

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.5 - phase: 0
(301 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
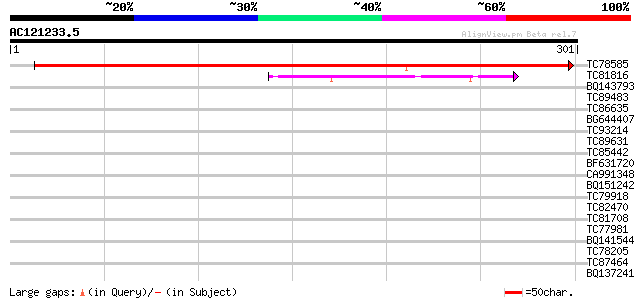
Score E
Sequences producing significant alignments: (bits) Value
TC78585 similar to SP|Q9LUJ5|EBP2_ARATH Probable rRNA processing... 487 e-138
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 41 6e-04
BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 35 0.027
TC89483 similar to GP|18377662|gb|AAL66981.1 unknown protein {Ar... 34 0.060
TC86635 weakly similar to GP|9758600|dbj|BAB09233.1 gene_id:MDF2... 34 0.079
BG644407 similar to GP|12697612|dbj contains ESTs C97999(C0353) ... 33 0.18
TC93214 similar to GP|10177918|dbj|BAB11329. kinesin-like protei... 32 0.30
TC89631 homologue to GP|10177250|dbj|BAB10718. gene_id:K19P17.4~... 32 0.30
TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 32 0.39
BF631720 weakly similar to GP|10798648|em putative DNAJ protein ... 31 0.51
CA991348 similar to PIR|D84685|D84 hypothetical protein At2g2848... 31 0.67
BQ151242 similar to PIR|G86292|G86 hypothetical protein AAF82153... 31 0.67
TC79918 similar to PIR|T06671|T06671 hypothetical protein T17F15... 30 0.87
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 30 0.87
TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protei... 30 1.1
TC77981 similar to PIR|T12970|T12970 hypothetical protein T6H20.... 30 1.5
BQ141544 similar to GP|7022293|dbj| unnamed protein product {Hom... 30 1.5
TC78205 homologue to PIR|T47559|T47559 60S ribosomal protein-lik... 30 1.5
TC87464 homologue to GP|7635498|emb|CAB88668.1 histone H2B {Cice... 29 1.9
BQ137241 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 29 1.9
>TC78585 similar to SP|Q9LUJ5|EBP2_ARATH Probable rRNA processing protein
EBP2 homolog. [Mouse-ear cress] {Arabidopsis thaliana},
partial (72%)
Length = 1280
Score = 487 bits (1253), Expect = e-138
Identities = 249/289 (86%), Positives = 262/289 (90%), Gaps = 3/289 (1%)
Frame = +3
Query: 14 MELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWP 73
ME+PLVN D MID E E L EEMSP ES+SEEDVKLAEPSKTAVNNRDA+LDKLGDISWP
Sbjct: 114 MEVPLVNGDPMIDDETEDLDEEMSPFESDSEEDVKLAEPSKTAVNNRDAILDKLGDISWP 293
Query: 74 ENVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYY 133
ENV+W HKLSIDIDQ QEVDVNDDLAREL+FYTQALEGTRQAFEKL+S+GLPFLRPADYY
Sbjct: 294 ENVDWKHKLSIDIDQGQEVDVNDDLARELSFYTQALEGTRQAFEKLESMGLPFLRPADYY 473
Query: 134 AEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIE 193
AEMVK DSHMEKVKSRLL EK+KM EA+ERRKAREAKRLSKEVQSQKLKERAKQKKDDIE
Sbjct: 474 AEMVKGDSHMEKVKSRLLDEKRKMEEADERRKAREAKRLSKEVQSQKLKERAKQKKDDIE 653
Query: 194 SVKKWRKQRQQSGFAD--DGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKG 251
SVKKWRKQRQQSGFAD D DK+L FEDGKVFERSKKKRPGVSPGDRSGGKAKQA GKG
Sbjct: 654 SVKKWRKQRQQSGFADGADAPDKSLSFEDGKVFERSKKKRPGVSPGDRSGGKAKQAMGKG 833
Query: 252 KKP-KKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKR 299
KKP KK D KN+KFGFGG+KG KKQNTADTTNDFGGFSK A G +KR
Sbjct: 834 KKPQKKRDTKNAKFGFGGRKGSKKQNTADTTNDFGGFSKGSAAGNKRKR 980
