

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121234.10 + phase: 0
(526 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
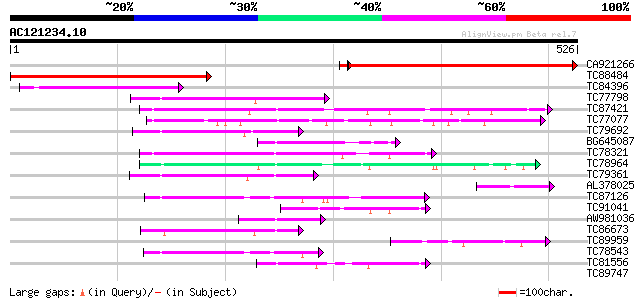
Score E
Sequences producing significant alignments: (bits) Value
CA921266 weakly similar to PIR|T01019|T010 transport protein hom... 406 e-113
TC88484 weakly similar to PIR|T01019|T01019 transport protein ho... 384 e-107
TC84396 similar to GP|9294345|dbj|BAB02242.1 organic anion trans... 96 4e-20
TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 69 6e-12
TC87421 sugar transporter 63 2e-10
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 63 3e-10
TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 59 3e-09
BG645087 similar to GP|17381265|gb AT5g18840/F17K4_90 {Arabidops... 57 2e-08
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 55 7e-08
TC78964 weakly similar to PIR|T00450|T00450 probable monosacchar... 54 2e-07
TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 48 8e-06
AL378025 similar to GP|9294345|dbj| organic anion transporter-li... 48 1e-05
TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {... 46 4e-05
TC91041 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 45 9e-05
AW981036 weakly similar to PIR|G84564|G84 probable sugar transpo... 44 1e-04
TC86673 weakly similar to PIR|S25015|S25015 monosaccharide trans... 44 1e-04
TC89959 similar to GP|15795137|dbj|BAB02515. transporter-like pr... 44 2e-04
TC78543 similar to PIR|T14545|T14545 probable sugar transporter ... 43 3e-04
TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar tran... 42 6e-04
TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported]... 36 0.032
>CA921266 weakly similar to PIR|T01019|T010 transport protein homolog
YUP8H12R.2 - Arabidopsis thaliana, partial (17%)
Length = 721
Score = 406 bits (1043), Expect(2) = e-113
Identities = 202/212 (95%), Positives = 207/212 (97%)
Frame = -3
Query: 315 LFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL 374
+FHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL
Sbjct: 698 IFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL 519
Query: 375 ENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELF 434
ENLRRKPSILVFSILGGICCV CAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELF
Sbjct: 518 ENLRRKPSILVFSILGGICCVMCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELF 339
Query: 435 PTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIG 494
PTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPET+G
Sbjct: 338 PTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETMG 159
Query: 495 IVLCDTMDQQEKKEIALCDAMNQQVRTENVSV 526
IVLCDTMDQQEKKEIA+ DAMNQ ++ NVSV
Sbjct: 158 IVLCDTMDQQEKKEIAMSDAMNQDEKSRNVSV 63
Score = 22.7 bits (47), Expect(2) = e-113
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -1
Query: 307 QLYSSIGELFHK 318
QLYSSIGE F K
Sbjct: 721 QLYSSIGEFFIK 686
>TC88484 weakly similar to PIR|T01019|T01019 transport protein homolog
YUP8H12R.2 - Arabidopsis thaliana, partial (20%)
Length = 619
