

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119417.2 + phase: 0
(898 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
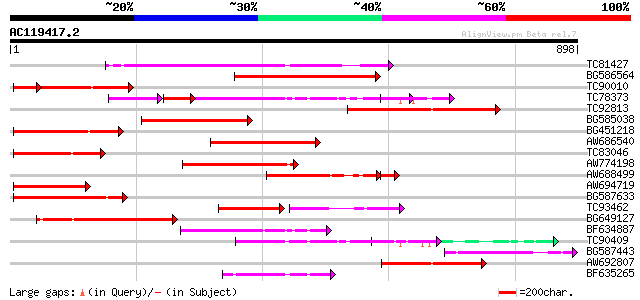
Score E
Sequences producing significant alignments: (bits) Value
TC81427 weakly similar to GP|11994216|dbj|BAB01338. disease resi... 265 4e-71
BG586564 weakly similar to GP|14348622|gb NBS-LRR resistance-lik... 250 1e-66
TC90010 similar to GP|5817349|gb|AAD52718.1| putative NBS-LRR ty... 183 7e-64
TC78373 weakly similar to GP|14348622|gb|AAK61318.1 NBS-LRR resi... 172 5e-63
TC92813 233 3e-61
BG585038 weakly similar to GP|11994216|db disease resistance com... 229 3e-60
BG451218 similar to GP|11994217|db disease resistance comples pr... 227 1e-59
AW686540 weakly similar to GP|15289776|db Lycopersicon esculentu... 221 1e-57
TC83046 weakly similar to GP|10280097|emb|CAC09980. nucleotide s... 208 6e-54
AW774198 182 6e-46
AW688499 similar to GP|4234953|gb|A NBS-LRR-like protein cD7 {Ph... 155 2e-44
AW694719 similar to GP|11994217|db disease resistance comples pr... 177 2e-44
BG587633 similar to GP|20385442|gb| resistance gene analog {Viti... 176 4e-44
TC93462 112 1e-43
BG649127 weakly similar to GP|11994217|db disease resistance com... 164 1e-40
BF634887 weakly similar to GP|10280097|em nucleotide sequence of... 163 2e-40
TC90409 weakly similar to GP|10280097|emb|CAC09980. nucleotide s... 152 7e-37
BG587443 weakly similar to PIR|T06405|T064 resistance complex pr... 146 3e-35
AW692807 134 1e-31
BF635265 similar to GP|4234955|gb|A NBS-LRR-like protein cD8 {Ph... 132 5e-31
>TC81427 weakly similar to GP|11994216|dbj|BAB01338. disease resistance
comples protein {Arabidopsis thaliana}, partial (6%)
Length = 1253
Score = 265 bits (678), Expect = 4e-71
Identities = 168/457 (36%), Positives = 242/457 (52%), Gaps = 2/457 (0%)
Frame = +3
Query: 153 TFFDFKCLRSFLPIGSRLQESYLSCKVIDDLIPSIKRLRMLSLSNYNITVLPNSINKLVQ 212
TF FK F P+ L S + L+ K LR+ SLS Y IT+LP+SI L+
Sbjct: 6 TFMPFK----FYPVVPSLGGISAS---VSTLLKKPKPLRVFSLSEYPITLLPSSIGHLLH 164
Query: 213 LRYLNLSHTDIKCLPDTTCDLYYLQTLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKK 272
LRYL+LS T I LPD+ C+LY L LLL GC L LP KLINLR LDIS + IKK
Sbjct: 165 LRYLDLSRTPITSLPDSICNLYNLXALLLVGCADLTLLPTKTSKLINLRQLDISGSGIKK 344
Query: 273 MPMQIVRLENLQTLTVFLVGKQKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANL 332
MP + +L++LQ+L F+V G ++ ELG+ LRG L I NL+N + A +A L
Sbjct: 345 MPTNLGKLKSLQSLPRFVVSNDG-GSNVGELGEMLELRGSLSIVNLENVLLKEXASNAGL 521
Query: 333 KHKVHLEELEVYWDQQTEESPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSN 392
K K +L E+E W T + +I + L P NLK+L I +GG FP+WLG S S
Sbjct: 522 KRKKYLHEVEFKWTTPTHSQESENIIFDMLXPHRNLKRLKINNFGGEKFPNWLGSNSGST 701
Query: 393 MVYLSIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSL 452
M+ L + C C++LP LGQ+ L+E+ I ++R++ +GPEFYG + F+ F SL
Sbjct: 702 MMSLYLDECGNCLSLPSLGQLSNLREIYITSVTRLQKVGPEFYG-------NGFEAFSSL 860
Query: 453 EKLEFNSMPSWREW-ISFRGSKFPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNH 511
++F M +W EW ++ + F L+ L + +C +L G LP +LPS++K+ I C
Sbjct: 861 RIIKFKDMLNWEEWSVNNQSGSEGFTLLQELYIENCPKLIGKLPGNLPSLDKLVITSCQT 1040
Query: 512 FPATLSTLHWLSSVKSLDLMCQGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMS-S 570
T+ + L +K I G +SL M +
Sbjct: 1041LSDTMPCVPRLRELK---------------------------ISGCEAFVSLSEQMMKCN 1139
Query: 571 TCLQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNL 607
CLQ + + SL + P + + +L+SL++ +CQ L
Sbjct: 1140DCLQTMAISNCPSLVSIPMDCVSRTLKSLKVSDCQKL 1250
Score = 37.7 bits (86), Expect = 0.020
Identities = 38/115 (33%), Positives = 54/115 (46%), Gaps = 1/115 (0%)
Frame = +3
Query: 594 TSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCS 653
