

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122163.4 - phase: 0
(431 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
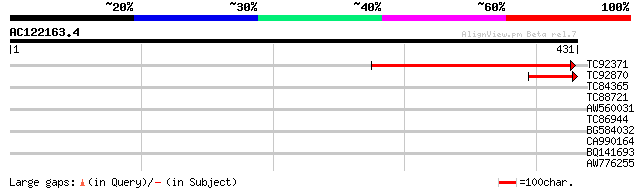
Sequences producing significant alignments: (bits) Value
TC92371 similar to GP|18377809|gb|AAL67091.1 AT3g20300/MQC12_5 {... 181 5e-46
TC92870 weakly similar to GP|21740393|emb|CAD40872. OSJNBa0064H2... 81 9e-16
TC84365 similar to GP|15811367|gb|AAL08940.1 zinc finger protein... 30 1.8
TC88721 similar to GP|18086472|gb|AAL57689.1 unknown protein {Ar... 30 2.4
AW560031 similar to GP|882341|gb|AA LRP1 {Arabidopsis thaliana},... 29 3.1
TC86944 similar to GP|15215806|gb|AAK91448.1 At3g14201 {Arabidop... 28 6.8
BG584032 similar to GP|10177858|dbj gene_id:K18P6.11~unknown pro... 28 6.8
CA990164 similar to GP|9757805|dbj CCR4-associated factor-like p... 28 8.9
BQ141693 weakly similar to GP|160968|gb|AA eggshell protein {Sch... 28 8.9
AW776255 weakly similar to PIR|T07672|T07 cyclin a2-type mitosi... 28 8.9
>TC92371 similar to GP|18377809|gb|AAL67091.1 AT3g20300/MQC12_5 {Arabidopsis
thaliana}, partial (29%)
Length = 647
Score = 181 bits (459), Expect = 5e-46
Identities = 92/158 (58%), Positives = 125/158 (78%), Gaps = 3/158 (1%)
Frame = +3
Query: 276 VDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNI-DGE 334
+++ K GEL L S+TL+S L I+ RSATKITHKAQ++TGLA+KWH+CAT++SFD + +GE
Sbjct: 3 LNVYKTGELALCSVTLLSALSIMFRSATKITHKAQAITGLAAKWHVCATLDSFDGVGEGE 182
Query: 335 TPTALIASAQAMAADISWGSSE-DESGDEEDELNNTKMMPIQT-RTISFHKRQALVTYME 392
T +A I+ + + + G SE D++G EEDE++ TKM+P + TIS+ KRQALV Y E
Sbjct: 183 TLSAQISRERIYPSVGTDGESETDDAGSEEDEIDTTKMIPSYSYSTISYQKRQALVNYFE 362
Query: 393 NNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVG 430
NN+AGITVYGFMLDR+ LH+IFGI+++L LWLL KT+G
Sbjct: 363 NNKAGITVYGFMLDRSTLHTIFGIEMSLVLWLLGKTIG 476
>TC92870 weakly similar to GP|21740393|emb|CAD40872. OSJNBa0064H22.17 {Oryza
sativa}, partial (8%)
Length = 535
Score = 80.9 bits (198), Expect = 9e-16
Identities = 37/37 (100%), Positives = 37/37 (100%)
Frame = +3
Query: 395 RAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVGI 431
RAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVGI
Sbjct: 3 RAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVGI 113
>TC84365 similar to GP|15811367|gb|AAL08940.1 zinc finger protein
{Arabidopsis thaliana}, partial (6%)
Length = 597
Score = 30.0 bits (66), Expect = 1.8
Identities = 16/68 (23%), Positives = 33/68 (48%), Gaps = 3/68 (4%)
Frame = -2
Query: 37 YVDQSNLCKAGISWSVFFTLAYVVPILSHFL---LDCSTTCDADHRRPYHVPVQISLSVF 93
Y+ + +C G +S++F + + H L + STTC + +H+P + SL+
Sbjct: 581 YIHCNKMCLKGELFSIYFPKTIIEQVNQHSLPSVMPGSTTCLSFEHCTFHIPYRDSLTTK 402
Query: 94 ATLSFISL 101
+ L + +
Sbjct: 401 SLLDSLQI 378
>TC88721 similar to GP|18086472|gb|AAL57689.1 unknown protein {Arabidopsis
thaliana}, complete
Length = 835
