

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.1 - phase: 0 /pseudo
(660 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
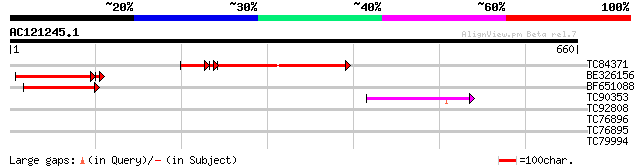
Score E
Sequences producing significant alignments: (bits) Value
TC84371 similar to GP|9759609|dbj|BAB11551.1 gene_id:MNA5.1~unkn... 225 2e-70
BE326156 similar to GP|9759609|dbj gene_id:MNA5.1~unknown protei... 148 6e-36
BF651088 similar to GP|9759609|dbj gene_id:MNA5.1~unknown protei... 137 1e-32
TC90353 weakly similar to GP|9759609|dbj|BAB11551.1 gene_id:MNA5... 49 5e-06
TC92808 similar to GP|9759609|dbj|BAB11551.1 gene_id:MNA5.1~unkn... 34 0.16
TC76896 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 30 3.9
TC76895 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 30 3.9
TC79994 weakly similar to GP|14423560|gb|AAK62462.1 Unknown prot... 28 8.6
>TC84371 similar to GP|9759609|dbj|BAB11551.1 gene_id:MNA5.1~unknown protein
{Arabidopsis thaliana}, partial (25%)
Length = 656
Score = 225 bits (574), Expect(3) = 2e-70
Identities = 109/155 (70%), Positives = 132/155 (84%)
Frame = +1
Query: 242 VKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAI 301
+KNY+RR STGW Y S +R +R GK+G++ +T+EF W+C+LRKQ+EK +AV+LG MS AI
Sbjct: 142 IKNYERRKSTGWEYNSSIRSDRTGKLGSLFNTLEFFWKCVLRKQVEKGMAVLLGIMSVAI 321
Query: 302 LLAEATILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYS 361
LLAEAT+LPS +DLSLFSIL+ + E+LVQ AFVPLMYMC+ TYYSLFK+GM MFYS
Sbjct: 322 LLAEATLLPS-IDLSLFSILIRSVRTQELLVQAFAFVPLMYMCIWTYYSLFKIGMLMFYS 498
Query: 362 LTPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
LTPRQTSSV+LLMICSMVARYA PISYNFLNLI +
Sbjct: 499 LTPRQTSSVNLLMICSMVARYAPPISYNFLNLIRL 603
Score = 58.9 bits (141), Expect(3) = 2e-70
Identities = 26/34 (76%), Positives = 29/34 (84%)
Frame = +2
Query: 199 DYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFV 232
DYDTDEKSMA LRR LR ARE+YYRY+SEY +V
Sbjct: 11 DYDTDEKSMAKLRRHLRNAREEYYRYKSEYITYV 112
Score = 21.6 bits (44), Expect(3) = 2e-70
Identities = 10/11 (90%), Positives = 10/11 (90%)
Frame = +3
Query: 233 LEALELEDTVK 243
LEALELEDT K
Sbjct: 114 LEALELEDTNK 146
>BE326156 similar to GP|9759609|dbj gene_id:MNA5.1~unknown protein
{Arabidopsis thaliana}, partial (22%)
Length = 522
Score = 148 bits (374), Expect(2) = 6e-36
Identities = 69/93 (74%), Positives = 79/93 (84%)
Frame = +3
Query: 7 NVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVAL 66
N ISF WS SYWSTFLLTWAVVPL+QG+EDAGDFTV RL+TS+H NLVFYL +GS+
Sbjct: 207 NGGISFFWSWSYWSTFLLTWAVVPLIQGFEDAGDFTVSERLKTSIHVNLVFYLIVGSIGF 386
Query: 67 FGLILLITLNKFWSGSVRGFAMACSNTFGLVTG 99
FGLILLI +++ WSGS+ G AMACSNTFGLVTG
Sbjct: 387 FGLILLIMMHRHWSGSILGLAMACSNTFGLVTG 485
Score = 21.2 bits (43), Expect(2) = 6e-36
Identities = 8/10 (80%), Positives = 9/10 (90%)
Frame = +2
Query: 101 FLLGFGMSEI 110
F LGFG+SEI
Sbjct: 491 FFLGFGLSEI 520
>BF651088 similar to GP|9759609|dbj gene_id:MNA5.1~unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 627
Score = 137 bits (345), Expect = 1e-32
Identities = 63/88 (71%), Positives = 73/88 (82%)
Frame = +1
Query: 17 SYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLITLN 76
SYWSTFLLTWAV PL+QG+EDAGDFTV RL+TS+H NLVFYL GS+ LFG+ILLI ++
Sbjct: 343 SYWSTFLLTWAVAPLIQGFEDAGDFTVSXRLKTSVHVNLVFYLIXGSIGLFGIILLIMMH 522
Query: 77 KFWSGSVRGFAMACSNTFGLVTGAFLLG 104
W+GS+ GFAM CSNTFGL TGAF G
Sbjct: 523 XTWTGSLMGFAMTCSNTFGLXTGAFXXG 606
