

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121239.4 - phase: 0
(442 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
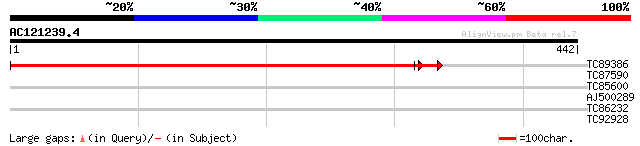
TC89386 similar to GP|6779209|emb|CAB70464.1 BETA-1 2-N-ACETYLGL... 649 0.0
TC87590 weakly similar to PIR|G96522|G96522 F11A17.16 [imported]... 32 0.37
TC85600 homologue to SP|P29828|PDI_MEDSA Protein disulfide isome... 30 1.4
AJ500289 29 4.1
TC86232 similar to GP|21593313|gb|AAM65262.1 protein disulfide i... 28 5.4
TC92928 similar to GP|2160170|gb|AAB60733.1| F21M12.19 gene prod... 28 9.2
>TC89386 similar to GP|6779209|emb|CAB70464.1 BETA-1
2-N-ACETYLGLUCOSAMINYLTRANSFERASE I {Nicotiana tabacum},
partial (72%)
Length = 1392
Score = 649 bits (1673), Expect = 0.0
Identities = 318/322 (98%), Positives = 318/322 (98%)
Frame = +3
Query: 1 MAKFWCDFRFLLFVAALVFIYIQMRLFASQSQYADRLAAAIEAENHCTAQIRSLIDQISL 60
MAKFWCDFRFLLFVAALVFIYIQMRLFASQSQYADRLAAAIEAENHCTAQIRSLIDQISL
Sbjct: 3 MAKFWCDFRFLLFVAALVFIYIQMRLFASQSQYADRLAAAIEAENHCTAQIRSLIDQISL 182
Query: 61 QQGRILDLQQERNRREQECSQIKSLVQDLERKDVRRLIDKVQVPVAAVVIMACNRADYLE 120
QQGRILDLQQERNRREQECSQIKSLVQDLERKDVRRLIDKVQVPVAAVVIMACNRADYLE
Sbjct: 183 QQGRILDLQQERNRREQECSQIKSLVQDLERKDVRRLIDKVQVPVAAVVIMACNRADYLE 362
Query: 121 RTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELI 180
RTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELI
Sbjct: 363 RTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELI 542
Query: 181 AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSS 240
AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSS
Sbjct: 543 AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSS 722
Query: 241 WNDNGQKQFVHDPYELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQ 300
WNDNGQKQFVHDPYELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQ
Sbjct: 723 WNDNGQKQFVHDPYELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQ 902
Query: 301 FIRPEVCRTYNFGEHGSSLGQF 322
FIRPEVCRTYNFGEH S L F
Sbjct: 903 FIRPEVCRTYNFGEHVSFLLYF 968
Score = 47.4 bits (111), Expect = 1e-05
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = +2
Query: 316 GSSLGQFFKQYLEPIKLNEVQV 337
GSSLGQFFKQYLEPIKLNEVQV
Sbjct: 1235 GSSLGQFFKQYLEPIKLNEVQV 1300
>TC87590 weakly similar to PIR|G96522|G96522 F11A17.16 [imported] -
Arabidopsis thaliana, partial (45%)
Length = 2425
Score = 32.3 bits (72), Expect = 0.37
Identities = 15/55 (27%), Positives = 32/55 (57%)
Frame = +2
Query: 41 IEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQIKSLVQDLERKDVR 95
IE ++ + + L++++ + + I LQ E N + E +Q+K L DLE ++++
Sbjct: 641 IEEDDPDVKEKKELLEKLEVSENLIKSLQSEINALKDELNQVKGLNIDLESQNIK 805
>TC85600 homologue to SP|P29828|PDI_MEDSA Protein disulfide isomerase
precursor (PDI) (EC 5.3.4.1). [Alfalfa] {Medicago
sativa}, complete
Length = 1915
Score = 30.4 bits (67), Expect = 1.4
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = -2
Query: 260 DFFPGLGWMLARSTWDELSPKWPKAYWDD 288
+F P L ++ R TW SPK+ KA W D
Sbjct: 1068 NFLPSLC*IIIRGTWSSFSPKYWKAPWLD 982
>AJ500289
Length = 484
Score = 28.9 bits (63), Expect = 4.1
Identities = 21/60 (35%), Positives = 29/60 (48%), Gaps = 1/60 (1%)
Frame = -1
Query: 221 YFEAMATLLDKDKSIMAVSSWND-NGQKQFVHDPYELYRSDFFPGLGWMLARSTWDELSP 279
YF A + K KS + S G+K FV L + + GLGW + +T D+LSP
Sbjct: 259 YFSAKEFYVTKGKSSIRYSRRKGIKGKKPFVAP---LPKGRLYRGLGWKPSLATVDDLSP 89
>TC86232 similar to GP|21593313|gb|AAM65262.1 protein disulfide isomerase
precursor-like {Arabidopsis thaliana}, partial (50%)
Length = 1315
Score = 28.5 bits (62), Expect = 5.4
Identities = 19/80 (23%), Positives = 36/80 (44%), Gaps = 3/80 (3%)
Frame = +3
Query: 113 CNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKR-KALSYDELSHMQ--HLDFE 169
C +LE N + K+ R I S + DG+ ++ R K+ + L + F+
Sbjct: 636 CGHCQHLEPIYNKLAKHLRSIDSL--VIAKMDGTQNEHPRAKSDGFPTLLFFPAGNKSFD 809
Query: 170 PVQTERPGELIAYYKIARHY 189
P+ E ++A+YK + +
Sbjct: 810 PITVETDRTVVAFYKFLKQH 869
>TC92928 similar to GP|2160170|gb|AAB60733.1| F21M12.19 gene product
{Arabidopsis thaliana}, partial (39%)
Length = 643
Score = 27.7 bits (60), Expect = 9.2
Identities = 29/129 (22%), Positives = 51/129 (39%), Gaps = 8/129 (6%)
Frame = +2
Query: 60 LQQGRILDLQQERNRREQECS-------QIKSLVQDLERKDVRRLIDKVQVPVAAVVIMA 112
L+ + L + +ECS +I + +++ + R I + V V +
Sbjct: 17 LESATMTQLNPKHTVEFEECSDNNNRPSKISKIEEEMNQHHHHRKIQRYLVAVEYIGTRF 196
Query: 113 CNRADYL-ERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPV 171
L +RT+ VL+ F FV Q S R LS++ H+D E +
Sbjct: 197 FGFQKQLTDRTVAGVLE------DAFCKFVGQPVSVVCSSRTDAGVHALSNVCHVDIERI 358
Query: 172 QTERPGELI 180
+PGE++
Sbjct: 359 SKRKPGEVL 385
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,383,757
Number of Sequences: 36976
Number of extensions: 180892
Number of successful extensions: 798
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 798
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 798
length of query: 442
length of database: 9,014,727
effective HSP length: 99
effective length of query: 343
effective length of database: 5,354,103
effective search space: 1836457329
effective search space used: 1836457329
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121239.4