Score = 384 bits (985), Expect = e-107
Identities = 185/187 (98%), Positives = 186/187 (98%)
Frame = +1
Query: 1 MEGSVQINVPESHSNIEQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSF 60
MEGSVQINVPESHSNIEQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSF
Sbjct: 58 MEGSVQINVPESHSNIEQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSF 237
Query: 61 ISIYTDNYPKWHCTNSICTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQ 120
ISIYTDNYPKWHCTNSICTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQ
Sbjct: 238 ISIYTDNYPKWHCTNSICTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQ 417
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSA+KFLIGF
Sbjct: 418 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSAMKFLIGF 597
Query: 181 WRSSIGT 187
WR SIGT
Sbjct: 598 WRXSIGT 618
>TC84396 similar to GP|9294345|dbj|BAB02242.1 organic anion transporter-like
protein {Arabidopsis thaliana}, partial (22%)
Length = 564
Score = 95.5 bits (236), Expect = 4e-20
Identities = 50/153 (32%), Positives = 77/153 (49%), Gaps = 1/153 (0%)
Frame = +1
Query: 10 PESHSNIEQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYP 69
P S ++ E EK I D ++EK FG +L ++A +A + + I+ D P
Sbjct: 97 PSSKTSNETEKICI---DEMLEKYCGEFGKWQLKHFILTSLAWALEAFHTMVMIFADREP 267
Query: 70 KWHCTNSI-CTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCL 128
W C + C+++ +C + SSW W + +S W+L C F GL Q+ FF GC+
Sbjct: 268 DWKCVEGMECSANGSVCDMASSSWQWVAGEGASTVSEWSLICGDKFKIGLVQALFFAGCM 447
Query: 129 LGSFFLAALADSSIGRKNMLIFSCVSMSITSML 161
+G+ L+DSS+GRK LI C +I L
Sbjct: 448 IGAGVFGHLSDSSLGRKGSLIVVCALNTIFGCL 546
>TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (88%)
Length = 2467
Score = 68.6 bits (166), Expect = 6e-12
Identities = 47/187 (25%), Positives = 93/187 (49%), Gaps = 3/187 (1%)
Frame = +3
Query: 113 TFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYS 172
T + GL + IG + + ++D +GR+ M+I S V + S+++++S NV++
Sbjct: 228 TTMEGLVVAMSLIGATVITTCSGPISDW-LGRRPMMIISSVLYFLGSLVMLWSPNVYVLC 404
Query: 173 ALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYI---NRN 229
+ L GF T V V ++E ++ R + + F+ + G ++ + +
Sbjct: 405 LARLLDGFGIGLAVTLVPVYISETAPSDIRGSLNTLPQFSGSGGMFLSYCMVFVMSLSPS 584
Query: 230 SSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVN 289
SW+ + S+P++ Y ++ F+ ESPRWLV +G+ E K+L+R+ ++ +
Sbjct: 585 PSWRIMLGVLSIPSLFYFLLTVFFLPESPRWLVSKGKMLEAKKVLQRLRGQDDVSGEMAL 764
Query: 290 LASNLPI 296
L L I
Sbjct: 765 LVEGLGI 785
>TC87421 sugar transporter
Length = 1753
Score = 63.2 bits (152), Expect = 2e-10
Identities = 88/417 (21%), Positives = 173/417 (41%), Gaps = 34/417 (8%)
Frame = +2
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SS ++ LL S +A+ GRK ++F + + +++ F+ +VW+ + L+GF
Sbjct: 287 SSLYLAALLSSL-VASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGF 463
Query: 181 WRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLP----GFAYINRNSSWKSLY 236
V + L+E ++R + I + T+G + FA I W+
Sbjct: 464 GIGFANQPVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSL 643