T LQ L I+ C L P + LV + D++ P L+ L I GC
Sbjct: 936 TLLQELYIENCPKLIGKLPGNLPSLDKLVITSCQTLSDTMPCV-----PRLRELKISGCE 1100
Query: 654 SLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLF-VLKSLTLDKLSLC 707
+ S+ SE+ + LQ++ +SNC SL S+P MD + LKSL K+S C
Sbjct: 1101 AFVSL--SEQMMKCN-DCLQTMAISNCPSLVSIP--MDCVSRTLKSL---KVSDC 1241
Score = 33.9 bits (76), Expect = 0.29
Identities = 37/116 (31%), Positives = 52/116 (43%), Gaps = 2/116 (1%)
Frame = +3
Query: 571 TCLQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNC- 629
T LQ L + L LP SL L I CQ L+ +T L L++ C
Sbjct: 936 TLLQELYIENCPKLIGKLPGNLP-SLDKLVITSCQTLS----DTMPCVPRLRELKISGCE 1100
Query: 630 -CDSLTSFQLNGFPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLR 684
SL+ + LQ ++I C SL SI + + TL+SLKVS+C+ L+
Sbjct: 1101 AFVSLSEQMMKCNDCLQTMAISNCPSLVSIPMDCVSR-----TLKSLKVSDCQKLQ 1253
Score = 30.8 bits (68), Expect = 2.4
Identities = 44/199 (22%), Positives = 80/199 (40%), Gaps = 18/199 (9%)
Frame = +3
Query: 666 SLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCCEVACLPPKLQFMHIESL 725
S S ST+ SL + C + SLP + L L+ + + ++ +V P+ E+
Sbjct: 684 SNSGSTMMSLYLDECGNCLSLPS-LGQLSNLREIYITSVTRLQKVG---PEFYGNGFEAF 851
Query: 726 GLATPVT--------EW------GFQSLCFLSDLHIGGDNIVNTLLKKKLLPPLLVSLTI 771
+ EW G + L +L+I +N + K P L L I
Sbjct: 852 SSLRIIKFKDMLNWEEWSVNNQSGSEGFTLLQELYI--ENCPKLIGKLPGNLPSLDKLVI 1025
Query: 772 TNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDFFPSF---LKSLVFINCPKLMSLP 828
T+ + + + + L+ L C + + L+++ NCP L+S+P
Sbjct: 1026TSCQTL----SDTMPCVPRLRELKISGCEAFVSLSEQMMKCNDCLQTMAISNCPSLVSIP 1193
Query: 829 -DMFPSSLETLEFDDCPRL 846
D +L++L+ DC +L
Sbjct: 1194MDCVSRTLKSLKVSDCQKL 1250
>BG586564 weakly similar to GP|14348622|gb NBS-LRR resistance-like protein
J78 {Phaseolus vulgaris}, partial (6%)
Length = 733
Score = 250 bits (639), Expect = 1e-66
Identities = 127/235 (54%), Positives = 160/235 (68%), Gaps = 3/235 (1%)
Frame = +2
Query: 356 EVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPF 415
E +L+ L P +NL +L+I FYGG SFPSWLGD SFSNMV L I++C YC+TLPPLGQ+
Sbjct: 29 ERVLDMLIPPVNLNRLNIYFYGGTSFPSWLGDSSFSNMVSLCIENCRYCVTLPPLGQLSS 208
Query: 416 LKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFP 475
LK+L I GMS +ETIGPEFYG+ GG +NS FQPF SLEKLEF +MP+W++W+ F+ P
Sbjct: 209 LKDLTIRGMSILETIGPEFYGIVGGGSNSSFQPFSSLEKLEFTNMPNWKKWLLFQDGILP 388
Query: 476 FPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDL---MC 532
FP LK+L L DCTELRG+LPSHL SIE+ C H + TL WLSS+K +D +
Sbjct: 389 FPCLKSLKLYDCTELRGNLPSHLSSIEEFVNKGCPHLLESPPTLEWLSSIKEIDFSGSLD 568
Query: 533 QGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAF 587
+ +DSPC LQ + F+ + SL M +SSTCL+ L L SLT F
Sbjct: 569 STETRWPFVESDSPCLLQCVALRFFDTIFSLSQMILSSTCLKFLKLHSGPSLTVF 733
>TC90010 similar to GP|5817349|gb|AAD52718.1| putative NBS-LRR type disease
resistance protein {Pisum sativum}, partial (88%)
Length = 1107
Score = 183 bits (464), Expect(2) = 7e-64
Identities = 97/153 (63%), Positives = 117/153 (76%), Gaps = 1/153 (0%)
Frame = +1
Query: 45 FPKGYPFNRKKLILLWMAEGFLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKF 104
FPK YP +RK+L+LLWMAEGFL+ S GK +EE+GDD F ELLSRSLI++ +DD EKF
Sbjct: 652 FPKDYPLDRKELVLLWMAEGFLDCSQRGKKMEELGDDCFAELLSRSLIQQLSDDDRGEKF 831
Query: 105 VMHDVVYDLATIASGKSCCRFGSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFL 164
VMHD+V DLAT SGKSCCR G I E+V H +YNQE YDIF KFE +FKCLRSFL
Sbjct: 832 VMHDLVNDLATFVSGKSCCRLECGD-IPENVRHFSYNQENYDIFMKFEKLHNFKCLRSFL 1008
Query: 165 PIG-SRLQESYLSCKVIDDLIPSIKRLRMLSLS 196
I +++YLS KV++DL+PS KRLR+LSLS
Sbjct: 1009FICLMTWRDNYLSFKVVNDLLPSHKRLRVLSLS 1107
Score = 80.5 bits (197), Expect(2) = 7e-64
Identities = 34/45 (75%), Positives = 41/45 (90%)
Frame = +3
