

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.15 - phase: 0
(342 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
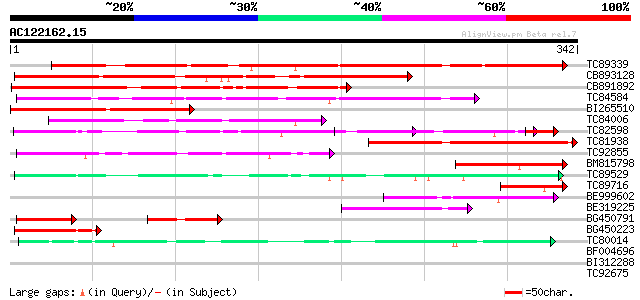
Score E
Sequences producing significant alignments: (bits) Value
TC89339 similar to GP|17380986|gb|AAL36305.1 unknown protein {Ar... 259 1e-69
CB893128 weakly similar to PIR|F96671|F966 hypothetical protein ... 202 1e-52
CB891892 178 3e-45
TC84584 similar to PIR|C84547|C84547 hypothetical protein At2g17... 154 5e-38
BI265510 weakly similar to PIR|C84555|C845 hypothetical protein ... 147 6e-36
TC84006 homologue to GP|9758228|dbj|BAB08727.1 gb|AAD22308.1~gen... 133 1e-31
TC82598 weakly similar to PIR|C84547|C84547 hypothetical protein... 112 2e-25
TC81938 similar to PIR|C84547|C84547 hypothetical protein At2g17... 98 5e-21
TC92855 84 8e-17
BM815798 homologue to GP|7292702|gb|A CG15741 gene product {Dros... 80 1e-15
TC89529 homologue to PIR|H86400|H86400 protein T17H3.7 [imported... 68 6e-12
TC89716 56 2e-08
BE999602 weakly similar to GP|6686405|gb|A F1E22.11 {Arabidopsis... 52 4e-07
BE319225 50 1e-06
BG450791 weakly similar to GP|9758228|dbj| gb|AAD22308.1~gene_id... 47 8e-06
BG450223 homologue to PIR|H71159|H711 hypothetical protein PH047... 46 2e-05
TC80014 weakly similar to GP|21536784|gb|AAM61116.1 unknown {Ara... 44 1e-04
BF004696 33 0.16
BI312288 similar to PIR|T49048|T490 hypothetical protein T5P19.1... 33 0.21
TC92675 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 32 0.35
>TC89339 similar to GP|17380986|gb|AAL36305.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 1290
Score = 259 bits (661), Expect = 1e-69
Identities = 143/316 (45%), Positives = 202/316 (63%), Gaps = 5/316 (1%)
Frame = +3
Query: 26 DLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPDSINNNNEIIDNINNANTSFCYL 85
DLIRFRSVCSTW SS +PN H ILP++FP+ K I N+ N FC+L
Sbjct: 3 DLIRFRSVCSTWRSSSVPNHHYILPYKFPVFKL-----------PFISEQNDKNPPFCHL 149
Query: 86 SKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGKTQLLNPPFVSLKSTAFSHLPNA- 144
SK++ FL+KPPQ+QQ+Q+TL+R PWLI++++N+ GKT+ +P F S LP
Sbjct: 150 SKQNIFLIKPPQQQQEQQTLLR--PWLIRVSQNTRGKTRFFHPLFCD------SPLPRIN 305
Query: 145 --VNFIKFSAQHLRTDFIIDEDDFTFQNQ--HGSYLYPQTVLAVTCHGKKPLVLGTLSYC 200
++F K S HL ++ + D Q+Q + Y++P+ V+AVTCHGK PL++ T+S
Sbjct: 306 LILDFRKLSLLHLGSNTFTRDFDTYSQHQPFNSDYMFPEKVIAVTCHGKNPLIVATIS-S 482
Query: 201 SSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRFYVLDQSGQTVTVGPDSSTELAAHPLY 260
+P+L +++W + ++S GDICLFKGR Y +D+ G+T+ VG DS+ L A PL
Sbjct: 483 HPEPLLSKCGNEKWKVIPDMSMEFGDICLFKGRAYAVDKIGKTIMVGSDSNVHLVAEPLD 662
Query: 261 QRCVRGKNRKLLVESEGELLLLDIHQTFFQFSMKIFKLDENRKKWEKLKKLGDRILFVGS 320