Query: 237 IWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPI 296
+ VPA+ + I L + ++P ++ +G LKR+ E D++ +L +
Sbjct: 644 GGAMVPALIIT-IGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVA---- 808
Query: 297 LPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMI----LGIGLGMVYFGMPLAVGNLGF- 351
+ Q+ + L +++ +AV+I G+ ++ F P+ ++GF
Sbjct: 809 ----ASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFK 976
Query: 352 ---NIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAK 408
++ AV+ + +CV+ Y ++ R+ L GG + C V AK
Sbjct: 977 DDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLE----GGAQMLICQVAVAAAIGAK 1144
Query: 409 ----------------VVLAMVAFFGACTAYN---VFLIYIIELFPTCVRNTTTSL---V 446
VV+ + + A A++ + + E+FP +R+ S+ V
Sbjct: 1145FGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSV 1324
Query: 447 RQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMDQ 503
F ++ K ++ + F V++M S + F LPET GI + + MD+
Sbjct: 1325NMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVM-SIYVFFLLPETKGIPI-EEMDR 1489
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 62.8 bits (151), Expect = 3e-10
Identities = 101/417 (24%), Positives = 174/417 (41%), Gaps = 47/417 (11%)
Frame = +1
Query: 128 LLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGT 187
L+GS LA IGR+ ++F+ + ++L+ FS N +F+ G IG
Sbjct: 262 LIGSC-LAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYNFLMFGRFVAGV---GIGY 429
Query: 188 CVLV--LLTEKVS--AEWRFRVGIVEYFT---FTMGYMSLPGFAYINRNSSWKSLYIWSS 240
+++ + T +VS + F E F +GY+S F+ ++ W+ + +
Sbjct: 430 ALMIAPVYTAEVSPASSRGFLTSFPEVFINSGILLGYISNYAFSKLSLKLGWRMMLGVGA 609
Query: 241 VPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLAS-------- 292
+P++ +V L + ESPRWLVM+GR + +K+L + S +S ++ + LA
Sbjct: 610 IPSVILAV-GVLAMPESPRWLVMRGRLGDAIKVLNKTS--DSKEEAQLRLAEIKQAAGIP 780
Query: 293 ---NLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGI-------GLGMVYFGM 342
N ++ K K + ++ ELF +R I + LGI G+ V
Sbjct: 781 EDCNDDVVEVKVKNTGEGVWK---ELFLYPTPAVRHIVIAALGIHFFQQASGVDAVVLYS 951
Query: 343 PLAVGNLGFN-----IYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGG--ICCV 395
P G N + + L VAT+ L+ R+P L+ + +GG + +
Sbjct: 952 PTIFKKAGINGDTHLLIATIAVGFVKTLFILVATFMLDRYGRRP--LLLTSVGGMVVSLL 1125
Query: 396 TCAVLENRVP------------AAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVR--NT 441
T AV + + VL+ VA F + A + +Y E+FP +R
Sbjct: 1126TLAVSLTIIDHSNTKLNWAIGLSIATVLSYVATF-SIGAGPITWVYSSEIFPLRLRAQGA 1302
Query: 442 TTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLS-NFTLFFLPETIGIVL 497
+V + G I FL + ++ +FG + + F LPET G L
Sbjct: 1303ACGVVVNRVTSGVISMTFLSLSKGITIGGAFFLFGGIATIGWIFFYIMLPETQGKTL 1473
>TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (49%)
Length = 1353
Score = 59.3 bits (142), Expect = 3e-09
Identities = 46/162 (28%), Positives = 79/162 (48%), Gaps = 4/162 (2%)
Frame = +2
Query: 115 ITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSAL 174
+ GL + IG + + AL+D GR+ MLI S + ++S+++ +S NV+I
Sbjct: 281 VEGLIVAMSLIGATVVTTCSGALSDL-FGRRPMLIISSLLYFLSSLVMFWSPNVYILLFA 457