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 40.8 bits (94), Expect = 6e-04
Identities = 42/148 (28%), Positives = 69/148 (46%), Gaps = 15/148 (10%)
Frame = +3
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREA---KRLSKEVQSQKLKERAKQKKDDIES 194
KTD +++ K + EK+K + EE K E ++ KE + ++ KE+ K+ KD
Sbjct: 216 KTD--VDEGKDKKDKEKKKKEKKEENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDGE 389
Query: 195 VKKWRKQRQQSGFADDGADKALDFEDGKVFERSK-KKRPGVSPGDRSGGK---------- 243
KK +K +++ ++ D+ EDG +++K KK ++ GK
Sbjct: 390 EKKSKKDKEKKKDKNEDDDEG---EDGSKKKKNKDKKEKKKEEDEKEEGKVSVRDIDIEE 560
Query: 244 -AKQAFGKGKKPKKGDVKNSKFGFGGKK 270
AK+ GK KK KK D + K G K
Sbjct: 561 TAKE--GKEKKKKKEDKEEKKKKSGKDK 638
>BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (19%)
Length = 1100
Score = 35.4 bits (80), Expect = 0.027
Identities = 23/65 (35%), Positives = 32/65 (48%)
Frame = -1
Query: 237 GDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGS 296
G R GG + +G+K +KG+ N+K GGK+ KK+ N KG GG
Sbjct: 515 GARGGGGQAKERKRGRKRRKGE--NAKREKGGKERGKKERKKGAGNGIYEGEYKGGGGGG 342
Query: 297 KKRKR 301
KRK+
Sbjct: 341 GKRKK 327
>TC89483 similar to GP|18377662|gb|AAL66981.1 unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 1711
Score = 34.3 bits (77), Expect = 0.060
Identities = 27/105 (25%), Positives = 47/105 (44%), Gaps = 14/105 (13%)
Frame = +1
Query: 140 DSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQ--------------KLKERA 185
+ +++ K R+L ++ E + RK RE KR +EV+ Q K KER
Sbjct: 154 ERELKREKERVLERYEREAERDRIRKEREQKRRIEEVERQFELQLKEWEYREREKEKERQ 333
Query: 186 KQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKK 230
+K+ + + +K RK+ DDG D + + E+ K+
Sbjct: 334 YEKEKEKDRERKRRKEILYDEEDDDG-DSRKRWRXNAIEEKRNKR 465
>TC86635 weakly similar to GP|9758600|dbj|BAB09233.1
gene_id:MDF20.10~pir||T08929~similar to unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1963
Score = 33.9 bits (76), Expect = 0.079
Identities = 21/65 (32%), Positives = 31/65 (47%)
Frame = +2
Query: 225 ERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDF 284
E SKK+R V G G A + G GK+ KK + G++K++T DT ++
Sbjct: 758 ESSKKRRRTVKRGTPRSGGASTSRGSGKRQKKNE---------DSSGVEKKSTTDTEDES 910
Query: 285 GGFSK 289
G K
Sbjct: 911 EGEEK 925
>BG644407 similar to GP|12697612|dbj contains ESTs C97999(C0353)
D15249(C0353)~similar to Arabidopsis thaliana chromosome
2 At2g01100~, partial (21%)
Length = 764
Score = 32.7 bits (73), Expect = 0.18
Identities = 30/105 (28%), Positives = 44/105 (41%), Gaps = 2/105 (1%)
Frame = +2
Query: 154 KQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDI--ESVKKWRKQRQQSGFADDG 211
K+ M EE++K K+ QKL+ A K D E K + +R+ +D G
Sbjct: 221 KKFMQLVEEKKKRALEKKEGHLKWEQKLEASANLKSDAEAGEKTKAAKHRRKSDSGSDSG 400
Query: 212 ADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKK 256
+D +FE K +S KK D K K +KPK+
Sbjct: 401 SDTDSEFEKKKSTRKSHKKHRKHHSSDLVDLDRKDKKSK-RKPKR 532
>TC93214 similar to GP|10177918|dbj|BAB11329. kinesin-like protein
{Arabidopsis thaliana}, partial (22%)
Length = 715
Score = 32.0 bits (71), Expect = 0.30