G+N+KLLVES+G+LLL +H F S+ +FKL+E KKW KL LGD++LF+G
Sbjct: 663 G----GENKKLLVESDGDLLLACVH-AFPVISVDLFKLNEKEKKWVKLTSLGDKVLFLGI 827
Query: 321 GCSFSASALDLCLPKG 336
CSFSAS DL + KG
Sbjct: 828 KCSFSASFSDLYVAKG 875
>CB893128 weakly similar to PIR|F96671|F966 hypothetical protein F13O11.14
[imported] - Arabidopsis thaliana, partial (8%)
Length = 768
Score = 202 bits (515), Expect = 1e-52
Identities = 124/253 (49%), Positives = 156/253 (61%), Gaps = 13/253 (5%)
Frame = +3
Query: 4 VDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDH-NILPFQFPLLKYVPAP 62
VDW+ELP+++L LISQ +D ELDLI FRS+CSTW SS IPN H NI F+FP KY+P P
Sbjct: 63 VDWSELPKELLNLISQCIDNELDLIHFRSICSTWRSSSIPNHHSNISSFKFP--KYIPPP 236
Query: 63 DSI-NNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITK---- 117
S N++N+II I N+N+ F SKRS FLVK P Q + RPWL+QI
Sbjct: 237 YSTKNDDNDIIHFIKNSNSPFDSFSKRSLFLVKAPPHQT-----VVLRPWLMQICPSSLP 401
Query: 118 NSSGKTQLL--NPPF-----VSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQN 170
N GK L +PP L+ T + ++F K S HL T +I N
Sbjct: 402 NKRGKIMLTYSHPPIRHESAKQLQETLIERA-DVLDFNKLSVLHLGTRYI--------SN 554
Query: 171 QHGSYLYPQTVLAVTCHGKKPLVLGTLSYCSSKPVLFHDQDKRWTPVSNLSTANGDICLF 230
Q+G P+T LAVTCHGK PL+LGTLS+ S PV+F +++W SNLST GDICLF
Sbjct: 555 QYGKC--PETALAVTCHGKNPLLLGTLSHFSHLPVIFRGCNEQWKLFSNLSTVRGDICLF 728
Query: 231 KGRFYVLDQSGQT 243
KGRFYV++ SG+T
Sbjct: 729 KGRFYVVEMSGRT 767
>CB891892
Length = 587
Score = 178 bits (451), Expect = 3e-45
Identities = 104/206 (50%), Positives = 136/206 (65%), Gaps = 1/206 (0%)
Frame = +2
Query: 2 ASVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDH-NILPFQFPLLKYVP 60
A DW+ LP D++ LISQ +D E+DLIRFRSVC W SS I H NILPF+FPL K+
Sbjct: 26 AMSDWSALPMDLVNLISQGIDTEIDLIRFRSVCCNWRSSSIRKHHLNILPFEFPLFKFRS 205
Query: 61 APDSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSS 120
DSI N+N I FCYLSK SFFL+KPP EQQQ TLIR RPWL++I + S+
Sbjct: 206 LFDSIINDNTI--------PFFCYLSKCSFFLIKPPHEQQQ--TLIRHRPWLVRIRQYSN 355
Query: 121 GKTQLLNPPFVSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGSYLYPQT 180
GKTQ+LN P +S +S F L ++++F K S HL TDFI+ +D + Y+YP+
Sbjct: 356 GKTQILN-PLISYQS-PFDSL-HSIDFNKLSVIHLGTDFIVHDDSLLY-----DYMYPEK 511
Query: 181 VLAVTCHGKKPLVLGTLSYCSSKPVL 206
++A T + KKP+VLG + +SKP+L
Sbjct: 512 IVAFTGNEKKPIVLGAFT-TTSKPIL 586
>TC84584 similar to PIR|C84547|C84547 hypothetical protein At2g17030
[imported] - Arabidopsis thaliana, partial (4%)
Length = 812
Score = 154 bits (389), Expect = 5e-38
Identities = 106/283 (37%), Positives = 152/283 (53%), Gaps = 4/283 (1%)
Frame = +3
Query: 5 DWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPDS 64
DW++LP+D+L LIS +LD E +RFRSVCS+W SS N H+ L L ++P P
Sbjct: 36 DWSQLPKDLLQLISSKLDSEFYQLRFRSVCSSWRSSVPKNHHHHL----TLPSHLPTPSD 203