Query: 175 KFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMG----YMSLPGFAYINRNS 230
+ L G T V + ++E E R + + F + G Y + G + + +
Sbjct: 458 RLLDGLGIGLAVTLVPLYISEIAPPEIRGSLNTLPQFAGSAGMFFSYCMVFGMS-LTKAP 634
Query: 231 SWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILK 272
SW+ + S+P++ Y + L + ESPRWLV +GR E K
Sbjct: 635 SWRLMLGVLSIPSLIYFALTLLVLPESPRWLVSKGRMLEAKK 760
>BG645087 similar to GP|17381265|gb AT5g18840/F17K4_90 {Arabidopsis
thaliana}, partial (46%)
Length = 805
Score = 56.6 bits (135), Expect = 2e-08
Identities = 37/133 (27%), Positives = 63/133 (46%), Gaps = 1/133 (0%)
Frame = +2
Query: 231 SWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEE-SADDDSVN 289
+W+ L + VP IC ++ F+ ESPRWL GREKE L+++ ++ D++
Sbjct: 74 NWRQLALAGLVPCICL-LVGLCFIPESPRWLAKVGREKEFQLALRKLRGKDIDISDEANE 250
Query: 290 LASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNL 349
+ N+ L K F L+ S + + +I+G+GL + F + + +
Sbjct: 251 ILDNIETLQSLPKTKFLDLFQS------------KYVRSVIIGVGL--MAFQQSVGINGI 388
Query: 350 GFNIYLAVVFSAS 362
GF Y A F A+
Sbjct: 389 GF--YTAETFVAA 421
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 55.1 bits (131), Expect = 7e-08
Identities = 62/294 (21%), Positives = 125/294 (42%), Gaps = 18/294 (6%)
Frame = +2
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
S+ G ++G+ + + GRK +I + + S+++ + N + +G
Sbjct: 362 STAIAGAIIGAA-IGGWINDRFGRKVSIIVADTLFLLGSIILAAAPNPATLIVGRVFVGL 538
Query: 181 WRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMG-YMS-LPGFAYINRNSSWKSLYIW 238
+ ++E R + + F T G ++S L A+ +W+ +
Sbjct: 539 GVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTKAPGTWRWMLGV 718
Query: 239 SSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPI-L 297
++ PA+ ++ L + ESPRWL +G+E+E +LK++ E D++ L ++ + L
Sbjct: 719 AAAPAVI-QIVLMLSLPESPRWLYRKGKEEEAKVILKKIYEVEDYDNEIQALKESVEMEL 895
Query: 298 PPKEKVSFFQ----------LYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVG 347
EK+S Q LY+ +G F +++ G+ V + P V
Sbjct: 896 KETEKISIMQLVKTTSVRRGLYAGVGLAFFQQFT------------GINTVMYYSPSIVQ 1039
Query: 348 NLGF-----NIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGGICCVT 396
GF + L+++ S S ++ YF++ RK L+ S+ G + +T
Sbjct: 1040LAGFASKRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALI-SLTGVVLSLT 1198
>TC78964 weakly similar to PIR|T00450|T00450 probable monosaccharide
transport protein T14N5.7 - Arabidopsis thaliana,
partial (98%)
Length = 1771
Score = 53.5 bits (127), Expect = 2e-07
Identities = 97/407 (23%), Positives = 156/407 (37%), Gaps = 35/407 (8%)
Frame = +1
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SS + L+ +FF + L + GRK +I +S I ++L + N+ + +G
Sbjct: 298 SSLYFSALVMTFFASYLTRNK-GRKATIIVGALSFLIGAILNAAAQNIPTLIIGRVFLGG 474
Query: 181 WRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRN---SSWKSLYI 237
V + L+E A R V + FT G + Y W+
Sbjct: 475 GIGFGNQAVPLYLSEMAPASSRGAVNQLFQFTTCAGILIANLVNYFTDKIHPHGWRISLG 654
Query: 238 WSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPIL 297
+ +PA+ ++ +F E+P LV QGR E K+L++V ++ D + +L
