%)
Length = 1224
Score = 33.1 bits (74), Expect = 0.040
Identities = 22/91 (24%), Positives = 39/91 (42%), Gaps = 1/91 (1%)
Frame = +2
Query: 37 DQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKE-AD 95
++ A + + +T K+ + G PP PK+ P RK K++ +E
Sbjct: 149 NRGDRRAQGAAARRARGRTTKKTRAGRGHPRTPPGDPKKTRPPPHTRKDTKKKRARENGR 328
Query: 96 KRISGTGRENKVDNLRELVGGEKTSVGMAKG 126
KR GR+++ + + GG + AKG
Sbjct: 329 KRRPRRGRKDENNTTDQGAGGNRDPEPKAKG 421
>TC88415 similar to GP|18057164|gb|AAL58187.1 putative retinoblastoma
binding protein {Oryza sativa}, partial (2%)
Length = 1410
Score = 31.2 bits (69), Expect = 0.15
Identities = 18/81 (22%), Positives = 33/81 (40%)
Frame = +1
Query: 28 IPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRF 87
I + DSD S + V+ P+S K+ KI H + ++V+P + R
Sbjct: 436 IKPIADNDSDDSDSGIFRVKRPSSLKAEKRNLKIMASKHSEQQGLKRLKKVLPEGKSNRQ 615
Query: 88 KQKLKEADKRISGTGRENKVD 108
+ + ++ +KVD
Sbjct: 616 QMNFRTSESSDKYNPVNDKVD 678
>AL381989
Length = 364
Score = 29.6 bits (65), Expect = 0.44
Identities = 16/71 (22%), Positives = 36/71 (50%)
Frame = +2
Query: 14 FFGLLLVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQP 73
F G L V +P++G QD+D ++N ++ PN++ T+++ P + P++
Sbjct: 122 FHGQLPVAPIPLANLPNMGEQDNDLNQNIQQPLQQPNNSQDTRRDV-----PQNSQPTET 286
Query: 74 KQQEVIPSNRK 84
+ + +N +
Sbjct: 287 EGRTTTETNEE 319
>TC80683 homologue to GP|8777424|dbj|BAA97014.1
gb|AAF56406.1~gene_id:K9P8.7~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (16%)
Length = 1360
Score = 29.6 bits (65), Expect = 0.44
Identities = 28/88 (31%), Positives = 41/88 (45%), Gaps = 2/88 (2%)
Frame = +1
Query: 45 SVESPNSAAKTKKEEKIYLGP--HGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTG 102
S E+ N ++K +K Y+ ++PK E PS+ + K KLK+ ISG
Sbjct: 667 SSENHNESSKPAVRDKPYISKAERRKLKNEPKHGEAHPSDGNGKDKSKLKD----ISGDL 834
Query: 103 RENKVDNLRELVGGEKTSVGMAKGSSPK 130
+NL+ GG+K S G KG K
Sbjct: 835 HAKDAENLK-TGGGKKISRGQ-KGKLKK 912
>AW689553
Length = 695
Score = 29.3 bits (64), Expect = 0.57
Identities = 26/71 (36%), Positives = 31/71 (43%), Gaps = 9/71 (12%)
Frame = -2
Query: 64 GPHGAPPSQPKQQEVIPSNRKQRFKQKL---------KEADKRISGTGRENKVDNLRELV 114
G PP PKQQE +P R + K+K+ KE K ISG K NLR
Sbjct: 355 GGTSPPPPCPKQQE-MPKGRPYQRKKKI*QRNNMSQSKEQQKGISG----GKRTNLRFTQ 191
Query: 115 GGEKTSVGMAK 125
KTS+ K
Sbjct: 190 *SRKTSIAEMK 158
>TC79106 weakly similar to PIR|A86426|A86426 hypothetical protein AAG29710.1
[imported] - Arabidopsis thaliana, partial (11%)
Length = 1058
Score = 26.9 bits (58), Expect = 2.8
Identities = 16/41 (39%), Positives = 21/41 (51%)
Frame = +3
Query: 52 AAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLK 92
A K K +EK+ P G+ P+ P Q + QRF QK K
Sbjct: 378 ANKIKLQEKVKT-PRGSEPTTPVQSGLSMKRSLQRFLQKRK 497
>AW584990 weakly similar to PIR|T40618|T406 probable cell wall protein -
fission yeast (Schizosaccharomyces pombe), partial (11%)
Length = 299
Score = 26.2 bits (56), Expect = 4.8
Identities = 11/35 (31%), Positives = 16/35 (45%)
Frame = +2
Query: 46 VESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIP 80
V+ A K + L P G PP PK +++ P
Sbjct: 185 VQKGGFGAPKKTRQNKNLSPRGGPPFSPKTRKIAP 289
>TC86963 similar to GP|15148918|gb|AAK84886.1 homeodomain leucine zipper
protein HDZ2 {Phaseolus vulgaris}, partial (87%)
Length = 1519
Score = 25.8 bits (55), Expect = 6.3
Identities = 23/83 (27%), Positives = 33/83 (39%), Gaps = 18/83 (21%)
Frame = +1
