

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
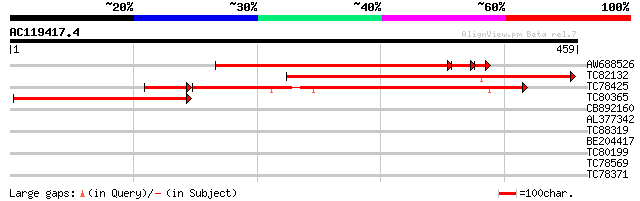
Query= AC119417.4 - phase: 0 /pseudo
(459 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW688526 similar to GP|20161335|db putative cytokinin oxidase {O... 394 e-119
TC82132 similar to PIR|T00807|T00807 probable cytokinin oxidase ... 394 e-110
TC78425 similar to GP|11120514|gb|AAG30908.1 cytokinin oxidase {... 234 7e-64
TC80365 similar to PIR|T49185|T49185 cytokinin oxidase-like prot... 139 3e-33
CB892160 similar to PIR|T49185|T491 cytokinin oxidase-like prote... 40 0.001
AL377342 similar to GP|13549123|gb putative short-chain type alc... 32 0.51
TC88319 similar to GP|19387174|gb|AAL87123.1 SEC10 {Arabidopsis ... 32 0.67
BE204417 similar to PIR|T06690|T06 galactonolactone dehydrogenas... 32 0.67
TC80199 weakly similar to GP|21593112|gb|AAM65061.1 unknown {Ara... 29 3.3
TC78569 similar to GP|21536701|gb|AAM61033.1 1-acylcerol-3-phosp... 28 5.7
TC78371 similar to GP|9558523|dbj|BAB03441.1 ESTs AU078742(C1188... 28 7.4
>AW688526 similar to GP|20161335|db putative cytokinin oxidase {Oryza sativa
(japonica cultivar-group)}, partial (36%)
Length = 673
Score = 394 bits (1011), Expect(3) = e-119
Identities = 192/192 (100%), Positives = 192/192 (100%)
Frame = +2
Query: 167 FGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLL 226
FGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLL
Sbjct: 2 FGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLL 181
Query: 227 NNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEVNQDIQKHLSHLNFIPSTL 286
NNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEVNQDIQKHLSHLNFIPSTL
Sbjct: 182 NNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEVNQDIQKHLSHLNFIPSTL 361
Query: 287 FQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKETSN 346
FQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKETSN
Sbjct: 362 FQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKETSN 541
Query: 347 GPVLIYPVHKSK 358
GPVLIYPVHKSK
Sbjct: 542 GPVLIYPVHKSK 577
Score = 46.2 bits (108), Expect(3) = e-119
Identities = 20/20 (100%), Positives = 20/20 (100%)
Frame = +3
Query: 358 KWDKRTSVVIPDEDIFYLVG 377
KWDKRTSVVIPDEDIFYLVG
Sbjct: 576 KWDKRTSVVIPDEDIFYLVG 635
Score = 28.1 bits (61), Expect(3) = e-119
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = +1
Query: 377 GFLASSSGPDELE 389
GFLASS GPDELE
Sbjct: 634 GFLASSXGPDELE 672
>TC82132 similar to PIR|T00807|T00807 probable cytokinin oxidase [imported]
- Arabidopsis thaliana, partial (37%)
Length = 995
Score = 394 bits (1011), Expect = e-110
Identities = 187/238 (78%), Positives = 209/238 (87%), Gaps = 4/238 (1%)
Frame = +2
Query: 225 LLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEVNQDIQKHLSHLNFIPS 284
+LNNWR SFNPQDPVQAS FKSDG+TLFCLELAKYFN ++ VNQD+++HLS LN+I S
Sbjct: 2 VLNNWRSSFNPQDPVQASHFKSDGKTLFCLELAKYFNFQQINIVNQDVERHLSRLNYIRS 181
Query: 285 TLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKET 344
TLFQTEVTYV+FLDRVH+SEVKLRSKGLWDVPHPWLNLFIPKSKIH+FA+ VFGNI+ +T
Sbjct: 182 TLFQTEVTYVEFLDRVHVSEVKLRSKGLWDVPHPWLNLFIPKSKIHSFAQFVFGNILTQT 361
Query: 345 SNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLA----SSSGPDELEHILSQNKRILE 400
SNGPVLIYPV KSKWD RTSVVIPDEDIFYLV FL SS+G D LEHILSQNKRILE
Sbjct: 362 SNGPVLIYPVKKSKWDNRTSVVIPDEDIFYLVAFLTSAVPSSNGTDGLEHILSQNKRILE 541
Query: 401 YCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPLAILAPGQGIFSKSIT 458
YC+R +LGVKQYLPH+ TQEEW+ H+G KWEIF QRK +YDP AILAPGQ IF K+IT