Identities = 27/102 (26%), Positives = 56/102 (54%), Gaps = 10/102 (9%)
Frame = +3
Query: 140 DSHMEKVKS---RLLAEKQKMVEA----EERRKAREA-KRLSKEVQSQKL-KERAKQK-K 189
D+H +K+K+ ++L K+K ++++K+ EA KRL E+QS K K + +QK K
Sbjct: 150 DTHSQKLKALEAQILDLKKKQESQVQLMKQKQKSDEATKRLQDEIQSIKAQKVQLQQKIK 329
Query: 190 DDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
+ E ++W+ R++ + ++E K+ ++++R
Sbjct: 330 QEAEQFRQWKASREKELLQLRKEGRKSEYEKHKLQALNQRQR 455
>TC89631 homologue to GP|10177250|dbj|BAB10718. gene_id:K19P17.4~unknown
protein {Arabidopsis thaliana}, partial (36%)
Length = 647
Score = 32.0 bits (71), Expect = 0.30
Identities = 15/39 (38%), Positives = 24/39 (61%)
Frame = +1
Query: 152 AEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKD 190
A K K+ +E +KA+E K KEV+ +E+ ++KKD
Sbjct: 187 AGKAKVAGVKEVKKAKEVKEAPKEVKEAAKEEKIEEKKD 303
>TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (62%)
Length = 585
Score = 31.6 bits (70), Expect = 0.39
Identities = 22/77 (28%), Positives = 37/77 (47%)
Frame = +3
Query: 160 AEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFE 219
A+E K E K+ + + +K KE AK++++ E KK ++++ K E
Sbjct: 54 AKEPEKKEEPKKEEPKKEEEK-KEEAKKEEEKKEGEKKEEPKKEEE-------KKEEKKE 209
Query: 220 DGKVFERSKKKRPGVSP 236
+G+ E KKK P P
Sbjct: 210 EGEKKEEEKKKEPAPDP 260
>BF631720 weakly similar to GP|10798648|em putative DNAJ protein {Nicotiana
tabacum}, partial (25%)
Length = 630
Score = 31.2 bits (69), Expect = 0.51
Identities = 21/81 (25%), Positives = 34/81 (41%), Gaps = 9/81 (11%)
Frame = +3
Query: 229 KKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQ--NTADTTN---- 282
K P +PG++ + + + + + K + G+ GLK N D ++
Sbjct: 141 KYHPDKNPGNKEAEEMFKKISEAYEVLSDEQKRQMYDQYGEDGLKNSGYNPGDASSLFEH 320
Query: 283 ---DFGGFSKKGAVGGSKKRK 300
FGGF G GG +KRK
Sbjct: 321 LFGGFGGFGGFGGFGGGRKRK 383
>CA991348 similar to PIR|D84685|D84 hypothetical protein At2g28480 [imported]
- Arabidopsis thaliana, partial (52%)
Length = 658
Score = 30.8 bits (68), Expect = 0.67
Identities = 30/124 (24%), Positives = 55/124 (44%), Gaps = 9/124 (7%)
Frame = +1
Query: 158 VEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADD------- 210
VEA + K + KRL + + +K K +A ++D K +K++Q+ ++
Sbjct: 85 VEAPRKDKWKTKKRLKMQRKREKEKRKAANRRDPRRLGVKGKKKKQKFANPEERIKFKIN 264
Query: 211 --GADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGG 268
+AL E K +E +K + P V P +G + KK K+S + G
Sbjct: 265 NARVKEALLIERLKRYEVAKAQGPEVKPDGLTGEERHYL-------KKMAQKSSNYLQVG 423
Query: 269 KKGL 272
++G+
Sbjct: 424 RRGI 435
>BQ151242 similar to PIR|G86292|G86 hypothetical protein AAF82153.1
[imported] - Arabidopsis thaliana, partial (5%)
Length = 1371
Score = 30.8 bits (68), Expect = 0.67
Identities = 21/74 (28%), Positives = 34/74 (45%)
Frame = +1
Query: 226 RSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFG 285
++KK++ G G+R G+ ++ GKG + G+ + + KG KK+
Sbjct: 91 KAKKRQGGEQEGERERGRGREREGKGGRGGGGEGRRT-------KGEKKRGGGRGGGREA 249
Query: 286 GFSKKGAVGGSKKR 299
KKG GSK R
Sbjct: 250 QGKKKGGGRGSKTR 291
Score = 29.3 bits (64), Expect = 1.9
