

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.8 + phase: 0 /pseudo
(427 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
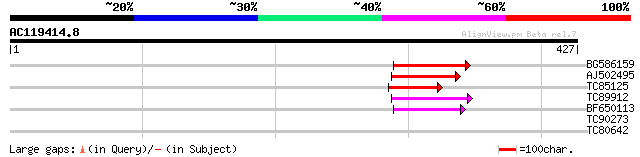
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 80 2e-15
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 68 6e-12
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 47 2e-05
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 46 2e-05
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 46 3e-05
TC90273 similar to GP|22136764|gb|AAM91701.1 unknown protein {Ar... 28 6.8
TC80642 similar to GP|18176376|gb|AAL60033.1 unknown protein {Ar... 28 8.8
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 79.7 bits (195), Expect = 2e-15
Identities = 34/58 (58%), Positives = 47/58 (80%)
Frame = +1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
KSTS YV+M+ +GA+SWSS K+P+VTLSTT+AEF+AAA C+C +W+RR+ +G Q
Sbjct: 409 KSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQSVWMRRVLEKLGYTQ 582
Score = 38.5 bits (88), Expect = 0.005
Identities = 27/79 (34%), Positives = 45/79 (56%), Gaps = 2/79 (2%)
Frame = +3
Query: 156 AR*SRNFHRSKQVCY*NIDKAWYGKLQYGV*PYSHW*KTSEG*N*EGC--GCNQV*ADGW 213
++* RN + K++C * + K W+G+ + Y + * *+ C GC +V*A+ W
Sbjct: 12 SK*GRNLYLPKEICN*LVRKVWHGEK*FV*ESYCS*VQVD--*R*KWCEGGCYKV*ANCW 185
Query: 214 MLDVITCYKT*SYIFSLSN 232
+ DV++ +KT*S + SN
Sbjct: 186 LSDVLSSHKT*SNVCVKSN 242
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 68.2 bits (165), Expect = 6e-12
Identities = 30/52 (57%), Positives = 39/52 (74%)
Frame = +2
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRI 339
T KSTS Y + +G GAISWSS K+P+V ST EAE++A+ SC+ +WLRRI
Sbjct: 26 TRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 46.6 bits (109), Expect = 2e-05
Identities = 21/41 (51%), Positives = 29/41 (70%)
Frame = +1
Query: 286 HMTEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAA 326
H KST+ YV+ + GA+SW S + +V LSTTEAE++AA
Sbjct: 109 HDKRKSTTGYVFTLAGGAVSWLSKLQTVVALSTTEAEYMAA 231
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 46.2 bits (108), Expect = 2e-05
Identities = 23/61 (37%), Positives = 36/61 (58%)
Frame = +1
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
T KS S +V+ + ISW + ++ +VTLSTT+AE++A IWL+ + +G Q
Sbjct: 130 TRKSLSGFVFTLYGTTISWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQ 309
Query: 348 E 348
E
Sbjct: 310 E 312
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 45.8 bits (107), Expect = 3e-05
Identities = 20/54 (37%), Positives = 32/54 (59%)
Frame = +1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
+STS YV+ + AISW + K+PI LS+ EAE++A + +WL + +
Sbjct: 319 RSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVIKEL 480
>TC90273 similar to GP|22136764|gb|AAM91701.1 unknown protein {Arabidopsis
thaliana}, partial (25%)
Length = 834
Score = 28.1 bits (61), Expect = 6.8
Identities = 16/40 (40%), Positives = 22/40 (55%), Gaps = 2/40 (5%)
Frame = +1
Query: 275 LDGLTQIIQEIHM--TEKSTSWYVYMVGNGAISWSSMKKP 312
LDGLT I+ +I + + T + VGNG S+ KKP
Sbjct: 37 LDGLTTILMQIEQMFSSEDTQLLISEVGNGIQEDSATKKP 156
>TC80642 similar to GP|18176376|gb|AAL60033.1 unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 1182
Score = 27.7 bits (60), Expect = 8.8
Identities = 18/50 (36%), Positives = 26/50 (52%)
Frame = -1
Query: 126 CV*NFQAFYKGKVCYDRFGKYEIFSWSRS*AR*SRNFHRSKQVCY*NIDK 175
C+ N + + + ++ Y F KY W R R S NF+RSK C+ DK
Sbjct: 360 CLINMR*YCRNQILY--FSKYR---WRR---RLSNNFYRSKVFCFFKRDK 235
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.372 0.166 0.671
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,175,880
Number of Sequences: 36976
Number of extensions: 231827
Number of successful extensions: 2206
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 859
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 1330
Number of HSP's gapped (non-prelim): 958
length of query: 427
length of database: 9,014,727
effective HSP length: 99
effective length of query: 328
effective length of database: 5,354,103
effective search space: 1756145784
effective search space used: 1756145784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC119414.8