Query: 65 INNNNEIIDNINNANTSFCYLSKRSFFLVKPP--QEQQQQETLIRCRPWLIQITKNSSGK 122
N N++++ LSKR+ FL+ PP QQ+TL PWLI+I +S +
Sbjct: 204 SN-------NLHHSKYITFPLSKRTIFLITPPTNHHHHQQQTL-NNNPWLIKIGPDSRDR 359
Query: 123 TQLLNPPFVSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGSYLYPQTVL 182
T+L NP +S +LP +NF + L +F+I F Y+ VL
Sbjct: 360 TRLWNP--LSRDKQLSIYLPQVINFNEHRVIDLGREFVIG----NFNEYSSLYMEKVIVL 521
Query: 183 AVTCHGKKP--LVLGTLSYCSSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRFYVLDQS 240
G K VL T+ + S K LF D+RWT V + + D+C+FKGR +D +
Sbjct: 522 DADMWGGKERCSVLFTI-HISGKIALFRCGDERWTIVPEMPSPFDDVCVFKGRPVAVDGT 698
Query: 241 GQTVTVGPDSSTELAAHPLYQRCVRGKNRKLLVESEGELLLLD 283
G+TV + PD S +L A P++ G ++K LVES+GELLL+D
Sbjct: 699 GRTVALRPDLSLDLVAEPVF-----GGDKKFLVESDGELLLVD 812
>BI265510 weakly similar to PIR|C84555|C845 hypothetical protein At2g17690
[imported] - Arabidopsis thaliana, partial (8%)
Length = 545
Score = 147 bits (371), Expect = 6e-36
Identities = 73/113 (64%), Positives = 90/113 (79%), Gaps = 2/113 (1%)
Frame = +2
Query: 1 MASVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVP 60
MA DW++LP+D+L LISQR+D E+DLIRFRS+CS W SS IPN HNILPF+FPLLK
Sbjct: 116 MAEADWSQLPKDLLTLISQRIDTEIDLIRFRSICSNWRSSSIPNHHNILPFKFPLLKC-- 289
Query: 61 APDSIN-NNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQ-QQQETLIRCRPW 111
+SI N+NEIID+INN ++ FCYLSKRS FLVKPP Q ++++TLI RPW
Sbjct: 290 EDNSIKYNDNEIIDSINNTDSLFCYLSKRSIFLVKPPPSQLEEKQTLIPRRPW 448
>TC84006 homologue to GP|9758228|dbj|BAB08727.1
gb|AAD22308.1~gene_id:MZF18.8~similar to unknown protein
{Arabidopsis thaliana}, partial (4%)
Length = 597
Score = 133 bits (334), Expect = 1e-31
Identities = 77/172 (44%), Positives = 103/172 (59%), Gaps = 4/172 (2%)
Frame = +2
Query: 24 ELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPDSINNNNEIIDNINNANTSFC 83
E+DLI FRSVCSTW SS +P H FQFPLLK+ PD IN N SFC
Sbjct: 137 EVDLIHFRSVCSTWRSSSVPKHHRNFRFQFPLLKFPFNPDFINRNK-----------SFC 283
Query: 84 YLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGKTQLLNPPFVSLKSTAFSHLPN 143
Y++K++ FL+KPP QQE + RP LI+I +N+ GK++L +P L L N
Sbjct: 284 YITKQNIFLIKPP----QQEQTLNLRPSLIRICQNARGKSKLYHP----LLQNGDKFLYN 439
Query: 144 -AVNFIKFSAQHLRTDFIIDEDDFTFQNQ---HGSYLYPQTVLAVTCHGKKP 191
A++F K S HL ++F + + DFTF ++ Y+YP+ V+ VTCHGKKP
Sbjct: 440 FALDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDYYMYPKKVVVVTCHGKKP 595
>TC82598 weakly similar to PIR|C84547|C84547 hypothetical protein At2g17030
[imported] - Arabidopsis thaliana, partial (2%)
Length = 1050
Score = 112 bits (281), Expect = 2e-25
Identities = 84/246 (34%), Positives = 121/246 (49%), Gaps = 2/246 (0%)
Frame = +1