Query: 6 AILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYP 50
+ILNS+IWN+ NDNI+P+L L+YQ+LPSHLKRCFAYCSIF KG P
Sbjct: 534 SILNSNIWNLRNDNILPALHLSYQYLPSHLKRCFAYCSIFSKGLP 668
>TC78373 weakly similar to GP|14348622|gb|AAK61318.1 NBS-LRR resistance-like
protein J78 {Phaseolus vulgaris}, partial (7%)
Length = 1463
Score = 172 bits (437), Expect(3) = 5e-63
Identities = 122/363 (33%), Positives = 178/363 (48%), Gaps = 10/363 (2%)
Frame = +1
Query: 288 VFLVGKQKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQ 347
+F+VG+Q G I+ELG +L+G+LCI +++ ID +A ANLK K H+EEL
Sbjct: 457 IFVVGEQS-GSDIKELGNLNHLQGELCISGMEHVIDPVDAAGANLKDKKHVEELIWSGSY 633
Query: 348 QTEESPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITL 407
+ + + LQP+ +LK+L I Y G F +W+ C N+V + + C C L
Sbjct: 634 KFNTNGRESDVFEALQPNSSLKRLIISHYKGNRFANWMRGCDLPNLVSIRLNLCALCSEL 813
Query: 408 PPLGQVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWI 467
PPLGQ+P LKE+ I G ++ IG EFYG STN PF+ SLE L F+SM W EW
Sbjct: 814 PPLGQLPCLKEISISGCDKIRIIGKEFYG--NNSTNVPFR---SLEILHFDSMSEWEEWS 978
Query: 468 SFRGSKFPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKS 527
G FP LK L ++ C +L+ LP HLPS++K+ I C A + ++
Sbjct: 979 HLEG----FPLLKELSIKKCPKLKRALPQHLPSLQKLEIKDCKKLEALIPK---CDNMIE 1137
Query: 528 LDL-MCQGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTA 586
LD+ C +L N+ P +L+ L N + + Q+L I +
Sbjct: 1138LDIKRCD-----RILVNELPTNLK--------SLFLCDNQYTEFSVDQNLINILFLEVLK 1278
Query: 587 FPANGLPTSLQSLRIDECQNLAFLRPETWSN---------YTSLVTLELKNCCDSLTSFQ 637
F G + SL + +L L + W + +TSL L L + C L SF
Sbjct: 1279FDFRGC-VNCPSLDLRCYNSLRDLSIKGWHSSSLPLELHLFTSLSALRLYH-CPELESFP 1452
Query: 638 LNG 640
+ G
Sbjct: 1453MGG 1461
Score = 61.6 bits (148), Expect(3) = 5e-63
Identities = 37/87 (42%), Positives = 50/87 (56%), Gaps = 2/87 (2%)
Frame = +2
Query: 157 FKCLRSFLPIGSRLQESYL--SCKVIDDLIPSIKRLRMLSLSNYNITVLPNSINKLVQLR 214
FK LRSFL + ++ Y S + DL +K LRMLS + + L I L LR
Sbjct: 56 FKGLRSFLAVPGGYRKDYFMTSNNLQRDLFSKLKYLRMLSFCGFKLKELSGEIGNLKLLR 235
Query: 215 YLNLSHTDIKCLPDTTCDLYYLQTLLL 241
YLNL+ + I+ LPD+ C LY L+TL+L
Sbjct: 236 YLNLTASLIERLPDSICKLYKLETLIL 316
Score = 47.4 bits (111), Expect(3) = 5e-63
Identities = 23/50 (46%), Positives = 33/50 (66%)
Frame = +3
Query: 244 CWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLTVFLVGK 293
C++L +LP K ++LRHL + IKKMP +I RL +LQTL+ F G+
Sbjct: 324 CFELTKLPSKFYKFVSLRHLHLEGCNIKKMPKKIGRLNHLQTLSDFCCGR 473
Score = 43.9 bits (102), Expect = 3e-04
Identities = 35/124 (28%), Positives = 61/124 (48%), Gaps = 7/124 (5%)
Frame = +1
Query: 588 PANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLV---TLELKNCCDSLTSFQ----LNG 640
P LP L+ + I C + + E + N ++ V +LE+ + DS++ ++ L G
Sbjct: 817 PLGQLPC-LKEISISGCDKIRIIGKEFYGNNSTNVPFRSLEILHF-DSMSEWEEWSHLEG 990
Query: 641 FPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLT 700
FP+L+ LSI+ C LK L +LQ L++ +CK L +L + D + L
Sbjct: 991 FPLLKELSIKKCPKLKRALPQH------LPSLQKLEIKDCKKLEALIPKCDNMIELDIKR 1152
Query: 701 LDKL 704
D++
Sbjct: 1153CDRI 1164
Score = 30.0 bits (66), Expect = 4.1
Identities = 21/45 (46%), Positives = 26/45 (57%), Gaps = 2/45 (4%)
Frame = +1
Query: 809 FPSFLKSLVFINCPKLM-SLPDMFPSSLETLEFDDCPRL-GLLPR 851
FP LK L CPKL +LP PS L+ LE DC +L L+P+
Sbjct: 991 FP-LLKELSIKKCPKLKRALPQHLPS-LQKLEIKDCKKLEALIPK 1119
>TC92813
Length = 784
Score = 233 bits (593), Expect = 3e-61
Identities = 135/247 (54%), Positives = 173/247 (69%), Gaps = 4/247 (1%)
Frame = +3
Query: 535 SPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPT 594
S +LSLL +DSPC +Q I KLL++P + + STCL HL L +SSLTAFP++GLPT
Sbjct: 27 SSQLSLLESDSPCMMQDVEIKKCVKLLAVPKLILKSTCLTHLGLDSLSSLTAFPSSGLPT 206
Query: 595 SLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSS 654
