

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
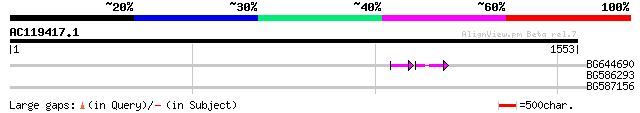
Query= AC119417.1 + phase: 0 /pseudo
(1553 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 48 7e-09
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 42 0.002
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 33 0.86
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 48.1 bits (113), Expect(2) = 7e-09
Identities = 36/98 (36%), Positives = 51/98 (51%), Gaps = 6/98 (6%)
Frame = -1
Query: 1111 SRLQSTRRH*LH*NICSSCKIGS----NQVTSILRN*SWHNI--ISNGCQKCLS*WCH*R 1164
+R+QS RR+ L * + C+ GS N I H + + NGC++C+ *W R
Sbjct: 395 ARIQSKRRNRL**GFFTCCQNGSY*NFNSFCCI------HGVQAVPNGCEECIY*WRSQR 234
Query: 1165 RSVC*TTSWV*GS*AS*PCL*T*EITIWLETSSQSLV* 1202
VC TSW+* + C+ TIW E SS+S+V*
Sbjct: 233 GGVCQATSWI*RCRGTKSCVQIE*DTIWSEASSKSMV* 120
Score = 31.6 bits (70), Expect(2) = 7e-09
Identities = 24/62 (38%), Positives = 32/62 (50%)
Frame = -2
Query: 1043 *A*NC*RSSLR*WMDISYARRAKSIPKE*CVGSGTQTLSEEHYWNKMGIQKQAE*TMRSN 1102
*A C*RS +D YARR S+ K+* + G+ T + WN +G KQA R+
Sbjct: 589 *AQEC*RSIA*CRLDQFYARRTPSV*KK*GMVPGSST*RQNSNWN*VGF*KQA*VITRNK 410
Query: 1103 QK 1104
K
Sbjct: 409 SK 404
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 42.0 bits (97), Expect = 0.002
Identities = 25/75 (33%), Positives = 38/75 (50%), Gaps = 2/75 (2%)
Frame = +1
Query: 1096 E*TMRSNQKQSQTCCSRLQSTRRH*LH*NICSSCKIGSNQVTSILRN*SWHNIISNGCQK 1155
E* +Q QS+ C RL+ T RH L ++C+SC ++ T + + W S+ C+
Sbjct: 127 E*RWNVDQIQSKASCKRLRETTRHRLRRSVCTSCSNRNHMTTLGVSSN*WMLDPSHRCKN 306
Query: 1156 CLS*W--CH*RRSVC 1168
C+ W C R S C
Sbjct: 307 CIPKWTLCGNRNSHC 351
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 33.1 bits (74), Expect = 0.86
Identities = 18/56 (32%), Positives = 31/56 (55%)
Frame = -3
Query: 1103 QKQSQTCCSRLQSTRRH*LH*NICSSCKIGSNQVTSILRN*SWHNIISNGCQKCLS 1158
++++ T R+ S LH*+IC+S + N L W I++NGC++C+S
Sbjct: 382 EEEN*TSSKRVYSDIWRGLH*DICTSSQATHN*NCFKLGCEPWMGIVANGCEECIS 215
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.386 0.173 0.716
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,517,325
Number of Sequences: 36976
Number of extensions: 652283
Number of successful extensions: 11868
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1369
Number of HSP's successfully gapped in prelim test: 222
Number of HSP's that attempted gapping in prelim test: 10323
Number of HSP's gapped (non-prelim): 1966
length of query: 1553
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1444
effective length of database: 4,984,343
effective search space: 7197391292
effective search space used: 7197391292
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 34 (21.5 bits)
S2: 65 (29.6 bits)
Medicago: description of AC119417.1