Query: 3 SVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAP 62
+ DW+ L +D+L LIS RLD E D+IRFRSVCSTW IPN + LPF+ P
Sbjct: 67 AADWSTLHKDLLLLISSRLDNEADIIRFRSVCSTWRFF-IPN--HSLPFKLPHF------ 219
Query: 63 DSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGK 122
+ ++ CYLSK + FL+KPPQ++QQQ RPWLI+I KNS +
Sbjct: 220 --------------HTSSLSCYLSKHNIFLIKPPQQEQQQ----TLRPWLIRIAKNSCRE 345
Query: 123 TQLLNPPFVSLKSTAFSHLPNAVNFIKFSAQHLRTDFIID--EDDFTFQNQHGSYLYPQT 180
TQL + P S+ S+ + H P+ ++ KFS L D ++ +D T + L
Sbjct: 346 TQLFH-PLRSVDSSPY-HFPHVLDLKKFSVVRLGADVVMGRYNEDKTVLERSKELLSQFE 519
Query: 181 VLAVTCHGKKPLVLGTLSYCSSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRFYVLDQS 240
+ + H + P LG ++ H ++K + L+ NG LF + LD
Sbjct: 520 LYQLKIHRQDPTPLGHGMGKKVVAIMQHGKNKETVALGTLN-QNGHPVLFHCK*KALDVD 696
Query: 241 GQTVTV 246
V V
Sbjct: 697 PMHVNV 714
Score = 76.6 bits (187), Expect(2) = 4e-18
Identities = 51/130 (39%), Positives = 73/130 (55%), Gaps = 7/130 (5%)
Frame = +2
Query: 197 LSYCSSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRFYVLDQSGQTVTVGPDSSTELAA 256
+ YCS+ +K WT + +ST DIC+FK RFY +++ G+TV G D S EL A
Sbjct: 653 IQYCSTA------SEKHWTLIPCMSTYFLDICVFKERFYAVNKIGRTVAFGLDYSVELVA 814
Query: 257 HPLYQRCVRGKNRKLLVESEGELLLLDIHQTF-FQF------SMKIFKLDENRKKWEKLK 309
V G + K LVE EGELLL+DI + F F +K+F+L+E +K+W + K
Sbjct: 815 -----EYVNGGDMKFLVECEGELLLVDICDSHCFGFPGEKGLKLKVFRLNERKKEW-RSK 976
Query: 310 KLGDRILFVG 319
L +R +G
Sbjct: 977 SLRNRGFVLG 1006
Score = 32.0 bits (71), Expect(2) = 4e-18
Identities = 14/20 (70%), Positives = 17/20 (85%)
Frame = +3
Query: 312 GDRILFVGSGCSFSASALDL 331
G +LF+G+GCSFSASA DL
Sbjct: 984 GIGVLFLGNGCSFSASASDL 1043
>TC81938 similar to PIR|C84547|C84547 hypothetical protein At2g17030
[imported] - Arabidopsis thaliana, partial (1%)
Length = 916
Score = 97.8 bits (242), Expect = 5e-21
Identities = 53/128 (41%), Positives = 85/128 (66%), Gaps = 2/128 (1%)
Frame = +1
Query: 217 VSNLSTANG-DICLFKGRFYVLDQSGQTVTVGPDSSTELAAHPLYQRCVRGKNRKLLVES 275
+ N+S ++G ++C+FKGR V+D++G+T +GPD +T L A P++ GK++ L+ S
Sbjct: 1 IPNMSNSHGGNVCVFKGRPCVVDKTGRTFMIGPDLTTHLIAEPVFG----GKSKFLVESS 168
Query: 276 EGELLLLDIHQTF-FQFSMKIFKLDENRKKWEKLKKLGDRILFVGSGCSFSASALDLCLP 334
E ELLL+D ++ + + + +LDE K+W KL LGD++LF+G+G SFSASA +L
Sbjct: 169 EFELLLVDRYENYRVPVWIDVLRLDEKEKRWVKLANLGDKVLFLGNGYSFSASASELGFA 348
Query: 335 KGKLCHLY 342
G C +Y
Sbjct: 349 NGN-CVIY 369
>TC92855
Length = 625
Score = 84.0 bits (206), Expect = 8e-17
Identities = 69/196 (35%), Positives = 96/196 (48%), Gaps = 4/196 (2%)
Frame = +2
Query: 5 DWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPN--DHNILPFQFPLLKYVPAP 62
DW+ LP ++L LISQ+LD L L+RFRSVC+TW SS IPN N P P P P
