

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119419.5 - phase: 0
(735 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
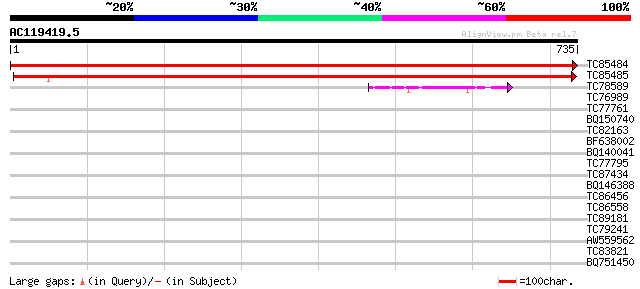
Sequences producing significant alignments: (bits) Value
TC85484 similar to PIR|S58083|S58083 transketolase (EC 2.2.1.1) ... 1484 0.0
TC85485 similar to GP|22136900|gb|AAM91794.1 putative transketol... 1228 0.0
TC78589 1-deoxy-D-xylulose 5-phosphate synthase 2 [Medicago trun... 48 2e-05
TC76989 homologue to SP|O24310|EFTU_PEA Elongation factor Tu ch... 42 0.001
TC77761 1-deoxy-D-xylulose 5-phosphate synthase 1 [Medicago trun... 38 0.012
BQ150740 similar to GP|15982842|gb| At2g45290/F4L23.20 {Arabidop... 36 0.061
TC82163 similar to PIR|T47804|T47804 hypothetical protein F24G16... 35 0.079
BF638002 weakly similar to GP|20260150|gb strong similarity to n... 35 0.10
BQ140041 similar to GP|5712672|gb|A HMG-CoA reductase {Artemisia... 35 0.10
TC77795 similar to GP|15450707|gb|AAK96625.1 At1g01090/T25K16_8 ... 34 0.18
TC87434 similar to SP|P52902|ODPA_PEA Pyruvate dehydrogenase E1 ... 33 0.30
BQ146388 weakly similar to GP|13877607|gb Unknown protein {Arabi... 33 0.30
TC86456 similar to SP|P52902|ODPA_PEA Pyruvate dehydrogenase E1 ... 33 0.30
TC86558 similar to GP|20805105|dbj|BAB92777. contains ESTs AU065... 32 0.67
TC89181 similar to PIR|T47628|T47628 sugar-phosphate isomerase-l... 32 0.67
TC79241 similar to GP|21740501|emb|CAD40825. OSJNBa0006B20.16 {O... 32 1.1
AW559562 similar to GP|14486705|gb phosphatidylinositol transfer... 31 1.5
TC83821 similar to GP|8953400|emb|CAB96673.1 1-D-deoxyxylulose 5... 31 2.0
BQ751450 similar to GP|21954534|db 6-phosphogluconate dehydrogen... 31 2.0
>TC85484 similar to PIR|S58083|S58083 transketolase (EC 2.2.1.1) precursor -
potato (fragment), partial (95%)
Length = 2685
Score = 1484 bits (3843), Expect = 0.0
Identities = 735/735 (100%), Positives = 735/735 (100%)
Frame = +1
Query: 1 MSSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHA 60
MSSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHA
Sbjct: 97 MSSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHA 276
Query: 61 SVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDEVMRYNP 120
SVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDEVMRYNP
Sbjct: 277 SVSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDEVMRYNP 456
Query: 121 KNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPG 180
KNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPG
Sbjct: 457 KNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPG 636
Query: 181 IEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACS 240
IEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACS
Sbjct: 637 IEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACS 816
Query: 241 LAGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAA 300
LAGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAA
Sbjct: 817 LAGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAA 996
