

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.10 + phase: 0 /pseudo
(330 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
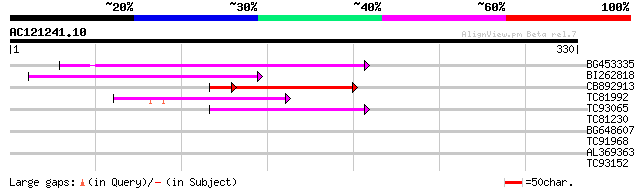
Score E
Sequences producing significant alignments: (bits) Value
BG453335 weakly similar to GP|18071369|g putative gag-pol polypr... 86 3e-17
BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 85 3e-17
CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabi... 65 5e-13
TC81992 weakly similar to PIR|T47841|T47841 hypothetical protein... 62 2e-10
TC93065 55 3e-08
TC81230 30 0.99
BG648607 weakly similar to PIR|F96614|F96 probable copia-type po... 30 1.7
TC91968 similar to PIR|T02550|T02550 NPK1-related protein kinase... 27 8.4
AL369363 weakly similar to GP|10177044|dbj RNA helicase {Arabido... 27 8.4
TC93152 27 8.4
>BG453335 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (11%)
Length = 660
Score = 85.5 bits (210), Expect = 3e-17
Identities = 51/181 (28%), Positives = 98/181 (53%), Gaps = 1/181 (0%)
Frame = +1
Query: 30 LRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQAISHDVLETILNK 89
LR K YW L + L+ + Q E R KE K + K + ++ + I
Sbjct: 25 LRHKIYWRLCKTVTLI*V--LMQPEVHRNAHKELKKKDCKELFLIQ*SLDEGNFKRISKS 198
Query: 90 DTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAH 149
K + ++ + G ++V + +LQ+LRR F+++ M++ + I Y S+ + + N++KA+
Sbjct: 199 TRSKEASNILEKYHEGDDKVEQIKLQSLRRKFKLMQMEDEKKIADYISKLINVVNQIKAY 378
Query: 150 SEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSL-VHEQRIKSNGEEE 208
E+++ I EKI+R++ + D++V +I ES ++ T+ I+EL+SSL H+ + E
Sbjct: 379 GEVVTDQQIVEKIMRTLSPRFDFIVVAIHESKDVKTLKIEELKSSLEAHKLMVYERSSER 558
Query: 209 N 209
+
Sbjct: 559 S 561
>BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (24%)
Length = 615
Score = 85.1 bits (209), Expect = 3e-17
Identities = 47/137 (34%), Positives = 76/137 (55%), Gaps = 1/137 (0%)
Frame = +1
Query: 12 IPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKT 70
+P FDG +Y WA +E +L + + W+ +E + V+ T AQ K KE+K K
Sbjct: 175 LPVFDGENYHIWAARIEAYLEANDLWEAVEEDYEVLPLSDNPTMAQIKNHKERKTRKSKA 354
Query: 71 KNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGE 130
+ LF A+S ++ I+ + IW+ +KT + G R+ Q L R+FE+ MKE E
Sbjct: 355 RATLFAAVSEEIFTRIMTIKSAFEIWNFLKTEYEGDERIRGMQALNLIREFEMQKMKESE 534
Query: 131 TINTYFSRTLTIANKMK 147
TI Y ++ ++IANK++
Sbjct: 535 TIKEYANKLISIANKVR 585
>CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabidopsis
thaliana}, partial (35%)
Length = 837
Score = 65.5 bits (158), Expect(2) = 5e-13
Identities = 33/74 (44%), Positives = 53/74 (71%), Gaps = 1/74 (1%)
Frame = +1
Query: 130 ETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTID 189
E + YFSR ++I+N++K +SE + I EKILRS+ K +++V IEE+ +L+TMTI+
Sbjct: 472 ERVRVYFSRVISISNQLKRNSEKLEDVRIMEKILRSLDPKFEHIVEKIEETKDLETMTIE 651
Query: 190 ELQSSL-VHEQRIK 202
+LQ SL +E++ K
Sbjct: 652 KLQGSLQAYEEKHK 693
Score = 25.8 bits (55), Expect(2) = 5e-13
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = +2
Query: 117 LRRDFEILHMKEGETI 132
LR +FE LHMKE E+I
Sbjct: 443 LRGEFESLHMKESESI 490
>TC81992 weakly similar to PIR|T47841|T47841 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (3%)
Length = 952
Score = 62.4 bits (150), Expect = 2e-10
Identities = 43/108 (39%), Positives = 62/108 (56%), Gaps = 5/108 (4%)
Frame = +3
Query: 61 KEQKLMNLKTKNYLFQAISH--DVLETILN---KDTYKNIWDSMKTFF*GSNRVIRAQLQ 115
+E+ + K +L Q I+ ++E L K ++ S+K + + R RAQLQ
Sbjct: 576 REENIG*AKVVRFLRQKITSFKQLIELCLKQF*KTRHRRRSGSLKK*YQVTARGKRAQLQ 755