Sbjct: 56 DWSHLPSELLQLISQKLDTVLCLLRFRSVCATWRSSSIPNHKHKNSSPLNIP---QFPEP 226
Query: 63 DSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGK 122
+ N I NI + L + FL+KPPQ Q PW I+I + GK
Sbjct: 227 NFDPFN---IRNIIKSKYITSRLITHTIFLIKPPQHQPS------LNPWWIRIGPDFKGK 379
Query: 123 TQLLNPPFVSLKSTAFSHLPNAVNFIKFSAQHL--RTDFIIDEDDFTFQNQHGSYLYPQT 180
TQL +P S S N ++F K S + L + F + +Q+ S+++
Sbjct: 380 TQLWHP--FSSGQNLSSDFNNVLDFNKLSFRVLGCMSYFYLHHLIGGYQS---SFIH--- 535
Query: 181 VLAVTCHGKKPLVLGT 196
+A C G++P V+ T
Sbjct: 536 CVAAVCLGEQPPVVAT 583
>BM815798 homologue to GP|7292702|gb|A CG15741 gene product {Drosophila
melanogaster}, partial (14%)
Length = 577
Score = 79.7 bits (195), Expect = 1e-15
Identities = 41/80 (51%), Positives = 51/80 (63%), Gaps = 13/80 (16%)
Frame = +3
Query: 270 KLLVESEGELLLLDIHQTFFQFSMKIFKLDENRKKWE-------------KLKKLGDRIL 316
K+LVESEG LLLL I ++ FS+ F+LDE +KKW KL+ GDR+
Sbjct: 36 KMLVESEGRLLLLGIEESSNYFSIDFFELDEKKKKWVRLMDFDEKEKKWVKLRNFGDRVF 215
Query: 317 FVGSGCSFSASALDLCLPKG 336
F+G GCSFSASA DLC+ KG
Sbjct: 216 FIGRGCSFSASASDLCIQKG 275
>TC89529 homologue to PIR|H86400|H86400 protein T17H3.7 [imported] -
Arabidopsis thaliana, partial (1%)
Length = 1745
Score = 67.8 bits (164), Expect = 6e-12
Identities = 92/358 (25%), Positives = 134/358 (36%), Gaps = 27/358 (7%)
Frame = +1
Query: 4 VDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPD 63
V+W ELP++I+ IS+RL + D IRFR VC TW+SS +P LP Q P+L
Sbjct: 448 VEWAELPQEIIESISKRLTIYADYIRFRCVCRTWNSS-VPKTPLHLPPQLPILML----- 609
Query: 64 SINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGKT 123
++ SF LS + P WL+ + + S+
Sbjct: 610 --------------SDESFFDLSTNKIHRLNFPLPSGPTRICGSSHGWLVILDETSN--L 741
Query: 124 QLLNPPFVSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGSYLYPQTVLA 183
LLNP V S L T + + F F N++ YL +
Sbjct: 742 NLLNP----------------VTHATLSLPSLHTFPYLVKKLFAF-NKNNLYL----IKV 858
Query: 184 VTCHGKKP---------LVLGTLSY----CSSKPVLFHDQDKRWTPVSNLSTANGDICLF 230
V P L L L++ C S +L + WT D
Sbjct: 859 VLSSNPSPNDDFAAFAILGLRHLAFCRKGCDSWVLLDANVGDHWT----------DAVYK 1008
Query: 231 KGRFYVLDQSGQT----VTVGPDSS-----TELAAHPLYQRCVRGKNRKLL---VESEGE 278
G F+ + SG T V GP S T L + ++ G+ L+ E E
Sbjct: 1009NGSFFAMSTSGITAVCDVVEGPRVSYIPPPTSLYYNDVFYAVFSGEEMLLIQRCCTEEYE 1188
Query: 279 LLLLDIHQTFFQFSMKIFKLDENRKKWEKLKKLGDRILFVGSGCSFSASALDL--CLP 334
+ + I+K++ N KWE+++ LG+ LF+G S S SA D C P
Sbjct: 1189PRDFEPYPDMSTVKFWIYKMNWNMLKWEEIQTLGEHSLFIGKNYSLSFSAADFAGCCP 1362
>TC89716
Length = 735
Score = 55.8 bits (133), Expect = 2e-08
Identities = 28/42 (66%), Positives = 31/42 (73%), Gaps = 2/42 (4%)
Frame = +2
Query: 297 KLDENRKKWEKLKKLGDRILFVGSG--CSFSASALDLCLPKG 336
KL+E KKW KL LGDRILF+G G CSFSA A DLC+ KG