SLQSL I C+NL+FL PETW NYTSLV+L+ CD+LTSF L+GFP LQ L+I C S
Sbjct: 207 SLQSLNIQCCENLSFLPPETWINYTSLVSLKFYRSCDTLTSFPLDGFPALQTLTICECRS 386
Query: 655 LKSIFISEKNSSLSLSTLQSLKVSNCKSLR--SLPQRMDTLFVLKSLTLDKLSLC-CEVA 711
L SI+ISE++S S S+L+SL++ + S+ + +MD L L+ LTLD + L CE
Sbjct: 387 LDSIYISERSSPRS-SSLESLEIISPDSIELFEVKLKMDMLTALERLTLDCVELSFCEGV 563
Query: 712 CLPPKLQFMHIESLGLATPVTEWGFQSLCFLSDLHI-GGDNIVNTLLKKKLLPPLLVSLT 770
CLPPKLQ + I + A PVTEWG Q L LSDL I GD+I NTL+K+ LLP LV+LT
Sbjct: 564 CLPPKLQSIKISTQKTAPPVTEWGLQYLTALSDLGIVKGDDIFNTLMKESLLPIXLVTLT 743
Query: 771 ITNLTEM 777
I +L+EM
Sbjct: 744 IRDLSEM 764
>BG585038 weakly similar to GP|11994216|db disease resistance comples protein
{Arabidopsis thaliana}, partial (5%)
Length = 528
Score = 229 bits (585), Expect = 3e-60
Identities = 116/175 (66%), Positives = 138/175 (78%)
Frame = +3
Query: 210 LVQLRYLNLSHTDIKCLPDTTCDLYYLQTLLLSGCWKLIELPIHVGKLINLRHLDISYTK 269
LV+LRYL+LS T IK LP+ TC+LY LQTL L+ C L ELP + GKLINLRHLDIS T
Sbjct: 3 LVELRYLDLSFTGIKSLPNATCNLYNLQTLNLTRCENLTELPPNFGKLINLRHLDISETN 182
Query: 270 IKKMPMQIVRLENLQTLTVFLVGKQKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACD 329
IK+MPMQIV L NLQTLTVF VGKQ GLS++E+ KFPNLRGKLCIKNLQN ID EA D
Sbjct: 183 IKEMPMQIVGLNNLQTLTVFSVGKQDTGLSLKEVCKFPNLRGKLCIKNLQNVIDAIEAYD 362
Query: 330 ANLKHKVHLEELEVYWDQQTEESPTNEVILNELQPSINLKKLSIKFYGGISFPSW 384
N+++K +EELE+ W +QTE+S + +L+ LQPS NL+KLSI+ YGG SFPSW
Sbjct: 363 VNMRNKEDIEELELQWSKQTEDSRIEKDVLDMLQPSFNLRKLSIRLYGGTSFPSW 527
>BG451218 similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (5%)
Length = 666
Score = 227 bits (579), Expect = 1e-59
Identities = 113/173 (65%), Positives = 137/173 (78%)
Frame = +1
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
ILNS++WN+PND I+P+L L+YQ LPSHLK CFAYCSIFPKG+ +RKKL+LLWMAEGFL
Sbjct: 142 ILNSNVWNLPNDKILPTLHLSYQCLPSHLKICFAYCSIFPKGHTHDRKKLVLLWMAEGFL 321
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCCRFG 126
++S K +EE+GDD F ELLSRSLI++SND+ EKF MHD+V DLAT+ SGKSCCRF
Sbjct: 322 DYSHGEKTMEELGDDCFAELLSRSLIQQSNDNGRGEKFFMHDLVNDLATVVSGKSCCRF- 498
Query: 127 SGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKV 179
G ISE+V HV+Y QEEYDI KF+ F + KCLR+FLPI +YLS KV
Sbjct: 499 ECGNISENVRHVSYIQEEYDIVTKFKPFHNLKCLRTFLPIHVWRCNNYLSFKV 657
>AW686540 weakly similar to GP|15289776|db Lycopersicon esculentum resistance
complex protein I2C-2 like protein, partial (2%)
Length = 566
Score = 221 bits (563), Expect = 1e-57
Identities = 103/174 (59%), Positives = 133/174 (76%)
Frame = +1
Query: 319 QNAIDVSEACDANLKHKVHLEELEVYWDQQTEESPTNEVILNELQPSINLKKLSIKFYGG 378
QN IDV+EA DA+LK K H+EEL + W +T++ + +L+ L+P +NL +L+I YGG
Sbjct: 1 QNVIDVAEAYDADLKSKEHIEELTLQWGVETDDPLKGKDVLDMLKPPVNLNRLNIDLYGG 180
Query: 379 ISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEFYGMT 438
SFPSWLGD SFSNMV LSI+ C YC+TLPPLGQ+ LK+L I GM +ETIGPEFYG+
Sbjct: 181 TSFPSWLGDSSFSNMVSLSIQHCGYCVTLPPLGQLSSLKDLSIRGMYILETIGPEFYGIV 360
Query: 439 GGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFPRLKTLMLRDCTELRG 492
GG +NS FQPFPSLEKL+F MP+W++W+ F+ FPFP LK+L+L +C ELRG
Sbjct: 361 GGGSNSSFQPFPSLEKLQFVKMPNWKKWLPFQDGIFPFPCLKSLILYNCPELRG 522
>TC83046 weakly similar to GP|10280097|emb|CAC09980. nucleotide sequence of
the I-2 resistance gene {Fusarium oxysporum}, partial
(8%)
Length = 884
Score = 208 bits (530), Expect = 6e-54
Identities = 100/146 (68%), Positives = 117/146 (79%)
Frame = +1
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
ILNSDIWN+PNDNI+P+L L+YQ+LPSHLKRCFAYCSIFPK + ++K+LILLWMAEGFL
Sbjct: 127 ILNSDIWNLPNDNILPALRLSYQYLPSHLKRCFAYCSIFPKDFSLDKKELILLWMAEGFL 306
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCCRFG 126
EHS K EEVG DYF ELLSRSLI++SNDD KEKFVMHD+V DLA + SG SC R