Sbjct: 542 YCQRENLGVKQYLPHHNTQEEWRDHFGTKWEIFSQRKFVYDPFAILAPGQRIFQKTIT 715
>TC78425 similar to GP|11120514|gb|AAG30908.1 cytokinin oxidase {Arabidopsis
thaliana}, partial (60%)
Length = 1478
Score = 234 bits (596), Expect(2) = 7e-64
Identities = 120/284 (42%), Positives = 174/284 (61%), Gaps = 13/284 (4%)
Frame = +1
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
++ QN +LF++ LGGLGQFG+ITRARI+L+ AP MV+WIRV+YS+F +T+D E L+
Sbjct: 640 NDNQNSDLFFASLGGLGQFGVITRARIVLQQAPDMVRWIRVIYSEFEDYTRDAEWLVTLP 819
Query: 209 KA--FDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKF-------KSDGRTLFCLELAKY 259
+ FDY+EGFV+ N N W P P+ +++ S G L+CLELA +
Sbjct: 820 EGDGFDYVEGFVVANNDDPCNGW-----PTIPMGSNQIFNPVCLPSSAGPVLYCLELALH 984
Query: 260 FNME-ETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHP 318
+ + EVN + + L L F+ F+ +V Y+DFL RV E ++KG+WD PHP
Sbjct: 985 YRKTARSSEVNTKVDRLLGGLRFVEGIKFEDDVKYMDFLLRVKRVEEDAKAKGIWDAPHP 1164
Query: 319 WLNLFIPKSKIHNFAEVVFGNIVKETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGF 378
WLN+F+ KS I +F VF I+K GP+L+YP+ +SKWD R SVV+PD +IFY++
Sbjct: 1165WLNMFVSKSDIADFDREVFKKILKHGVGGPILVYPLLRSKWDDRHSVVVPDSNIFYIIAL 1344
Query: 379 LASSSGPDE---LEHILSQNKRILEYCERAHLGVKQYLPHYTTQ 419
L P + + +++QN I++ C K YLPHYT+Q
Sbjct: 1345LRFIPPPPKGPPTDKLVAQNNAIIQLCYNKGFNFKLYLPHYTSQ 1476
Score = 28.5 bits (62), Expect(2) = 7e-64
Identities = 19/38 (50%), Positives = 23/38 (60%)
Frame = +2
Query: 110 GQIICI*QLVVLYPMLVLVVRLLGMVHRSVMFRRWRLL 147
G+II *+LV Y VLVVRL MV + R W+LL
Sbjct: 500 GRIILG*RLVERYLTPVLVVRLSVMVLKRQT*RNWKLL 613
>TC80365 similar to PIR|T49185|T49185 cytokinin oxidase-like protein -
Arabidopsis thaliana, partial (28%)
Length = 647
Score = 139 bits (349), Expect = 3e-33
Identities = 89/145 (61%), Positives = 95/145 (65%), Gaps = 1/145 (0%)
Frame = +1
Query: 4 KILATGTNLIL*QYYIQNQFLILQLL*SIYGI*VQVLT*LLQLEDMATHFKGKHKLKREL 63
KILA TN+I Q QNQFLILQ *+IYGI L LLQ EDM HFK HKL EL
Sbjct: 211 KILAIDTNIIQWQ*CTQNQFLILQQQ*NIYGIWDIALISLLQQEDMDIHFKDNHKLMEEL 390
Query: 64 *SIWNH*MLKKLRCMVENFL-MWMCQEVNYG*RFCMRH*SMVWHQDLGQIICI*QLVVLY 122
*S WNH K + M+ L MWM VN G* CMR *+M WHQD GQIICI*QLVVL
Sbjct: 391 *SKWNHSKYLKCKSMLGILLHMWMSLVVNCG*TSCMRL*NMGWHQDHGQIICI*QLVVLC 570
Query: 123 PMLVLVVRLLGMVHRSVMFRRWRLL 147
PMLVLV + L MV RSVMF RLL
Sbjct: 571 PMLVLVDKHLSMVLRSVMFSSLRLL 645
>CB892160 similar to PIR|T49185|T491 cytokinin oxidase-like protein -
Arabidopsis thaliana, partial (13%)
Length = 697
Score = 40.4 bits (93), Expect = 0.001
Identities = 29/68 (42%), Positives = 36/68 (52%)
Frame = +1
Query: 84 MWMCQEVNYG*RFCMRH*SMVWHQDLGQIICI*QLVVLYPMLVLVVRLLGMVHRSVMFRR 143
M M + NYG CM+H SM H LG II * L MLVLV + MV + ++F
Sbjct: 493 MLMLEGNNYGLMCCMKHLSMDLHLFLGLIIFT*LLEGHSLMLVLVDKPFVMVLKLLVFIN 672
Query: 144 WRLLQSEE 151
W LL +E
Sbjct: 673 WMLLLEKE 696
>AL377342 similar to GP|13549123|gb putative short-chain type alcohol
dehydrogenase {Solanum tuberosum}, partial (36%)
Length = 457
Score = 32.0 bits (71), Expect = 0.51
Identities = 26/83 (31%), Positives = 41/83 (49%), Gaps = 4/83 (4%)
Frame = +1
Query: 279 LNFIPSTLFQTEVTYV----DFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSKIHNFAE 334
+N I LF++E+T D+L+ V I V LR W +P L I + IH+ +E
Sbjct: 67 VNSISPGLFKSEITESLMKKDWLNNVAIRTVPLRE---WGTSNPALTS-IARYLIHDSSE 234
Query: 335 VVFGNIVKETSNGPVLIYPVHKS 357
V GNI + + +P++ S
Sbjct: 235 YVTGNIFIADAGATLPGFPIYSS 303
>TC88319 similar to GP|19387174|gb|AAL87123.1 SEC10 {Arabidopsis thaliana},