Identities = 23/97 (23%), Positives = 43/97 (43%), Gaps = 1/97 (1%)
Frame = +1
Query: 162 ERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWR-KQRQQSGFADDGADKALDFED 220
+RR+ + K+ K++ SQ K + +Q + ++ R ++R+ G G +
Sbjct: 31 KRRELGKKKKKKKKIFSQDAKAKKRQGGEQEGERERGRGREREGKGGRGGGGEGRRT--- 201
Query: 221 GKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
+ +KKR G G R K+ G+G K + G
Sbjct: 202 -----KGEKKRGGGRGGGREAQGKKKGGGRGSKTRAG 297
>TC79918 similar to PIR|T06671|T06671 hypothetical protein T17F15.10 -
Arabidopsis thaliana, partial (39%)
Length = 1269
Score = 30.4 bits (67), Expect = 0.87
Identities = 28/142 (19%), Positives = 62/142 (42%), Gaps = 11/142 (7%)
Frame = +1
Query: 141 SHMEKVKSRLLAEKQKMVEAEERRKA-REAKRLSKEVQSQKLKERAKQKKDDIESVKKWR 199
S ++ ++++L +K+ E+ K R+ + + ++ + R++++ D +S ++ R
Sbjct: 268 SQLQSIQNKL--KKKPTTESRVSEKTNRDYSSRDSDRRKRRSRSRSRERYRDRDSRRRSR 441
Query: 200 KQRQQSGFADDGADKALDFEDGKVFERS----------KKKRPGVSPGDRSGGKAKQAFG 249
+ + S ++D D+ D + G+ RS +K+ G G R
Sbjct: 442 SRERNSRYSDRDRDR--DRDSGRSRSRSDSDVDRGRDRRKRERGREKGRRRSXSVSPRKQ 615
Query: 250 KGKKPKKGDVKNSKFGFGGKKG 271
+ + K DVK G +KG
Sbjct: 616 RRSEVKSSDVKERGKGSEMRKG 681
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 30.4 bits (67), Expect = 0.87
Identities = 45/233 (19%), Positives = 93/233 (39%), Gaps = 27/233 (11%)
Frame = +2
Query: 25 IDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLD----KLGDISWPENVEWIH 80
I + E E++ + E + ++ E K D + K G + E++ H
Sbjct: 203 IKRDIEECCEDLENKKKEIRDVGRIIEARKKMQGKIDECVKDFVAKEGQLGLMEDLIGEH 382
Query: 81 KLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTD 140
K + + + V D+++++ +Q E K + + E+ +
Sbjct: 383 KKELKTKELELRQVMDNISKQKELESQVKELVNDLVSKQKHF-------ESHIKELESKE 541
Query: 141 SHME-KVKSRLLAEKQ-----KMVEAEERRKAREAKRLSKEVQSQK--LKERAKQKKD-- 190
+E ++K L EK+ +E++ER E + ++ ++ K +KE A ++K
Sbjct: 542 RQLEGRLKEHELEEKEFEGRMNELESKERHFKSEVEEINAKLMPLKGQIKELASKEKQLN 721
Query: 191 ----DIESVKKW---------RKQRQQSGFADDGADKALDFEDGKVFERSKKK 230
++ES K K++Q G + A K +FE + ++ KKK
Sbjct: 722 GQVKELESKKNQFENRIKELESKEKQHEGRVKEHASKEREFESQVMEQQFKKK 880
>TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protein {Oryza
sativa}, partial (6%)
Length = 605
Score = 30.0 bits (66), Expect = 1.1
Identities = 32/108 (29%), Positives = 42/108 (38%), Gaps = 4/108 (3%)
Frame = +1
Query: 188 KKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQA 247
KK + KK + ++Q+ G D G +K S KK+ G GG +
Sbjct: 88 KKGGDDGGKKNKGEKQKGGGGDWGDEK-----------NSSKKKNGKGKNG-GGGFLVKF 231
Query: 248 FGKGKKPKKG----DVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKG 291
G GKK KKG D N K GG ++ N G KKG
Sbjct: 232 LGLGKKSKKGGGAADTTNKKKNNGG--------GGNSNNSKGKDGKKG 351
Score = 26.9 bits (58), Expect = 9.6
Identities = 19/61 (31%), Positives = 24/61 (39%)
Frame = +1
Query: 240 SGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKR 299
+GG K GKK K K +G +K K+ N GGF K G K +
Sbjct: 76 NGGGKKGGDDGGKKNKGEKQKGGGGDWGDEKNSSKKKNGKGKNGGGGFLVKFLGLGKKSK 255
Query: 300 K 300
K
Sbjct: 256 K 258
>TC77981 similar to PIR|T12970|T12970 hypothetical protein T6H20.190 -