Query: 301 IKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPYEPFHVP 360
IKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPYEPFHVP
Sbjct: 997 IKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPYEPFHVP 1176
Query: 361 EDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALPTYTP 420
EDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALPTYTP
Sbjct: 1177EDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALPTYTP 1356
Query: 421 EIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAERNVRFGV 480
EIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAERNVRFGV
Sbjct: 1357EIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAERNVRFGV 1536
Query: 481 REHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGE 540
REHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGE
Sbjct: 1537REHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGE 1716
Query: 541 DGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPNLAG 600
DGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPNLAG
Sbjct: 1717DGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPNLAG 1896
Query: 601 TSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVSFVSWEL 660
TSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVSFVSWEL
Sbjct: 1897TSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVSFVSWEL 2076
Query: 661 FDEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEF 720
FDEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEF
Sbjct: 2077FDEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEF 2256
Query: 721 GITKEAVIAAAKELI 735
GITKEAVIAAAKELI
Sbjct: 2257GITKEAVIAAAKELI 2301
>TC85485 similar to GP|22136900|gb|AAM91794.1 putative transketolase
{Arabidopsis thaliana}, partial (90%)
Length = 2728
Score = 1228 bits (3177), Expect = 0.0
Identities = 607/733 (82%), Positives = 656/733 (88%), Gaps = 3/733 (0%)
Frame = +2
Query: 5 SSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQ---SHHLRHTAIHAS 61
+S+SL+L+ L R R +SL ST +L+ ++ S+ S AI A+
Sbjct: 194 ASSSLHLSQAFLARAVYLNPEIRAASLSSTSSLSFPALKSSSSSKPPTSRRRTTPAIRAT 373
Query: 62 VSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDEVMRYNPK 121
TT++SLVEKSVNTIRFLA+D+VEKANSGHPGLPMGCAPMGH+LYDEVMRYNPK
Sbjct: 374 AVETLDKTTEASLVEKSVNTIRFLAIDAVEKANSGHPGLPMGCAPMGHILYDEVMRYNPK 553
Query: 122 NPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPGI 181
NP WFNRDRFVLSAGHGCMLQYALLHLAGYDSV+ +DLKQFRQW S+TPGHPENFET GI
Sbjct: 554 NPSWFNRDRFVLSAGHGCMLQYALLHLAGYDSVQEDDLKQFRQWGSKTPGHPENFETYGI 733
Query: 182 EVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSL 241
EVTTGPLGQGIAN VGLALAEKHLAARFNKPD+EI+DHYTY ILGDGCQMEGI+NEACSL
Sbjct: 734 EVTTGPLGQGIANGVGLALAEKHLAARFNKPDSEIVDHYTYVILGDGCQMEGISNEACSL 913
Query: 242 AGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAI 301
AGHWGLGKLIAFYDDNHISIDGDTEIAFTE+V++RFE LGWHVIWVKNGNTG+DEIRAAI
Sbjct: 914 AGHWGLGKLIAFYDDNHISIDGDTEIAFTENVDQRFEALGWHVIWVKNGNTGYDEIRAAI 1093
Query: 302 KEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPYEPFHVPE 361