partial (52%)
Length = 1313
Score = 31.6 bits (70), Expect = 0.67
Identities = 38/140 (27%), Positives = 60/140 (42%), Gaps = 27/140 (19%)
Frame = +1
Query: 122 YPMLVLVVRLLGMVHRSVMFRRW-------RLLQSEEQNGELFYSVLGGLGQFGIITRAR 174
+P+L L L MVH ++ ++W RLL +++ N L S L F + R +
Sbjct: 931 FPILYLGFCFLWMVHMLLLVKKWLQQCPVQRLLLTKDYNSVLKLSWLR*NDCFQLSRRLQ 1110
Query: 175 ILLEPAPTMVKWI-----------------RVLYSDFTAFTKDQEKLIFAEKAFDYIEGF 217
I+ P M++W+ RVL S FTA ++ +E
Sbjct: 1111 II---NPLMMEWLPTTGQLMPAQGVVAYLSRVLESAFTALEGLNKQAFLSE--------- 1254
Query: 218 VIKNR--TGLLNNW-RLSFN 234
+ NR GLLN+W + +FN
Sbjct: 1255 -LGNRLHKGLLNHWQKYTFN 1311
>BE204417 similar to PIR|T06690|T06 galactonolactone dehydrogenase (EC
1.3.2.3) - Arabidopsis thaliana, partial (34%)
Length = 632
Score = 31.6 bits (70), Expect = 0.67
Identities = 38/175 (21%), Positives = 70/175 (39%), Gaps = 10/175 (5%)
Frame = +1
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S+E++ ELFY GLG G++ + +V+ S K+ +KL+
Sbjct: 124 SKEKDPELFYLARCGLGGLGVVAEVTLQCVDRQELVE--HTFVSTMDEIKKNHKKLLSEN 297
Query: 209 KAFDYI-----EGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDG-----RTLFCLELAK 258
K Y+ E V+ R ++ W+ P K+ D R L+ + K
Sbjct: 298 KHVKYLYIPYTESAVVV-RCNPVSKWK-----GPPKFKPKYTKDEAIQHVRDLYRESIQK 459
Query: 259 YFNMEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLW 313
Y + + D ++++ L+F T + + +D L++ HI +V W
Sbjct: 460 YRVEGSRNKSSDDDEQNIDELSF---TELRDRLIALDPLNKNHIVKVNQAEAEFW 615
>TC80199 weakly similar to GP|21593112|gb|AAM65061.1 unknown {Arabidopsis
thaliana}, partial (41%)
Length = 934
Score = 29.3 bits (64), Expect = 3.3
Identities = 19/72 (26%), Positives = 34/72 (46%)
Frame = -1
Query: 320 LNLFIPKSKIHNFAEVVFGNIVKETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFL 379
+NL I KI+ + N++K+ + + VHK W K T + +E+ ++ L
Sbjct: 841 INLHIILIKIYRQTQEKIANVIKKDLHKFITFSMVHKIYWIKDT--IRRNENKHCMINKL 668
Query: 380 ASSSGPDELEHI 391
S P +EH+
Sbjct: 667 GSHI*PKHMEHV 632
>TC78569 similar to GP|21536701|gb|AAM61033.1 1-acylcerol-3-phosphate
acyltransferase-like protein {Arabidopsis thaliana},
partial (92%)
Length = 1503
Score = 28.5 bits (62), Expect = 5.7
Identities = 14/50 (28%), Positives = 27/50 (54%)
Frame = -2
Query: 223 TGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEVNQDI 272
+G +NNW L+ Q +Q S + T+ + L K+F E+ L++ + +
Sbjct: 311 SGSVNNWILTPAHQSKIQTSSSQRSSATILLIRLYKFF--EKGLKITKHV 168
>TC78371 similar to GP|9558523|dbj|BAB03441.1 ESTs AU078742(C11888)
C26225(C11888) correspond to a region of the predicted
gene.~Similar to Oryza, partial (2%)
Length = 815
Score = 28.1 bits (61), Expect = 7.4
Identities = 13/38 (34%), Positives = 23/38 (60%), Gaps = 2/38 (5%)
Frame = +2
Query: 174 RILLEPAPTMVKWIRVLYSDFTAFTKDQE--KLIFAEK 209
RIL+ PT +KW+ + + +T F+ DQ ++F E+
Sbjct: 584 RILVNELPTSLKWLFLCDNPYTEFSVDQNLINILFLEE 697
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.336 0.148 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,691,655
Number of Sequences: 36976
Number of extensions: 237167
Number of successful extensions: 1773
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1705
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1767
length of query: 459
length of database: 9,014,727
effective HSP length: 99
effective length of query: 360
effective length of database: 5,354,103
effective search space: 1927477080
effective search space used: 1927477080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC119417.4