Sbjct: 655 LAGIPAVLM-LLGGIFCAETPNSLVEQGRLDEARKVLEKVRGTKNVDAEFEDL------- 810
Query: 298 PPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILG-------IGLGMVYFGMPLAVGNLG 350
K+ Q S ++ KR ++I + LG G + F P+ + G
Sbjct: 811 --KDASELAQAVKSPFKVLLKRKYRPQLI-IGALGTPAFQQLTGNNSILFYAPVIFQSSG 981
Query: 351 FNIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGG---ICC--VTCAVLENRVP 405
F A+ S VAT L K F + G ICC +T VL
Sbjct: 982 FGSNAALFSSFITNGALLVATVISMFLVDKFGTRKFFLEAGFEMICCMIITAVVLAVEFG 1161
Query: 406 AAKVVLAMVAFFGACTAYNVFLIY-----------IIELFPTCVRNTTTSLVRQAIVFGC 454
K + ++ F + L Y ELFP +R+ S+ A+
Sbjct: 1162HGKELSKGISAFLVIMIFWFVLAYGRSWGPLGWLVPSELFPLEIRSAAQSI---AVCVNM 1332
Query: 455 IFCP-----FLISAGRKNNIYSYGVF----GVVIMLSNFTLFFLPET 492
IF FL+S YG+F G+++++S F F LPET
Sbjct: 1333IFTALVAQLFLLSLCH----LKYGIFLLFGGLIVVMSVFVFFLLPET 1461
>TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (55%)
Length = 949
Score = 48.1 bits (113), Expect = 8e-06
Identities = 40/177 (22%), Positives = 76/177 (42%), Gaps = 2/177 (1%)
Frame = +1
Query: 112 STFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIY 171
S F+ S G ++G+ F L D+ GRK + + V + ++L+ + + ++
Sbjct: 283 SNFLQETIVSMAIAGAIVGAAFGGWLNDA-YGRKKATLLADVIFILGAILMAAAPDPYVL 459
Query: 172 SALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMS--LPGFAYINRN 229
A + L+G V + E +E R + T G L +
Sbjct: 460 IAGRLLVGLGVGIASVTAPVYIAEVAPSEIRGSLVSTNVLMITGGQFVSYLVNLVFTQVP 639
Query: 230 SSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDD 286
+W+ + S VPA+ I LF+ ESPRWL ++ R+ E + ++ ++ +D+
Sbjct: 640 GTWRWMLGVSGVPALI-QFICMLFLPESPRWLFIKNRKNEAVDVISKIYDLSRLEDE 807
>AL378025 similar to GP|9294345|dbj| organic anion transporter-like protein
{Arabidopsis thaliana}, partial (12%)
Length = 443
Score = 47.8 bits (112), Expect = 1e-05
Identities = 27/72 (37%), Positives = 36/72 (49%)
Frame = +2
Query: 434 FPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETI 493
FPT VRN QA G I PF++ G + +GVF V +L F+LPET+
Sbjct: 2 FPTVVRNAALGSATQAAQMGAILAPFVVVLG---SWLPFGVFAVCGILGGMFAFYLPETL 172
Query: 494 GIVLCDTMDQQE 505
+ L DT + E
Sbjct: 173 NMPLYDTFNGLE 208
>TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {Arabidopsis
thaliana}, partial (79%)
Length = 1524
Score = 45.8 bits (107), Expect = 4e-05
Identities = 65/276 (23%), Positives = 115/276 (41%), Gaps = 12/276 (4%)
Frame = +3
Query: 126 GCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSI 185
G ++G+ +A+ +GRK L+ + + I + I F+ + + L GF I
Sbjct: 6 GAMVGAIASGQIAEY-VGRKGSLMIASIPNIIGWLAISFAKDSSFLFMGRLLEGFGVGII 182
Query: 186 GTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYI-NRNSSWKSLYIWSSVPAI 244
V V + E R +G V + T+G M AY+ ++W+ L I +P
Sbjct: 183 SYVVPVYIAEIAPENMRGSLGSVNQLSVTIGIM----LAYLLGLFANWRVLAILGILP-- 344
Query: 245 CYSVIAYLF-VTESPRWLVMQGREKEI---LKMLKRVSSEESADDDSVN--LASN----- 293
C +I LF + ESPRWL G +E L++L+ ++ S + + +ASN
Sbjct: 345 CTVLIPGLFFIPESPRWLAKMGMMEEFETSLQVLRGFDTDISVEVHEIKKAVASNGKRAT 524
Query: 294 LPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNI 353
+ + K +F L IG L + + GI + Y A + +