KEAKAV D+PTLIK TTTIG+GSPNK+NSYSVHG+ALGAKEVDATR NLGWPYEPFHVPE
Sbjct: 1094KEAKAVTDKPTLIKVTTTIGFGSPNKANSYSVHGAALGAKEVDATRQNLGWPYEPFHVPE 1273
Query: 362 DVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALPTYTPE 421
DVKKHWSRH EGAALE+EWNAKFA+YEKKYKEEAA LK+II+ +LPA WEKALPTYTPE
Sbjct: 1274DVKKHWSRHTPEGAALEAEWNAKFAEYEKKYKEEAAELKAIINRELPADWEKALPTYTPE 1453
Query: 422 IPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAERNVRFGVR 481
PADATRNLSQQNLNALA VLPGLIGGSADLASSNMTLLK FGDFQ +P ERNVRFGVR
Sbjct: 1454TPADATRNLSQQNLNALAKVLPGLIGGSADLASSNMTLLKSFGDFQKATPEERNVRFGVR 1633
Query: 482 EHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHDSIGLGED 541
EH MGAICNGIALHS G IPYCATFFVFTDYMR AIR+SAL EAGVIYVMTHDSIGLGED
Sbjct: 1634EHAMGAICNGIALHSPGFIPYCATFFVFTDYMRAAIRISALCEAGVIYVMTHDSIGLGED 1813
Query: 542 GPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPNLAGT 601
GPTHQPIEHLASFRAMPN+LMLRPADGNETAG+Y+VAVLN+KRPSILALSRQKLPNL GT
Sbjct: 1814GPTHQPIEHLASFRAMPNVLMLRPADGNETAGSYRVAVLNQKRPSILALSRQKLPNLPGT 1993
Query: 602 SIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVSFVSWELF 661
SIEGVEKGGY +SDNS+GNKPDVILIGTGSELEIA A +DLRKEGK VRVVSFVSWELF
Sbjct: 1994SIEGVEKGGYTISDNSSGNKPDVILIGTGSELEIAAAAADDLRKEGKTVRVVSFVSWELF 2173
Query: 662 DEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEFG 721
D+QS YKESVLP +VTARVSIEAGSTFGW KIVGSKGK IGIDRFGASAPAG IYKE+G
Sbjct: 2174DDQSDEYKESVLPASVTARVSIEAGSTFGWHKIVGSKGKTIGIDRFGASAPAGKIYKEYG 2353
Query: 722 ITKEAVIAAAKEL 734
ITKEAVI+AAKE+
Sbjct: 2354ITKEAVISAAKEV 2392
>TC78589 1-deoxy-D-xylulose 5-phosphate synthase 2 [Medicago truncatula]
Length = 2760
Score = 47.8 bits (112), Expect = 2e-05
Identities = 57/197 (28%), Positives = 82/197 (40%), Gaps = 10/197 (5%)
Frame = +3
Query: 466 FQSDSPAERNVRFGVREHGMGAICNGIALHSRGLIPYCATFFVFTDYMRG---AIRLSAL 522
FQ P +R G+ E G+A + GL P+CA + F RG + L
Sbjct: 1467 FQKRFP-DRCFDVGIAEQHAVTFAAGLA--AEGLKPFCAIYSSFLQ--RGYDQVVHDVDL 1631
Query: 523 SEAGVIYVMTHDSIGL-GEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLN 581
+ V + M D GL G DGPTH + +PN++++ P+D E A
Sbjct: 1632 QKLPVRFAM--DRAGLVGADGPTHCGAFDITFMACLPNMIVMAPSDEAELMNMVATAAAI 1805
Query: 582 RKRPSILALSR-----QKLP-NLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEI 635
RPS R LP N GT +E + KG ++ + V ++G G ++
Sbjct: 1806 DDRPSCFRFPRGNGIGANLPLNNKGTPLE-IGKGRILLEGSR------VAILGYGCMVQQ 1964
Query: 636 AYKAGEDLRKEGKAVRV 652
KA E LR G V V
Sbjct: 1965 CMKAAEMLRAVGVYVTV 2015
>TC76989 homologue to SP|O24310|EFTU_PEA Elongation factor Tu chloroplast
precursor (EF-Tu). [Garden pea] {Pisum sativum}, partial
(92%)
Length = 1907
Score = 41.6 bits (96), Expect = 0.001
Identities = 26/72 (36%), Positives = 41/72 (56%)
Frame = +3
Query: 1 MSSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHA 60
+SSSSST L L++P+ HT + +T+ +S+ + T +R THLS S T +H
Sbjct: 156 ISSSSSTKLKLSSPY-PPHTHTFTTSSSASISNPSTTHRPKLTLTHLSSSFLNPSTILHL 332
Query: 61 SVSAPPSTTTDS 72
+ S+ P+ T S
Sbjct: 333 TPSSKPTPTRSS 368
>TC77761 1-deoxy-D-xylulose 5-phosphate synthase 1 [Medicago truncatula]