Arabidopsis thaliana, partial (39%)
Length = 1191
Score = 29.6 bits (65), Expect = 1.5
Identities = 23/74 (31%), Positives = 38/74 (51%), Gaps = 3/74 (4%)
Frame = +2
Query: 127 LRPADYYAEMVKTDSHME---KVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKE 183
LRP D + D + ++ + AE++ VEAE KAREA +K+++ + LK
Sbjct: 446 LRPVDELFSTIPEDGRRKAYAEILEKAKAEEEARVEAE---KAREAAATTKKLEEEALK- 613
Query: 184 RAKQKKDDIESVKK 197
+KQ+ VK+
Sbjct: 614 LSKQEAQASNLVKE 655
>BQ141544 similar to GP|7022293|dbj| unnamed protein product {Homo sapiens},
partial (5%)
Length = 1284
Score = 29.6 bits (65), Expect = 1.5
Identities = 11/42 (26%), Positives = 25/42 (59%)
Frame = +2
Query: 149 RLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKD 190
R+ +++ RR+ RE ++ KE++S+K + ++K+D
Sbjct: 1049 RVAGRERREGRRRRRRRVRERGKIKKEIESEKERRERERKRD 1174
>TC78205 homologue to PIR|T47559|T47559 60S ribosomal protein-like -
Arabidopsis thaliana, partial (93%)
Length = 858
Score = 29.6 bits (65), Expect = 1.5
Identities = 22/92 (23%), Positives = 39/92 (41%), Gaps = 10/92 (10%)
Frame = +1
Query: 133 YAEMVKTDSHMEKVKSRLLAEKQKM----------VEAEERRKAREAKRLSKEVQSQKLK 182
Y + K D E VK R A K+ V ++R + E + ++E +++K
Sbjct: 223 YRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRSEKPEVRDAAREAALREIK 402
Query: 183 ERAKQKKDDIESVKKWRKQRQQSGFADDGADK 214
ER K+ KD+ ++ K + + Q K
Sbjct: 403 ERVKKTKDEKKAKKAESQAKAQKSVGKGNVSK 498
>TC87464 homologue to GP|7635498|emb|CAB88668.1 histone H2B {Cicer
arietinum}, complete
Length = 677
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/36 (38%), Positives = 22/36 (60%)
Frame = +3
Query: 152 AEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQ 187
AEK+ +A +K + K++SKE S K K+R K+
Sbjct: 87 AEKKPAEKAPAEKKPKAEKKISKEGSSDKKKKRTKK 194
>BQ137241 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (8%)
Length = 1176
Score = 29.3 bits (64), Expect = 1.9
Identities = 27/140 (19%), Positives = 57/140 (40%), Gaps = 2/140 (1%)
Frame = +2
Query: 164 RKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKV 223
R R ++ KE ++ + K+ ++K + +++ G AD+
Sbjct: 245 RTTRNSRPRPKETRAARPKKTPNERKRGRRRGTQTHNKKEPRGGADEKP----------- 391
Query: 224 FERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTND 283
ER+ K+ G R G K ++ + + K+G G++G + + ++D TN+
Sbjct: 392 -ERTHKRHKNKQQGGRRGKKGERRGRERAREKRG----------GERGQETEQSSDATNN 538
Query: 284 --FGGFSKKGAVGGSKKRKR 301
G ++K R+R
Sbjct: 539 EKSGTHTRKRRAATDASRQR 598
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.130 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,116,764
Number of Sequences: 36976
Number of extensions: 60711
Number of successful extensions: 480
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 461
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 473
length of query: 301
length of database: 9,014,727
effective HSP length: 96
effective length of query: 205
effective length of database: 5,465,031
effective search space: 1120331355
effective search space used: 1120331355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC121233.5