Sbjct: 525 IRFADLQRKRYWFPLSVGIGLL----------VLQQLSGINGVLFYSTSIFANAGISSSN 674
Query: 354 YLAVVFSASMELPSCVATYFLENLRRKPSILVFSIL 389
V A + + VAT+ ++ R+ +++ S L
Sbjct: 675 AATVGLGAIQVIATGVATWLVDKSGRRVLLIISSSL 782
>TC91041 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 564
Score = 44.7 bits (104), Expect = 9e-05
Identities = 42/149 (28%), Positives = 72/149 (48%), Gaps = 10/149 (6%)
Frame = +3
Query: 252 LFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSS 311
+ + ESPRWL +G+E+E +L+R+ S E A+ + +N L KE S + +S
Sbjct: 18 IMLPESPRWLFRKGKEEEAKALLRRIYSPEEAEAE-INTLKESVELEIKESESSDK--AS 188
Query: 312 IGELFHKRWAVIRMIAVMILGI-----GLGMVYFGMPLAVGNLGF-----NIYLAVVFSA 361
I ++ + + A M L I G+ V + P + GF + L++V S
Sbjct: 189 IIKMLKTKTVRRGLYAGMGLQIFQQFVGINTVMYYSPAIIQLAGFASNQTALLLSLVTSG 368
Query: 362 SMELPSCVATYFLENLRRKPSILVFSILG 390
S ++ YF++ RK +L+FS+ G
Sbjct: 369 LNAFGSILSIYFIDKTGRK-KLLLFSLSG 452
>AW981036 weakly similar to PIR|G84564|G84 probable sugar transporter
[imported] - Arabidopsis thaliana, partial (25%)
Length = 575
Score = 44.3 bits (103), Expect = 1e-04
Identities = 26/81 (32%), Positives = 42/81 (51%)
Frame = +2
Query: 213 FTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILK 272
F +GY+S ++ W+ + S+P+I VI L + ESPRWLVMQGR + K
Sbjct: 140 FLLGYLSNYFLGKLSLRLGWRIMLAIPSIPSIGL-VILMLQLVESPRWLVMQGRLGDAKK 316
Query: 273 MLKRVSSEESADDDSVNLASN 293
+L +S+ + + + N
Sbjct: 317 VLLLISNSKQEAEQRMKEIKN 379
>TC86673 weakly similar to PIR|S25015|S25015 monosaccharide transport
protein MST1 - common tobacco, partial (65%)
Length = 1162
Score = 44.3 bits (103), Expect = 1e-04
Identities = 39/159 (24%), Positives = 70/159 (43%), Gaps = 8/159 (5%)
Frame = +2
Query: 122 SFFIGCLLGSFFLAALADSSI----GRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFL 177
+ F L S +A L SSI GR+ +I + ++L + N+ + A + L
Sbjct: 278 TLFTSSLYISALVAGLGASSITRALGRRTTMILGGIFFVSGALLNGLAMNIAMLIAGRLL 457
Query: 178 IGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAY----INRNSSWK 233
+GF V + L+E ++R + + + T+G + F Y I W+
Sbjct: 458 LGFGIGCANQAVPIYLSEMAPYKYRGALNMCFQLSITIGIFTANMFNYYFSKILNGEGWR 637
Query: 234 SLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILK 272
+VPA+ + +I + + +SP LV +GR +E K
Sbjct: 638 LSLGLGAVPAVVF-IIGSICLPDSPNSLVTRGRHEEARK 751
>TC89959 similar to GP|15795137|dbj|BAB02515. transporter-like protein
{Arabidopsis thaliana}, partial (87%)
Length = 1681
Score = 43.5 bits (101), Expect = 2e-04
Identities = 42/157 (26%), Positives = 72/157 (45%), Gaps = 9/157 (5%)
Frame = +1
Query: 354 YLAVVFSASMELPSCV-ATYFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLA 412
Y V ++ ELP + + ++ L RK S+ SI+ +CC+ + +P L
Sbjct: 1126 YKGVFIASFAELPGLLLSAVAVDKLGRKLSM---SIMFFMCCIFLLPVTFYLPED---LT 1287
Query: 413 MVAFFGA----CTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNN 468
FGA + + IY E++PT VR T + G + CP L++ G +
Sbjct: 1288 TGLLFGARICITVTFTIVYIYAPEIYPTSVRTTGVGVASSVGRIGGMLCP-LVAVGLVHG 1464
Query: 469 IYSYG---VFGVVIMLSNFTLFFLP-ETIGIVLCDTM 501
+ +F +V ++S + F+P ET+G L DT+