>TC90353 weakly similar to GP|9759609|dbj|BAB11551.1 gene_id:MNA5.1~unknown
protein {Arabidopsis thaliana}, partial (13%)
Length = 1119
Score = 49.3 bits (116), Expect = 5e-06
Identities = 46/136 (33%), Positives = 59/136 (42%), Gaps = 10/136 (7%)
Frame = +1
Query: 416 NGKH**CCSLFRERIQQNLPSYHGYIHFTNSRQFL*PGHQLLRELENIQV***CRRYGWI 475
NG+H**CC I QNL + +GYIH +Q L* LE I * RYGW+
Sbjct: 10 NGEH**CCPFLWR*I*QNLSTNNGYIHPLGCKQLL*QSI*FSGGLEEIYFQN*S*RYGWL 189
Query: 476 *SIWSNNFAERALSSSARAQSWRTCFPFSQKF--------QHEYGC*VCQQQGQGSG*EC 527
SI + AER + + R S R ++F Q YG C + C
Sbjct: 190 QSIRVDYLAERTVLA*TRT*SRRASCSTCKEF**Y*R*IKQQYYGAE*C*NERNFHFN*C 369
Query: 528 RK--RR*NNYHGRDQK 541
R N+ GRD+K
Sbjct: 370 RN*WASVKNFEGRDKK 417
>TC92808 similar to GP|9759609|dbj|BAB11551.1 gene_id:MNA5.1~unknown protein
{Arabidopsis thaliana}, partial (6%)
Length = 673
Score = 34.3 bits (77), Expect = 0.16
Identities = 19/41 (46%), Positives = 24/41 (58%)
Frame = +1
Query: 457 LRELENIQV***CRRYGWI*SIWSNNFAERALSSSARAQSW 497
L LE I * * YGW SI +N F +RA+ + RAQ+W
Sbjct: 19 LGSLEEIYF*N*SGGYGWFGSIRNNYFTKRAVLARTRAQNW 141
>TC76896 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (35%)
Length = 1348
Score = 29.6 bits (65), Expect = 3.9
Identities = 19/87 (21%), Positives = 37/87 (41%)
Frame = +1
Query: 137 KLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGES 196
+++ AH D N ++ + + + N + L+ N S +GGR
Sbjct: 385 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL---LIMTSNVGSSVIEKGGRRIGF 555
Query: 197 DMDYDTDEKSMASLRRRLRRAREQYYR 223
D+DYD + S ++ + +QY+R
Sbjct: 556 DLDYDEKDSSYNRIKSLVTEELKQYFR 636
>TC76895 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], complete
Length = 3425
Score = 29.6 bits (65), Expect = 3.9
Identities = 19/87 (21%), Positives = 37/87 (41%)
Frame = +1
Query: 137 KLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGES 196
+++ AH D N ++ + + + N + L+ N S +GGR
Sbjct: 2377 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL---LIMTSNVGSSVIEKGGRRIGF 2547
Query: 197 DMDYDTDEKSMASLRRRLRRAREQYYR 223
D+DYD + S ++ + +QY+R
Sbjct: 2548 DLDYDEKDSSYNRIKSLVTEELKQYFR 2628
>TC79994 weakly similar to GP|14423560|gb|AAK62462.1 Unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 1121
Score = 28.5 bits (62), Expect = 8.6
Identities = 27/111 (24%), Positives = 45/111 (40%)
Frame = +3
Query: 238 LEDTVKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFM 297
+ + +NY RR R S LRP + VG + +L + G +
Sbjct: 189 IPEASRNYTRRVLRWSRQGSSLRPLLLISVGTI------------------ALVALTGLL 314
Query: 298 SFAILLAEATILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTY 348
+F +LL ATI + + +SLF L A G + L + + + V +
Sbjct: 315 TFMLLLLVATI--NAIIVSLFISLAVAGGFLALFFALVTAIYIGALSVAIF 461
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.348 0.152 0.521
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,588,606
Number of Sequences: 36976
Number of extensions: 322363
Number of successful extensions: 3078
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1660
Number of HSP's successfully gapped in prelim test: 172
Number of HSP's that attempted gapping in prelim test: 1381
Number of HSP's gapped (non-prelim): 1895
length of query: 660
length of database: 9,014,727
effective HSP length: 102
effective length of query: 558
effective length of database: 5,243,175
effective search space: 2925691650
effective search space used: 2925691650
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC121245.1