Query: 116 ALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKIL 163
AL RD+ IL MK GE+I YF+RT+TI NK++ H E MS + EKIL
Sbjct: 756 AL*RDW*ILQMKNGESIIDYFARTVTIVNKIRMHGEQMSNLTVIEKIL 899
Score = 31.2 bits (69), Expect = 0.58
Identities = 17/36 (47%), Positives = 21/36 (58%)
Frame = +1
Query: 30 LRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKL 65
L SKEYW+LI I GV TE ++K + E KL
Sbjct: 499 LASKEYWNLIYPGIPQQAVGVVLTENEKKTLDELKL 606
>TC93065
Length = 783
Score = 55.5 bits (132), Expect = 3e-08
Identities = 30/93 (32%), Positives = 51/93 (54%)
Frame = +2
Query: 117 LRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCS 176
LRR+FE L MKE ET+ + R + +++ E +S + EKIL + + + S
Sbjct: 215 LRREFEALKMKETETVREFSDRISKVVTQIRLLGEDLSDQRVVEKILVCLPEMFEAKISS 394
Query: 177 IEESNNLDTMTIDELQSSLVHEQRIKSNGEEEN 209
+EE+ N +T+ EL ++L ++ +S EEN
Sbjct: 395 LEENKNFSEITVAELVNALQASEQRRSLRMEEN 493
>TC81230
Length = 958
Score = 30.4 bits (67), Expect = 0.99
Identities = 26/130 (20%), Positives = 54/130 (41%), Gaps = 1/130 (0%)
Frame = +1
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQT-EAQRKLIKEQKLMNLKTKNYLFQ 76
+Y+HWA M FL+ + W + + G + T +A ++E N + +
Sbjct: 250 NYNHWAESMCGFLKGRRLWRYVTGDKKCPTKGKDDTADAFADKLEEWDSKNHQIITWFRN 429
Query: 77 AISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYF 136
+ + K +WD +K + S+ + Q Q L +D L + G+ + +
Sbjct: 430 TSIPSIHMQFGRFENAKEVWDHLKQRYTISD--LSHQYQLL-KDLSNLKQQSGQPVYEFL 600
Query: 137 SRTLTIANKM 146
++ I N++
Sbjct: 601 AQMEVIWNQL 630
>BG648607 weakly similar to PIR|F96614|F96 probable copia-type polyprotein
T18I24.5 [imported] - Arabidopsis thaliana, partial (2%)
Length = 563
Score = 29.6 bits (65), Expect = 1.7
Identities = 16/38 (42%), Positives = 23/38 (60%)
Frame = +2
Query: 158 ITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSL 195
I EKI + + +V +IEE NL TM+I++LQ L
Sbjct: 71 IMEKIFPWLDINFENIVVTIEEIKNL*TMSIEQLQEPL 184
>TC91968 similar to PIR|T02550|T02550 NPK1-related protein kinase homolog
T26B15.7 - Arabidopsis thaliana, partial (20%)
Length = 662
Score = 27.3 bits (59), Expect = 8.4
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = -1
Query: 192 QSSLVHEQRIKSNGEEENGGDNPTT 216
Q SL H QRI+S+ G NP T
Sbjct: 647 QESLAHRQRIQSDTRPRQGQKNPAT 573
>AL369363 weakly similar to GP|10177044|dbj RNA helicase {Arabidopsis
thaliana}, partial (16%)
Length = 504
Score = 27.3 bits (59), Expect = 8.4
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = -3
Query: 72 NYLFQAISHDVLETILNKDTYKNIWDSMKT 101
N LF + D+L+T L KD +N W S+K+
Sbjct: 274 NNLFGT*TSDILKTHLAKDKPRNSWRSIKS 185
>TC93152
Length = 647
Score = 27.3 bits (59), Expect = 8.4
Identities = 12/42 (28%), Positives = 21/42 (49%), Gaps = 3/42 (7%)
Frame = +3
Query: 1 ENDSSNFVHSII---PKFDGHYDHWAMFMENFLRSKEYWDLI 39
E SSN + I P +YD+W+ + N+L ++ W +
Sbjct: 102 EGTSSNISGNAIVFEPLTKDNYDYWSCLVRNYLLGQDLWGFV 227
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.334 0.144 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,191,932
Number of Sequences: 36976
Number of extensions: 104246
Number of successful extensions: 878
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 870
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 877
length of query: 330
length of database: 9,014,727
effective HSP length: 96
effective length of query: 234
effective length of database: 5,465,031
effective search space: 1278817254
effective search space used: 1278817254
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC121241.10