Sbjct: 1465 CHQTAAVLLFIIVALVSGICVVFIPFETMGQELQDTV 1575
>TC78543 similar to PIR|T14545|T14545 probable sugar transporter protein -
beet, partial (93%)
Length = 1653
Score = 43.1 bits (100), Expect = 3e-04
Identities = 44/172 (25%), Positives = 77/172 (44%), Gaps = 5/172 (2%)
Frame = +1
Query: 125 IGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSS 184
+G ++G+ +A+ +GRK L+ + + I ++I F+ + + L GF
Sbjct: 292 VGAMVGAIASGQIAEY-MGRKGSLMIASIPNIIGWLMISFANDSSFLYMGRLLEGFGVGI 468
Query: 185 IGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYI-NRNSSWKSLYIWSSVPA 243
I V V + E R + V + T+G M AY+ W+ L I +P
Sbjct: 469 ISYTVPVYIAEISPQNLRGSLVSVNQLSVTLGIM----LAYLLGLFVEWRFLAILGIIP- 633
Query: 244 ICYSVIAYLF-VTESPRWLVMQGREKEI---LKMLKRVSSEESADDDSVNLA 291
C +I LF + ESPRWL G +E L++L+ ++ S + + + A
Sbjct: 634 -CTLLIPGLFFIPESPRWLAKMGMTEEFENSLQVLRGFETDISVEVNEIKTA 786
>TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar transporter
{Prunus armeniaca}, partial (59%)
Length = 1074
Score = 42.0 bits (97), Expect = 6e-04
Identities = 40/171 (23%), Positives = 77/171 (44%), Gaps = 10/171 (5%)
Frame = +1
Query: 230 SSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESA-----D 284
S+W++++ + VP+I + + ESPRWL QG+ E K +K + +E D
Sbjct: 13 SAWRTMFGIAVVPSILLA-LGMAISPESPRWLFQQGKIAEAEKAIKTLYGKERVATVMYD 189
Query: 285 DDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMIL-----GIGLGMVY 339
+ + S+ P + +F L+SS + W V+ + A + L GI + Y
Sbjct: 190 LRTASQGSSEP------EAGWFDLFSS------RYWKVVSVGAALFLFQQLAGINAVVYY 333
Query: 340 FGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILG 390
+ ++ + + AS + + +A+ ++ RK S+L+ S G
Sbjct: 334 STSVFRSAGIASDVAASALVGASNVIGTAIASSLMDKQGRK-SLLITSFSG 483
>TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1372
Score = 36.2 bits (82), Expect = 0.032
Identities = 38/159 (23%), Positives = 58/159 (35%), Gaps = 1/159 (0%)
Frame = +3
Query: 112 STFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIY 171
+T GL S G L G +AD+ +GR+ + M I + + + N++
Sbjct: 483 NTLAEGLVVSICLGGALFGCLLSGWIADA-VGRRRAFQLCALPMIIGAAMSAATNNLFGM 659
Query: 172 SALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYM-SLPGFAYINRNS 230
+ +G + +TE A R G + G + SL + S
Sbjct: 660 LVGRLFVGTGLGLGPPVAALYVTEVSPAFVRGTYGALIQIATCFGILGSLFIGIPVKEIS 839
Query: 231 SWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKE 269
W + W S +A F ESP WL QGR E
Sbjct: 840 GWWRVCFWVSTIPAAILALAMDFCAESPHWLYKQGRTAE 956
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.138 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,574,797
Number of Sequences: 36976
Number of extensions: 329278
Number of successful extensions: 2609
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 2577
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2604
length of query: 526
length of database: 9,014,727
effective HSP length: 101
effective length of query: 425
effective length of database: 5,280,151
effective search space: 2244064175
effective search space used: 2244064175
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC121234.10