Sbjct: 2 KLNEKEKKWVKLTSLGDRILFLGLGLVCSFSACASDLCVAKG 127
>BE999602 weakly similar to GP|6686405|gb|A F1E22.11 {Arabidopsis thaliana},
partial (4%)
Length = 579
Score = 51.6 bits (122), Expect = 4e-07
Identities = 39/125 (31%), Positives = 58/125 (46%), Gaps = 19/125 (15%)
Frame = +1
Query: 226 DICLFKGRFYVLDQSGQTVTVGPDSSTELAAHPLYQRCVRGKNRKLLVESEGELLLLDIH 285
D+ +FKG+FYV D+ G + S + P C G N+K LVES G L ++D +
Sbjct: 22 DLIVFKGQFYVTDKWGTISWIDISSLKLIQFSP--PLCGFG-NKKHLVESCGNLYVVDRY 192
Query: 286 QTFFQFSM-------------------KIFKLDENRKKWEKLKKLGDRILFVGSGCSFSA 326
+ +M K++KLDE KW +K L +R + C+FS
Sbjct: 193 SYYEDENMGGNYDGRRHQARYEVVECFKVYKLDEEWGKWVDVKDLKNRAFILSPSCNFSV 372
Query: 327 SALDL 331
SA +L
Sbjct: 373 SAKEL 387
>BE319225
Length = 242
Score = 50.1 bits (118), Expect = 1e-06
Identities = 31/80 (38%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Frame = +2
Query: 201 SSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRFYVLDQSGQTVTVGPDSST-ELAAHPL 259
+SKP+L ++ + ++ DIC+FKGR YV+ ++G+T T+G D T EL A L
Sbjct: 11 TSKPILLKCSNENRNGIPVMTAWFEDICIFKGRPYVVHKTGRTATLGLDDFTFELVAESL 190
Query: 260 YQRCVRGKNRKLLVESEGEL 279
G + K LVES+G+L
Sbjct: 191 VS---GGGDIKFLVESDGDL 241
>BG450791 weakly similar to GP|9758228|dbj|
gb|AAD22308.1~gene_id:MZF18.8~similar to unknown protein
{Arabidopsis thaliana}, partial (9%)
Length = 646
Score = 47.4 bits (111), Expect = 8e-06
Identities = 22/36 (61%), Positives = 28/36 (77%)
Frame = -2
Query: 5 DWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSS 40
+W++LP + L LISQ+L+ EL LIR RSVCS W SS
Sbjct: 363 EWSQLPHEFLQLISQKLNRELYLIRSRSVCSWWRSS 256
Score = 43.9 bits (102), Expect = 9e-05
Identities = 22/45 (48%), Positives = 28/45 (61%)
Frame = -1
Query: 84 YLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGKTQLLNP 128
YL K FL KPP+ QQ + RPWLI+I + GKT+LL+P
Sbjct: 160 YLIKHDIFLFKPPKHQQN----LHQRPWLIRIGPDFDGKTKLLHP 38
>BG450223 homologue to PIR|H71159|H711 hypothetical protein PH0477 -
Pyrococcus horikoshii, partial (8%)
Length = 604
Score = 46.2 bits (108), Expect = 2e-05
Identities = 24/52 (46%), Positives = 34/52 (65%)
Frame = +1
Query: 4 VDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPL 55
VDW+ LP+++ I + LD +D++RFRSVC ++ SS IP H P FPL
Sbjct: 274 VDWSHLPKELWSKIGKYLDNHIDILRFRSVCESFRSS-IPPSHPNSP-SFPL 423
>TC80014 weakly similar to GP|21536784|gb|AAM61116.1 unknown {Arabidopsis
thaliana}, partial (25%)
Length = 1587
Score = 43.5 bits (101), Expect = 1e-04
Identities = 76/339 (22%), Positives = 125/339 (36%), Gaps = 15/339 (4%)
Frame = +2
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPA---- 61
W +LP ++L +I RL +E D +R +VC +W+ + N ++ Q P L Y P
Sbjct: 167 WADLPAELLEMIISRLALE-DNVRASAVCKSWNF--VANAVRMVN-QSPWLMYFPKFGQW 334