Sbjct: 307 EHSQCNKTAEEVGHDYFIELLSRSLIQQSNDD-GKEKFVMHDLVNDLALVVSGTSCFRLE 483
Query: 127 SGGRISEDVHHVTYNQEEYDIFNKFE 152
GG +S++V H +YNQ YD KFE
Sbjct: 484 CGGNMSKNVRHFSYNQGVYDFLKKFE 561
>AW774198
Length = 554
Score = 182 bits (461), Expect = 6e-46
Identities = 96/186 (51%), Positives = 121/186 (64%), Gaps = 2/186 (1%)
Frame = +2
Query: 274 PMQIVRLENLQTLTVFLVGKQKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLK 333
P+QI +LENL TL+ F++ K GL + ELGKFP+L GKL I LQN D SEA ANL
Sbjct: 5 PVQIAKLENLHTLSDFVISKHNGGLKVAELGKFPHLHGKLYISQLQNVNDPSEAFQANLN 184
Query: 334 HKVHLEELEVYWDQQTE--ESPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFS 391
K + EL + WD + +S + +L LQPS NLK L+IK YGG SFP+WLGD F
Sbjct: 185 TKERIHELALEWDCGSTFLDSQAHSAVLKHLQPSTNLKSLTIKGYGGTSFPNWLGDNVFG 364
Query: 392 NMVYLSIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPS 451
NM+YL I +C C+ LP LGQ+ LKEL ID + ++++G EFYG G + FQPFPS
Sbjct: 365 NMLYLRISNCVNCLWLPSLGQLGNLKELVIDSLLSIKSVGTEFYGNDG---HPSFQPFPS 535
Query: 452 LEKLEF 457
LE L F
Sbjct: 536 LETLHF 553
>AW688499 similar to GP|4234953|gb|A NBS-LRR-like protein cD7 {Phaseolus
vulgaris}, partial (2%)
Length = 677
Score = 155 bits (393), Expect(2) = 2e-44
Identities = 89/184 (48%), Positives = 116/184 (62%), Gaps = 2/184 (1%)
Frame = +1
Query: 407 LPPLGQVPFLKELKIDGMSRVETIGPEFY--GMTGGSTNSPFQPFPSLEKLEFNSMPSWR 464
LPPLG++P LK L+I M +ETIGPEFY + GS++S FQPFPSLE ++F+++P+W
Sbjct: 43 LPPLGKLPSLKNLEICDMEMLETIGPEFYYVQIEEGSSSS-FQPFPSLECIKFDNIPNWN 219
Query: 465 EWISFRGSKFPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSS 524
EWI F G KF FPRL+ + LR+C +L+GHLPSHLP IE+I I T TLHWLSS
Sbjct: 220 EWIPFEGIKFAFPRLRAMELRNCPKLKGHLPSHLPCIEEIEIE--GRLLETGPTLHWLSS 393
Query: 525 VKSLDLMCQGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSL 584
+K + + N L+ + L S+P + M STCL HL L +SSL
Sbjct: 394 IKKVKI------------NGLRAMLEKCVM-----LSSMPKLIMRSTCLTHLALYSLSSL 522
Query: 585 TAFP 588
TAFP
Sbjct: 523 TAFP 534
Score = 43.1 bits (100), Expect(2) = 2e-44
Identities = 21/31 (67%), Positives = 25/31 (79%)
Frame = +2
Query: 587 FPANGLPTSLQSLRIDECQNLAFLRPETWSN 617
FP++GLPTSLQSL I +NL+FL PET SN
Sbjct: 530 FPSSGLPTSLQSLNILXGENLSFLPPETXSN 622
>AW694719 similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (6%)
Length = 511
Score = 177 bits (448), Expect = 2e-44
Identities = 83/123 (67%), Positives = 101/123 (81%)
Frame = +1
Query: 6 AILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGF 65
+ILNS IWN+ NDNI+P+L L+YQ+LPSHLKRCFAYCSIFPK P ++K+L+LLWMAEGF
Sbjct: 130 SILNSSIWNLRNDNILPALHLSYQYLPSHLKRCFAYCSIFPKDCPLDKKQLVLLWMAEGF 309
Query: 66 LEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCCRF 125
L+ S GK +EE+GDD F ELLSRSLI++ +DD EKFVMHD++ DLAT SGKSCCR
Sbjct: 310 LDCSQGGKKLEELGDDCFAELLSRSLIQQLSDDDRGEKFVMHDLINDLATFVSGKSCCRL 489
Query: 126 GSG 128
G
Sbjct: 490 ECG 498
>BG587633 similar to GP|20385442|gb| resistance gene analog {Vitis vinifera},
partial (22%)
Length = 677
Score = 176 bits (446), Expect = 4e-44
Identities = 96/183 (52%), Positives = 123/183 (66%), Gaps = 2/183 (1%)
Frame = +3
Query: 6 AILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGF 65
+ILNSDIWN+ NDNI+P+L L+YQ+LP HLKRCFAYCSIFPK YP +RK+L+LLWMAEGF
Sbjct: 135 SILNSDIWNLSNDNILPALHLSYQYLPCHLKRCFAYCSIFPKDYPLDRKQLVLLWMAEGF 314
Query: 66 LEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCCRF 125
L+ S GKA+EE+GDD F ELLSRSLI++ ++D EKFVMHD+V DLAT+ SG+SC
Sbjct: 315 LDCSHGGKAMEELGDDCFAELLSRSLIQQLSNDARGEKFVMHDLVNDLATVISGQSCFNL 494
Query: 126 GSGGRISEDV-HHVTYNQEEYD-IFNKFETFFDFKCLRSFLPIGSRLQESYLSCKVIDDL 183
+ V H+ N + + T C S+L R + K +DDL
Sbjct: 495 DVVTSLKGFVMFHIIKNCMTFS*SLRNYST--SKSCEASYLFTPQRRMINTYP*KWVDDL 668
Query: 184 IPS 186
+PS
Sbjct: 669 LPS 677