Length = 2595
Score = 38.1 bits (87), Expect = 0.012
Identities = 44/174 (25%), Positives = 68/174 (38%), Gaps = 8/174 (4%)
Frame = +3
Query: 487 AICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSA-LSEAGVIYVMTHDSIGL-GEDGPT 544
A+ L GL P+CA + F + L + V + M D GL G DGPT
Sbjct: 1473 AVTFAAGLACEGLKPFCAIYSSFLQRAYDQVVHDVDLQKLPVRFAM--DRAGLVGSDGPT 1646
Query: 545 HQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSR-----QKLP-NL 598
H + +PN++++ P+D E A RPS R +LP
Sbjct: 1647 HSGSFDVTFMACLPNMVVMAPSDEAELCHMVATAAAIDDRPSCFRYPRGNGIGVELPTEY 1826
Query: 599 AGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRV 652
G +E + KG ++ V L+G GS ++ A + + G + V
Sbjct: 1827 KGIPLE-IGKGRILIEGER------VALLGYGSAVQNCLAAASLVEQHGLRLTV 1967
>BQ150740 similar to GP|15982842|gb| At2g45290/F4L23.20 {Arabidopsis
thaliana}, partial (2%)
Length = 981
Score = 35.8 bits (81), Expect = 0.061
Identities = 21/41 (51%), Positives = 22/41 (53%), Gaps = 1/41 (2%)
Frame = +2
Query: 137 HGCMLQYALLHLAGYDSVKVEDL-KQFRQWESRTPGHPENF 176
H MLQYALLHLAGYDSV D F E + NF
Sbjct: 26 HEGMLQYALLHLAGYDSVHEYDFHSSFHNGEPKLLDISNNF 148
>TC82163 similar to PIR|T47804|T47804 hypothetical protein F24G16.70 -
Arabidopsis thaliana, partial (56%)
Length = 528
Score = 35.4 bits (80), Expect = 0.079
Identities = 24/76 (31%), Positives = 38/76 (49%)
Frame = +1
Query: 357 FHVPEDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALP 416
FH PE W H A+L++ + +Y+KK KEE + G+L A +K +
Sbjct: 310 FHSPE-----W--HAARLASLKTSHTITWEEYKKKQKEE-----ELKKGELEADADKMMR 453
Query: 417 TYTPEIPADATRNLSQ 432
Y ++ A+ +R LSQ
Sbjct: 454 EYRAQLDAERSRKLSQ 501
>BF638002 weakly similar to GP|20260150|gb strong similarity to naringenin
3-dioxygenase {Arabidopsis thaliana}, partial (40%)
Length = 652
Score = 35.0 bits (79), Expect = 0.10
Identities = 26/99 (26%), Positives = 40/99 (40%), Gaps = 2/99 (2%)
Frame = +2
Query: 5 SSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQS--HHLRHTAIHASV 62
SS+S Y + H S P + SLPS ++ TP+ TH +S HH RH +
Sbjct: 38 SSSSHYSDGTYRRTH-SGPIHNPIGSLPSPTRIHPTPSKPTHQHRSHCHHPRHRPLQLQH 214
Query: 63 SAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLP 101
P + +N R + + + HP +P
Sbjct: 215 QPPQ--------LHPRINLTRLPRMGRIPRNQPRHPTIP 307
>BQ140041 similar to GP|5712672|gb|A HMG-CoA reductase {Artemisia annua},
partial (11%)
Length = 302
Score = 35.0 bits (79), Expect = 0.10
Identities = 18/54 (33%), Positives = 24/54 (44%)
Frame = +3
Query: 13 NPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAPP 66
N H R + PSTIT + +P+T L HHL H + V +PP
Sbjct: 60 NHHRRRRDQPSFPPSQITFPSTITKSLRCSPSTSLPHQHHLLHALLRRRVLSPP 221
>TC77795 similar to GP|15450707|gb|AAK96625.1 At1g01090/T25K16_8
{Arabidopsis thaliana}, partial (82%)
Length = 1690
Score = 34.3 bits (77), Expect = 0.18
Identities = 31/117 (26%), Positives = 50/117 (42%)
Frame = +2
Query: 188 LGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHWGL 247
+G+GI A G A + K+ N+ D+ D+ T GDG G E ++A W L