Query: 62 ---PDSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKN 118
D + I+ + CY L +P + + + P+ + K
Sbjct: 335 YEFYDPVQRKTYSIEFPELNGSRVCYTKDGWLLLYRP-----RTDRVFFFNPFTRETIKM 499
Query: 119 SSGKTQLLNPPF-VSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGSYLY 177
P F ++ + AFS P + + + F+ +H+
Sbjct: 500 ---------PRFEMTYQIVAFSCAPTSPDCVLFTVKHVS--------------------- 589
Query: 178 PQTVLAVTCH-GKKPLVLGTLSYCSSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRFYV 236
P V TCH G V T++Y + P + S+ + G FY
Sbjct: 590 PTIVAISTCHPGATEWV--TVNYQNRLPFV--------------SSIWNKLVFCNGLFYC 721
Query: 237 LDQSGQTVTVGPDSSTELAAHPLYQRCVRG---KN---RKLLVESEGELLLLDIHQTFFQ 290
L +G P T +C KN K + E EG+++++ T
Sbjct: 722 LSLTGWLGVFDPSERTWSVLSVPPPKCPENFFAKNWWKGKFMTEQEGDVIVM---YTCSS 892
Query: 291 FSMKIFKLDENRKKWEKLKKLGDRILFVGSGCSFSASAL 329
+ IFKLD+ +WE+LK L LF S S + L
Sbjct: 893 ENPIIFKLDQASMEWEELKTLDGATLFASFLSSHSRTDL 1009
>BF004696
Length = 593
Score = 33.1 bits (74), Expect = 0.16
Identities = 23/76 (30%), Positives = 35/76 (45%), Gaps = 15/76 (19%)
Frame = +2
Query: 25 LDLIRFRSVCSTWHS--SPIPNDHNI-LP------FQFPLLKYVPAP------DSINNNN 69
+D++RFR++C W S P HN+ +P Q + P+P S +N
Sbjct: 8 IDVVRFRTICRFWRSLVPAPPGSHNLCIPHQNYFLLQTKIYHIEPSPHDHNPSTSCSNQG 187
Query: 70 EIIDNINNANTSFCYL 85
II N+N+S YL
Sbjct: 188GIIKGFQNSNSSKLYL 235
>BI312288 similar to PIR|T49048|T490 hypothetical protein T5P19.120 -
Arabidopsis thaliana, partial (18%)
Length = 705
Score = 32.7 bits (73), Expect = 0.21
Identities = 21/55 (38%), Positives = 28/55 (50%)
Frame = +2
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVP 60
W +LP ++L L RLD+ D IR +VC W S + +L Q P L Y P
Sbjct: 68 WADLPAEVLELFLSRLDIG-DNIRASAVCKRWCS--VATSVRVLD-QSPWLMYFP 220
>TC92675 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (17%)
Length = 630
Score = 32.0 bits (71), Expect = 0.35
Identities = 18/54 (33%), Positives = 26/54 (47%)
Frame = +3
Query: 1 MASVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFP 54
M + +P +++ I RL V+ LIRF+SVC +W S F FP
Sbjct: 48 METTTLPHMPNELMIQILLRLPVK-SLIRFKSVCKSWFSLVSDTHFAYSHFNFP 206
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,957,306
Number of Sequences: 36976
Number of extensions: 207425
Number of successful extensions: 1245
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1205
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1217
length of query: 342
length of database: 9,014,727
effective HSP length: 97
effective length of query: 245
effective length of database: 5,428,055
effective search space: 1329873475
effective search space used: 1329873475
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC122162.15