Query: 72 QPKQQEVIPSNRKQRFKQKLKEAD---KRISGTGRENKVDNL---------------REL 113
QP+Q + NR+ RFK K E D + S ++ DNL +L
Sbjct: 844 QPRQVAIWFQNRRARFKTKQLEKDYGTLKASFDSLKDDYDNLLQENDKLKEEVNSLKNKL 1023
Query: 114 VGGEKTSVGMAKGSSPKDWLDPH 136
+ +K V SSP+ PH
Sbjct: 1024IPRDKEKVNSEDKSSPEAINSPH 1092
>BQ135708
Length = 997
Score = 25.4 bits (54), Expect = 8.2
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = -2
Query: 42 NAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNR 83
N AS E ++ ++ +IY HG PS+ + SNR
Sbjct: 474 NGASTEDSSALNYPRRNIRIYAESHGRDPSRFARLSASTSNR 349
>TC84411 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersicon
esculentum}, partial (38%)
Length = 804
Score = 25.4 bits (54), Expect = 8.2
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Frame = -3
Query: 15 FGLLLVVDTDGFVIPSLGIQ----DSDQSKNNAASVESPNSAAKTKKEEK 60
F LLL+++T+GF I G+ Q+ AA SP T K +K
Sbjct: 442 FLLLLILNTEGFAITH*GVHPQVYQQGQALLQAADA*SPQQQLLTCKAQK 293
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 25.4 bits (54), Expect = 8.2
Identities = 20/98 (20%), Positives = 42/98 (42%), Gaps = 3/98 (3%)
Frame = +3
Query: 35 DSDQSKNNAASVESPNSAAKTKK---EEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKL 91
+ D+ K E + A +K EEK G+ +PK + P +++ ++ +
Sbjct: 570 EEDEKKTEEVVEEKKDEAVVEEKKVDEEK------GSTSEEPKVETAEPEKEEKKVEETV 731
Query: 92 KEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSP 129
E ++I+ + E+ + E + SV + +G P
Sbjct: 732 VEVVEKIAASTEEDGAKTV-EAIQESIVSVPVTEGEQP 842
>TC82232 similar to GP|10177316|dbj|BAB10642. kinesin-like protein
{Arabidopsis thaliana}, partial (21%)
Length = 1181
Score = 25.4 bits (54), Expect = 8.2
Identities = 12/40 (30%), Positives = 24/40 (60%)
Frame = +3
Query: 70 PSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDN 109
P + ++E+ S+ +Q+ ++LKE DK++ E K+ N
Sbjct: 156 PVEIHEKELEHSSAQQKLDRELKELDKKLEQKEAEMKLVN 275
>TC86107 similar to SP|P09444|LE34_GOSHI Late embryogenesis abundant protein
D-34 (LEA D-34). [Upland cotton] {Gossypium hirsutum},
partial (67%)
Length = 1444
Score = 25.4 bits (54), Expect = 8.2
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = +3
Query: 64 GPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDNLRE 112
GP P + Q +V+ + R ++KQK + +KR GT RE ++ R+
Sbjct: 1122 GPRCKPHNFCFQTKVMHARRTVKYKQKER--NKRKRGTKREEQLGQKRK 1262
>TC85154 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (83%)
Length = 1811
Score = 25.4 bits (54), Expect = 8.2
Identities = 20/98 (20%), Positives = 42/98 (42%), Gaps = 3/98 (3%)
Frame = +3
Query: 35 DSDQSKNNAASVESPNSAAKTKK---EEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKL 91
+ D+ K E + A +K EEK G+ +PK + P +++ ++ +
Sbjct: 114 EEDEKKTEEVVEEKKDEAVVEEKKVDEEK------GSTSEEPKVETAEPEKEEKKVEETV 275
Query: 92 KEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSP 129
E ++I+ + E+ + E + SV + +G P
Sbjct: 276 VEVVEKIAASTEEDGAKTV-EAIQESIVSVPVTEGEQP 386
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,225,334
Number of Sequences: 36976
Number of extensions: 43503
Number of successful extensions: 346
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 345
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 346
length of query: 144
length of database: 9,014,727
effective HSP length: 87
effective length of query: 57
effective length of database: 5,797,815
effective search space: 330475455
effective search space used: 330475455
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC122165.8