Sbjct: 746 IGEGIPVATGAAFSMKYKREVLNQADS---DNVTLAFFGDGTCNNGQFFECLNMAALWKL 916
Query: 248 GKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEA 304
I F +N++ G + + T E +G + + V ++R KEA
Sbjct: 917 P--IVFVVENNLWAIGMSHLRATSDPEIWKKGPAFGMPGVHVDGMDVLKVREVAKEA 1081
>TC87434 similar to SP|P52902|ODPA_PEA Pyruvate dehydrogenase E1 component
alpha subunit mitochondrial precursor (EC 1.2.4.1)
(PDHE1-A)., complete
Length = 1829
Score = 33.5 bits (75), Expect = 0.30
Identities = 35/161 (21%), Positives = 71/161 (43%), Gaps = 4/161 (2%)
Frame = +2
Query: 186 GPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHW 245
G +G + GLA +K+L NE + T+ + GDG +G EA ++A W
Sbjct: 740 GIVGAQVPLGCGLAFGQKYLK-------NESV---TFALYGDGAANQGQLFEALNIAALW 889
Query: 246 GLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAK 305
L ++ ++++ + A + + KR G +V +K ++ A+K AK
Sbjct: 890 DLPAILVCENNHYGMGTAEWRAAKSPAYYKR----GDYVPGLKVDGMDVLAVKQAVKFAK 1057
Query: 306 --AVKDRPTLIKFTT--TIGYGSPNKSNSYSVHGSALGAKE 342
A+++ P +++ T G+ + ++Y G ++
Sbjct: 1058EHALQNGPIILEMDTYRYHGHSMSDPGSTYRTRDEISGVRQ 1180
>BQ146388 weakly similar to GP|13877607|gb Unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 633
Score = 33.5 bits (75), Expect = 0.30
Identities = 15/35 (42%), Positives = 24/35 (67%), Gaps = 6/35 (17%)
Frame = +2
Query: 32 PSTITLNRTPTPTTHLSQSHHLRH------TAIHA 60
PST+T++ TP P+TH++Q +L H +AIH+
Sbjct: 5 PSTLTVSPTPRPSTHIAQPSNLNHHIPSPSSAIHS 109
>TC86456 similar to SP|P52902|ODPA_PEA Pyruvate dehydrogenase E1 component
alpha subunit mitochondrial precursor (EC 1.2.4.1)
(PDHE1-A)., partial (92%)
Length = 1781
Score = 33.5 bits (75), Expect = 0.30
Identities = 37/161 (22%), Positives = 69/161 (41%), Gaps = 4/161 (2%)
Frame = +2
Query: 186 GPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGIANEACSLAGHW 245
G +G I VGLA +K +NK N T+ + GDG +G EA ++A W
Sbjct: 713 GIVGAQIPLGVGLAFGQK-----YNKDPN-----VTFTLYGDGAANQGQLFEALNIAALW 862
Query: 246 GLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAK 305
L ++ ++++ + A + + KR G + +K ++ A K AK
Sbjct: 863 DLPAILVCENNHYGMGTAEWRSAKSPAYYKR----GDYAPGLKVDGMDVLAVKQACKFAK 1030
Query: 306 --AVKDRPTLIKFTT--TIGYGSPNKSNSYSVHGSALGAKE 342
A+K+ P +++ T G+ + ++Y G ++
Sbjct: 1031EHALKNGPLILEMDTYRYHGHSMSDPGSTYRTRDEISGVRQ 1153
>TC86558 similar to GP|20805105|dbj|BAB92777. contains ESTs AU065506(C0784)
AU083422(C0784)~unknown protein, partial (47%)
Length = 1346
Score = 32.3 bits (72), Expect = 0.67
Identities = 22/89 (24%), Positives = 41/89 (45%), Gaps = 2/89 (2%)
Frame = +1
Query: 285 IWVKNGNTGFDEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVD 344
+W+ NT + RA KEA+ ++ + + F PN+ + Y++H
Sbjct: 475 VWLGTFNTAEEAARAYDKEARKIRGKKAKVNF--------PNEDDEYTIH---------- 600
Query: 345 ATR--NNLGWPYEPFHVPEDVKKHWSRHI 371
ATR NN P P +P ++H+ +++
Sbjct: 601 ATRRYNNNPPPIRPQKLPLYHQQHYQKNL 687
>TC89181 similar to PIR|T47628|T47628 sugar-phosphate isomerase-like protein
- Arabidopsis thaliana, partial (54%)
Length = 817
Score = 32.3 bits (72), Expect = 0.67