>TC93462
Length = 814
Score = 112 bits (281), Expect(2) = 1e-43
Identities = 57/105 (54%), Positives = 73/105 (69%), Gaps = 2/105 (1%)
Frame = +1
Query: 332 LKHKVHLEELEVYWD--QQTEESPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCS 389
+K K L+EL + W+ + S + V+L L+PS NLK L+IK YGGISF +WLGD
Sbjct: 1 VKMKEQLDELALEWNCCSTSSNSQSQSVVLEHLRPSTNLKNLTIKGYGGISFSNWLGDSL 180
Query: 390 FSNMVYLSIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEF 434
F NMVYL I SC++C+ LPPLGQ+ LK+L I+GM VETIG EF
Sbjct: 181 FRNMVYLRISSCDHCLWLPPLGQLGNLKKLIIEGMQSVETIGVEF 315
Score = 83.6 bits (205), Expect(2) = 1e-43
Identities = 56/183 (30%), Positives = 78/183 (42%), Gaps = 1/183 (0%)
Frame = +2
Query: 444 SPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFPRLKTLMLRDCTELR-GHLPSHLPSIE 502
S FQPFPSLE L F M W EW G+ FP LKTL L C +LR G++ PS+
Sbjct: 332 SSFQPFPSLETLHFEDMQEWEEWNLIEGTTTEFPSLKTLSLSKCPKLRVGNIADKFPSLT 511
Query: 503 KITILWCNHFPATLSTLHWLSSVKSLDLMCQGSPELSLLGNDSPCHLQVSTIFGFNKLLS 562
++ + C L V ++ ++L
Sbjct: 512 ELEL--------------------------------------RECPLLVQSVRSSGRVLR 577
Query: 563 LPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLV 622
+ + CLQ L + FP +GLP +L+ L+I C+NL FL E +YTSL
Sbjct: 578 --QLMLPLNCLQQLTIDGFPFPVCFPTDGLPKTLKFLKISNCENLEFLPHEYLDSYTSLE 751
Query: 623 TLE 625
L+
Sbjct: 752 ELK 760
Score = 33.1 bits (74), Expect = 0.49
Identities = 31/93 (33%), Positives = 38/93 (40%), Gaps = 12/93 (12%)
Frame = +2
Query: 790 TLKNLSFKCCSTLET--CKDFFPSFLKSLVFINCPKLMSLPD---------MFP-SSLET 837
+LK LS C L D FPS L L CP L+ M P + L+
Sbjct: 434 SLKTLSLSKCPKLRVGNIADKFPS-LTELELRECPLLVQSVRSSGRVLRQLMLPLNCLQQ 610
Query: 838 LEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLK 870
L D P P G P +LK L IS+C L+
Sbjct: 611 LTIDGFPFPVCFPTDGLPKTLKFLKISNCENLE 709
>BG649127 weakly similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (6%)
Length = 688
Score = 164 bits (415), Expect = 1e-40
Identities = 102/228 (44%), Positives = 139/228 (60%), Gaps = 4/228 (1%)
Frame = +2
Query: 43 SIFPKGYPFNRK-KLILLWMAEGFLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVK 101
SIFPK N+K LI LWMAEG L K +E+V ++ F LLSRS +S
Sbjct: 8 SIFPKD*--NKKWNLIYLWMAEGILPQQRTDKRMEDVREECFEVLLSRSFFYQSTYHA-- 175
Query: 102 EKFVMHDVVYDLATIASGKSCCRFGSGG--RISEDVHHVTYNQEEYDIFNKFETFFDFKC 159
++MHD+++D+A +G+ C +I+ V H++Y Q YD KFE F +FK
Sbjct: 176 SHYMMHDLIHDVAQFVAGEFCYNLDDNNPRKITTIVRHLSYLQGIYDDPEKFEIFSEFKQ 355
Query: 160 LRSFLPIG-SRLQESYLSCKVIDDLIPSIKRLRMLSLSNYNITVLPNSINKLVQLRYLNL 218
LR+F+P S S ++ L+P +KRLR+LSLS+Y IT L +SI L+ +RYL+L
Sbjct: 356 LRTFIPFKFSYFVYSSSITSMVSILLPKLKRLRVLSLSHYPITNLSDSIGVLMHMRYLDL 535
Query: 219 SHTDIKCLPDTTCDLYYLQTLLLSGCWKLIELPIHVGKLINLRHLDIS 266
S+T I+CLPD+ LY L+TLLLSGC L LP ++ LINLR LDIS
Sbjct: 536 SYTGIECLPDSVSTLYNLETLLLSGCRCLTILPENMSNLINLRQLDIS 679
>BF634887 weakly similar to GP|10280097|em nucleotide sequence of the I-2
resistance gene {Fusarium oxysporum}, partial (3%)
Length = 694
Score = 163 bits (413), Expect = 2e-40
Identities = 98/242 (40%), Positives = 140/242 (57%), Gaps = 3/242 (1%)
Frame = +1
Query: 271 KKMPMQIVRLENLQTLTVFLVGKQKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDA 330
KKMP QI L +LQTL+ F+VG++ G +I++LG L+GKLCI L++ I+ +A A
Sbjct: 1 KKMPKQIGSLNHLQTLSHFVVGEEN-GSNIQQLGNLNCLQGKLCISGLEHVINPKDAAGA 177
Query: 331 NLKHKVHLEELEVYWDQ--QTEESPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDC 388
NLK K H+EEL + + + + + L P+ NLK+L I Y G +FP+W+ C
Sbjct: 178 NLKDKKHVEELNMEYSDNFKFNNNGRESNVFEALHPNSNLKRLYIGRYQGNNFPNWITGC 357
Query: 389 SFSNMVYLSIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQP 448
N+V L +K C C +PP+ Q+P LKE I + ++ I EFYG STN F+
Sbjct: 358 HLPNLVSLQLKGCG-CSHMPPIEQLPSLKEFSISNCNGIKIITEEFYG--NSSTNVSFR- 525