Score = 29.6 bits (65), Expect = 2.4
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -1
Query: 176 SGIILCTLELCSWFYRTSIFFLVCV 200
+GI+L TLE C W +T+ F CV
Sbjct: 82 NGIVLRTLERCCWRKKTTTLFRTCV 8
>AW560031 similar to GP|882341|gb|AA LRP1 {Arabidopsis thaliana}, partial
(24%)
Length = 756
Score = 29.3 bits (64), Expect = 3.1
Identities = 15/60 (25%), Positives = 32/60 (53%)
Frame = +2
Query: 197 LVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLL 256
+ ++ R++GYL ++ + F QR + +L+ L + RNL ++ H+ +L +L
Sbjct: 572 IALLMLRVLGYLLLEEGNVFLLPRQRRWLLVVVLVVQLLVLRNLDLLLHKLLLILLLLIL 751
>TC86944 similar to GP|15215806|gb|AAK91448.1 At3g14201 {Arabidopsis
thaliana}, partial (50%)
Length = 2552
Score = 28.1 bits (61), Expect = 6.8
Identities = 14/51 (27%), Positives = 26/51 (50%)
Frame = -3
Query: 321 ICATINSFDNIDGETPTALIASAQAMAADISWGSSEDESGDEEDELNNTKM 371
+ TI+ E+P + + + D+S+GSS E DE+ + +TK+
Sbjct: 1878 VSPTISPLTTFSDESPP----TGELVIDDLSYGSSSHEFADEDSQSRSTKL 1738
>BG584032 similar to GP|10177858|dbj gene_id:K18P6.11~unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 789
Score = 28.1 bits (61), Expect = 6.8
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = -1
Query: 13 NKENSGIYHDGKELRRFRAYLRWVYVDQ 40
+KE +G Y D + ++R Y + VYVD+
Sbjct: 696 SKEGNGFYDDEEGMKRMMYYYQLVYVDE 613
>CA990164 similar to GP|9757805|dbj CCR4-associated factor-like protein
{Arabidopsis thaliana}, partial (45%)
Length = 731
Score = 27.7 bits (60), Expect = 8.9
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = -1
Query: 61 PILSHFLLDCSTTCDADHRRPYH 83
P H C T+ +A HR+PYH
Sbjct: 239 PCPIHHANSCKTSSNASHRKPYH 171
>BQ141693 weakly similar to GP|160968|gb|AA eggshell protein {Schistosoma
japonicum}, partial (12%)
Length = 1226
Score = 27.7 bits (60), Expect = 8.9
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = -3
Query: 61 PILSHFLLDCSTTCDADHRRPYHVPVQISLSVF 93
P L+ FL CS++CD R HV +++ +F
Sbjct: 915 PPLAPFLRSCSSSCDDQLRESLHVHLRLWSPIF 817
>AW776255 weakly similar to PIR|T07672|T07 cyclin a2-type mitosis-specific -
soybean, partial (22%)
Length = 704
Score = 27.7 bits (60), Expect = 8.9
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = +2
Query: 124 LKIQRGYAIQMKRTMKLILLWGLPCFICQ 152
LKI + Q K K LLW +PCF C+
Sbjct: 323 LKILQHQQ*QRKSPCKCSLLWKIPCFQCR 409
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,504,917
Number of Sequences: 36976
Number of extensions: 254779
Number of successful extensions: 2086
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 2065
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2083
length of query: 431
length of database: 9,014,727
effective HSP length: 99
effective length of query: 332
effective length of database: 5,354,103
effective search space: 1777562196
effective search space used: 1777562196
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC122163.4


