

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.2 - phase: 0 /pseudo
(547 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
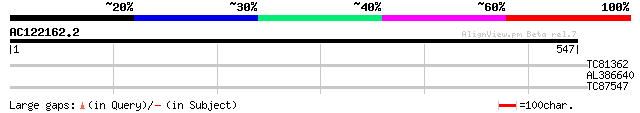
Score E
Sequences producing significant alignments: (bits) Value
TC81362 similar to GP|19911201|dbj|BAB86927. glucosyltransferase... 30 1.8
AL386640 similar to GP|23237918|dbj contains ESTs AU161005(C5315... 30 2.4
TC87547 weakly similar to PIR|F84784|F84784 probable glucosyl tr... 28 6.9
>TC81362 similar to GP|19911201|dbj|BAB86927. glucosyltransferase-9 {Vigna
angularis}, partial (70%)
Length = 1347
Score = 30.4 bits (67), Expect = 1.8
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +1
Query: 174 NFDELFASLYNDYTQKQEDKIWCI 197
+F+EL NDY + + DK+WC+
Sbjct: 751 SFEELEKEYVNDYKKVRNDKVWCV 822
>AL386640 similar to GP|23237918|dbj contains ESTs AU161005(C53159)
C27824(C53159)~exportin tRNA-like~nuclear export
receptor for, partial (4%)
Length = 491
Score = 30.0 bits (66), Expect = 2.4
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 4/65 (6%)
Frame = +2
Query: 485 KTIIQWLAASQAGRPFIAYYTFGSGALQNLDKILVGVVAGLVL----DFVAGMDSWRPME 540
+++I+ L Q G YT+ +G + +L I V VAG +L DF SW E
Sbjct: 95 QSLIENLRLQQNGSLVFR*YTYNNGQISSLRSIRV*FVAGRLLVRGNDFGLQKQSWNGGE 274
Query: 541 HASGV 545
+ S V
Sbjct: 275 YCSAV 289
>TC87547 weakly similar to PIR|F84784|F84784 probable glucosyl transferase
[imported] - Arabidopsis thaliana, partial (29%)
Length = 878
Score = 28.5 bits (62), Expect = 6.9
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +2
Query: 174 NFDELFASLYNDYTQKQEDKIWCI 197
+F+EL + DY + + DK+WCI
Sbjct: 725 SFEELEPTYARDYKKVRNDKVWCI 796
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,036,352
Number of Sequences: 36976
Number of extensions: 236301
Number of successful extensions: 1301
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1289
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1301
length of query: 547
length of database: 9,014,727
effective HSP length: 101
effective length of query: 446
effective length of database: 5,280,151
effective search space: 2354947346
effective search space used: 2354947346
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC122162.2