Identities = 21/71 (29%), Positives = 36/71 (50%), Gaps = 10/71 (14%)
Frame = +3
Query: 8 SLYLTNPHLTRHTSSPSTTRL----------SSLPSTITLNRTPTPTTHLSQSHHLRHTA 57
SL +NP+ T TSS + + + S+LP+ + + P + L++SH +
Sbjct: 105 SLISSNPNKTISTSSSTESTIPKPYHSPALSSTLPAPSSSPASANPASSLTKSHKHSSPS 284
Query: 58 IHASVSAPPST 68
+ A S+PPST
Sbjct: 285 VSAPPSSPPST 317
>TC79241 similar to GP|21740501|emb|CAD40825. OSJNBa0006B20.16 {Oryza
sativa}, partial (77%)
Length = 1285
Score = 31.6 bits (70), Expect = 1.1
Identities = 28/90 (31%), Positives = 41/90 (45%), Gaps = 5/90 (5%)
Frame = +3
Query: 232 EGIANEACSLAGHWGLGKLIAFYDDNHISIDGDTEI-AF----TESVEKRFEGLGWHVIW 286
+G A SLAG W K +A + +S DGD +I AF +++ R E L
Sbjct: 477 DGGAGGGISLAGTWWDKKALAIAKEVTMSFDGDLQIYAFKTLVNSTIQVRIEKLS----- 641
Query: 287 VKNGNTGFDEIRAAIKEAKAVKDRPTLIKF 316
K+G+ ++I A +A D L KF
Sbjct: 642 NKSGSPTMEDIEAFSTAYRAKLDEAELAKF 731
>AW559562 similar to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (34%)
Length = 668
Score = 31.2 bits (69), Expect = 1.5
Identities = 20/77 (25%), Positives = 34/77 (43%), Gaps = 2/77 (2%)
Frame = +3
Query: 576 KVAVLNRKRPSIL--ALSRQKLPNLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSEL 633
K+ VL K S L + +LP G + ++GG + SD P+++ + E
Sbjct: 138 KIHVLGNKYQSKLLEVIDASELPEFLGGTCSCADEGGCLRSDKGPWKNPEILKMVLNGEP 317
Query: 634 EIAYKAGEDLRKEGKAV 650
A + + L EGK +
Sbjct: 318 RRARQVVKVLNSEGKVI 368
>TC83821 similar to GP|8953400|emb|CAB96673.1 1-D-deoxyxylulose 5-phosphate
synthase-like protein {Arabidopsis thaliana}, partial
(46%)
Length = 1172
Score = 30.8 bits (68), Expect = 2.0
Identities = 29/119 (24%), Positives = 45/119 (37%), Gaps = 4/119 (3%)
Frame = +1
Query: 538 LGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPN 597
+G DGP + +PN++++ P+D E A +P R L
Sbjct: 553 VGSDGPLQCGAFDITFMSCLPNMIVMAPSDEAELVHMVATAAHINDQPVCFRYPRGALVG 732
Query: 598 LAGTSIEGVE----KGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRV 652
++G+ KG +V DV L+G GS ++ KA L G V V
Sbjct: 733 KDEAILDGIPIEIGKGRILVEGK------DVALLGYGSMVQNCLKAYSLLANLGIEVTV 891
>BQ751450 similar to GP|21954534|db 6-phosphogluconate dehydrogenase
{Aspergillus oryzae}, partial (36%)
Length = 661
Score = 30.8 bits (68), Expect = 2.0
Identities = 22/49 (44%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = -1
Query: 3 SSSSTSLYLTNPHLTRHTSSPST--TRLSSLPSTITLNRTPTPTTHLSQ 49
SSS TS LT P T TSS +T TRLS+ PS P P + +S+
Sbjct: 469 SSSRTSPTLTAPTPTSRTSSSTTSSTRLSTRPSPAGETSLPRPLSLVSR 323
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,334,488
Number of Sequences: 36976
Number of extensions: 301210
Number of successful extensions: 2037
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 1972
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2020
length of query: 735
length of database: 9,014,727
effective HSP length: 103
effective length of query: 632
effective length of database: 5,206,199
effective search space: 3290317768
effective search space used: 3290317768
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC119419.5