Query: 449 FPSLEKLEFNSMPSWREWISFRGSKFPFPRLKTLMLRDCTELR-GHLPSHLPSIEKITIL 507
SLE L+F M +W EW G FP LK + + +C +L+ LP HLPSI+K+ I
Sbjct: 526 --SLEVLKFEEMNNWEEWFCPEG----FPLLKEIYIWNCPKLKXALLPQHLPSIQKLKIC 687
Query: 508 WC 509
C
Sbjct: 688 DC 693
>TC90409 weakly similar to GP|10280097|emb|CAC09980. nucleotide sequence of
the I-2 resistance gene {Fusarium oxysporum}, partial
(5%)
Length = 1136
Score = 152 bits (383), Expect = 7e-37
Identities = 122/355 (34%), Positives = 174/355 (48%), Gaps = 28/355 (7%)
Frame = +3
Query: 358 ILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLK 417
+ LQP+ NL +L I Y G SFP W+ C N+V L ++SC C+ LPPLGQ+P LK
Sbjct: 57 VFEALQPNNNLNRLYISQYKGKSFPKWIRGCHLPNLVSLKLQSCGSCLHLPPLGQLPCLK 236
Query: 418 ELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFP 477
EL I ++ IG EF+G NS PF SLE L+F M SW EW+ G FP
Sbjct: 237 ELAICDCHGIKIIGEEFHG-----NNSTNVPFLSLEVLKFVKMNSWEEWLCLEG----FP 389
Query: 478 RLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLM-CQGSP 536
LK L ++ C ELR LP HLPS++K+ I+ C A++ ++ LDL C
Sbjct: 390 LLKELSIKSCPELRSALPQHLPSLQKLEIIDCELLEASIPK---GDNIIELDLQRCD--- 551
Query: 537 ELSLLGNDSPCHLQVSTIFGFN--KLLSLPNMFMSSTCLQHLDLIYISSLTAFPAN-GLP 593
+L N+ P L+ +F N S+ + +++T L+ L +I S+ +
Sbjct: 552 --HILINELPTSLK-RFVFRENWFAKFSVEQILINNTILEELKFDFIGSVKCLSLDLRCY 722
Query: 594 TSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPV-LQILSIEGC 652
+SL+ L I + + P +T+L +L+L N C L SF G P L+ L I C
Sbjct: 723 SSLRDLSITGWHSSSL--PLELHLFTNLHSLKLYN-CPRLDSFPNGGLPSNLRGLVIWNC 893
Query: 653 ---------------SSLKSIFISEK--------NSSLSLSTLQSLKVSNCKSLR 684
+SLKS F+S++ SL TL L ++NC LR
Sbjct: 894 PELIALRQEWGLFRLNSLKSFFVSDEFENVESFPEESLLPPTLTYLNLNNCSKLR 1058
Score = 44.3 bits (103), Expect = 2e-04
Identities = 70/303 (23%), Positives = 110/303 (36%), Gaps = 6/303 (1%)
Frame = +3
Query: 573 LQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNLAFLRPETWSN------YTSLVTLEL 626
L L L S P G L+ L I +C + + E N + SL L+
Sbjct: 162 LVSLKLQSCGSCLHLPPLGQLPCLKELAICDCHGIKIIGEEFHGNNSTNVPFLSLEVLKF 341
Query: 627 KNCCDSLTSFQLNGFPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSL 686
L GFP+L+ LSI+ C L+S L +LQ L++ +C+ L +
Sbjct: 342 VKMNSWEEWLCLEGFPLLKELSIKSCPELRSALPQH------LPSLQKLEIIDCELLEAS 503
Query: 687 PQRMDTLFVLKSLTLDKLSLCCEVACLPPKLQFMHIESLGLATPVTEWGFQSLCFLSDLH 746
+ D + L D HI L T + + F+ F
Sbjct: 504 IPKGDNIIELDLQRCD------------------HILINELPTSLKRFVFRENWFAK--- 620
Query: 747 IGGDNIVNTLLKKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCK 806
V + N T + LK + + + L +L +C S+L
Sbjct: 621 ------------------FSVEQILINNTILEELKFDFIGSVKCL-SLDLRCYSSLRDL- 740
Query: 807 DFFPSFLKSLVFINCPKLMSLPDMFPSSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHC 866
S+ + L +F ++L +L+ +CPRL P G PS+L+ L I +C
Sbjct: 741 --------SITGWHSSSLPLELHLF-TNLHSLKLYNCPRLDSFPNGGLPSNLRGLVIWNC 893
Query: 867 PLL 869
P L
Sbjct: 894 PEL 902
>BG587443 weakly similar to PIR|T06405|T064 resistance complex protein I2C-3
- tomato (fragment), partial (13%)
Length = 720
Score = 146 bits (369), Expect = 3e-35
Identities = 99/214 (46%), Positives = 120/214 (55%), Gaps = 4/214 (1%)
Frame = +3
Query: 689 RMDTLFVLKSLTL--DKLSLCCEVACLPPKLQFMHIESLGLATPVTEWGFQSLCFLSDLH 746
+MD L L+ L + KLS C E CLPPKLQ + S + PVTEWG Q L LS L
Sbjct: 39 KMDMLTALEKLHMKCQKLSFC-EGVCLPPKLQSIWFSSRRITPPVTEWGLQYLTALSLLT 215
Query: 747 IG-GDNIVNTLLKKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETC 805
I GD+I NTL+K+ LLP LV L IT+L+EM GN L+H+S+L+ L F C LET
Sbjct: 216 IQKGDDIFNTLMKESLLPISLVYLYITDLSEMKSFDGNGLRHLSSLQTLCFWFCDQLETL 395
Query: 806 -KDFFPSFLKSLVFINCPKLMSLPDMFPSSLETLEFDDCPRLGLLPRSGFPSSLKLLSIS 864
++ PS LKSL C KL SLP+ P SLK L I
Sbjct: 396 PENCLPSSLKSLDLWKCEKLESLPE----------------------DSLPDSLKQLRIR 509
Query: 865 HCPLLKSRYATKRNDHLSKISHIPVVKINGIVTI 898
CPLL+ RY KR +H SKI+HIPV+ IN VTI
Sbjct: 510 ECPLLEERY--KRKEHWSKIAHIPVIDINDEVTI 605
Score = 38.5 bits (88), Expect = 0.012
Identities = 26/82 (31%), Positives = 42/82 (50%), Gaps = 2/82 (2%)
Frame = +3
Query: 549 LQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNL- 607
LQ + ++L +LP + S+ L+ LDL L + P + LP SL+ LRI EC L
Sbjct: 351 LQTLCFWFCDQLETLPENCLPSS-LKSLDLWKCEKLESLPEDSLPDSLKQLRIRECPLLE 527
Query: 608 -AFLRPETWSNYTSLVTLELKN 628
+ R E WS + +++ +
Sbjct: 528 ERYKRKEHWSKIAHIPVIDIND 593
>AW692807
Length = 516
Score = 134 bits (338), Expect = 1e-31
Identities = 87/172 (50%), Positives = 106/172 (61%), Gaps = 5/172 (2%)
Frame = +2
Query: 589 ANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILS 648
A GLPTSLQSL I C+NL+FL PETWSNYTSLV L+L CD+LTSF L+GFP LQ L
Sbjct: 2 ARGLPTSLQSLNILWCENLSFLPPETWSNYTSLVRLDLCQSCDALTSFPLDGFPALQTLW 181
Query: 649 IEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLR--SLPQRMDTLFVLKSLTLDKLSL 706
I+ C SL SI I E S S S L+ L + + S+ + +MD L L+ L L L
Sbjct: 182 IQNCRSLVSICILESPSCQS-SRLEELVIRSHDSIELFEVKLKMDMLTALEKLILRCAQL 358
Query: 707 C-CEVACLPPKLQFMHIESLGLATPVTEWGFQSL-CFLSDLHI-GGDNIVNT 755
CE CLPPKLQ + I S + PVTEWG ++ +L I GD+I NT
Sbjct: 359 SFCEGVCLPPKLQTIVISSQRITPPVTEWGPSNI*QHFPNLSIEKGDDIFNT 514
>BF635265 similar to GP|4234955|gb|A NBS-LRR-like protein cD8 {Phaseolus
vulgaris}, partial (3%)
Length = 673
Score = 132 bits (333), Expect = 5e-31
Identities = 74/180 (41%), Positives = 99/180 (54%)
Frame = +1
Query: 337 HLEELEVYWDQQTEESPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYL 396
H E + DQ E + +L LQP+INL L+IK Y G SFP+WLGD N+V L
Sbjct: 43 HTMNGEKWMDQ*XEAQAS---VLEALQPNINLTSLTIKDYRGGSFPNWLGDRHLPNLVSL 213
Query: 397 SIKSCEYCITLPPLGQVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLE 456
+ C+ LPPLGQ P LK+ I +E IG EF G NS PF SLE L
Sbjct: 214 ELLGCKIHSQLPPLGQFPSLKKCSISSCDGIEIIGTEFLGY-----NSSDVPFRSLETLR 378
Query: 457 FNSMPSWREWISFRGSKFPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATL 516
F +M W+EW+ G FP L+ L ++ C +L+ LP HLPS++K+ I+ C A++
Sbjct: 379 FENMAEWKEWLCLEG----FPLLQKLCIKHCPKLKSALPQHLPSLQKLEIIDCQELAASI 546
Score = 40.0 bits (92), Expect = 0.004
Identities = 35/123 (28%), Positives = 53/123 (42%), Gaps = 7/123 (5%)
Frame = +1
Query: 573 LQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNLAFLRPETWSN------YTSLVTLEL 626
L L+L+ + P G SL+ I C + + E + SL TL
Sbjct: 202 LVSLELLGCKIHSQLPPLGQFPSLKKCSISSCDGIEIIGTEFLGYNSSDVPFRSLETLRF 381
Query: 627 KNCCDSLTSFQLNGFPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSL-RS 685
+N + L GFP+LQ L I+ C LKS L +LQ L++ +C+ L S
Sbjct: 382 ENMAEWKEWLCLEGFPLLQKLCIKHCPKLKSALPQH------LPSLQKLEIIDCQELAAS 543
Query: 686 LPQ 688
+P+
Sbjct: 544 IPK 552
Score = 31.2 bits (69), Expect = 1.8
Identities = 23/58 (39%), Positives = 32/58 (54%), Gaps = 2/58 (3%)
Frame = +1
Query: 805 CKDFFPSFLKSLVFINCPKLMS-LPDMFPSSLETLEFDDCPRLGL-LPRSGFPSSLKL 860
C + FP L+ L +CPKL S LP P SL+ LE DC L +P++ + L+L
Sbjct: 412 CLEGFP-LLQKLCIKHCPKLKSALPQHLP-SLQKLEIIDCQELAASIPKAANITELEL 579
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,121,612
Number of Sequences: 36976
Number of extensions: 582792
Number of successful extensions: 3930
Number of sequences better than 10.0: 219
Number of HSP's better than 10.0 without gapping: 3526
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3796
length of query: 898
length of database: 9,014,727
effective HSP length: 105
effective length of query: 793
effective length of database: 5,132,247
effective search space: 4069871871
effective search space used: 4069871871
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC119417.2


