

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
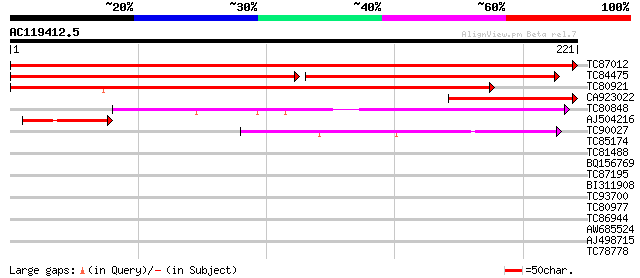
Query= AC119412.5 + phase: 0
(221 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87012 similar to GP|9294029|dbj|BAB01986.1 vesicle transport v... 355 6e-99
TC84475 similar to GP|19571103|dbj|BAB86528. putative vesicle tr... 164 4e-76
TC80921 similar to GP|9295720|gb|AAF87026.1| T24P13.5 {Arabidops... 250 3e-67
CA923022 similar to GP|19571103|dbj putative vesicle transport v... 71 4e-13
TC80848 similar to GP|9758926|dbj|BAB09463.1 golgi SNARE protein... 46 1e-05
AJ504216 similar to GP|9294029|dbj| vesicle transport v-SNARE (v... 45 2e-05
TC90027 similar to GP|4206789|gb|AAD11809.1| syntaxin-related pr... 41 3e-04
TC85174 homologue to SP|P30075|CHS4_MEDSA Chalcone synthase 4 (E... 30 0.56
TC81488 GP|17985051|gb|AAL54093.1 EXOPOLYSACCHARIDE PRODUCTION P... 30 0.73
BQ156769 homologue to GP|13750761|em ORF2 {Rhodococcus erythropo... 29 1.6
TC87195 similar to GP|16604629|gb|AAL24107.1 putative bZIP prote... 28 3.6
BI311908 similar to GP|10177105|d gene_id:MXC20.14~pir||T13959~s... 27 4.8
TC93700 27 4.8
TC80977 similar to GP|19310773|gb|AAL85117.1 unknown protein {Ar... 27 4.8
TC86944 similar to GP|15215806|gb|AAK91448.1 At3g14201 {Arabidop... 27 4.8
AW685524 homologue to GP|15982737|gb| AT4g23750/F9D16_220 {Arabi... 27 4.8
AJ498715 similar to PIR|A96772|A9677 hypothetical protein F1M20.... 27 8.1
TC78778 weakly similar to PIR|S71277|S71277 serine/threonine-spe... 27 8.1
>TC87012 similar to GP|9294029|dbj|BAB01986.1 vesicle transport v-SNARE
(vesicle soluble NSF attachment protein receptor)
protein, complete
Length = 1079
Score = 355 bits (912), Expect = 6e-99
Identities = 180/221 (81%), Positives = 200/221 (90%)
Frame = +2
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS VFEGYERQYCELS+ LSK CTAA ALNGEQKKQ +S++K+GIDEAEALIRKMDLEAR
Sbjct: 203 MSTVFEGYERQYCELSSKLSKLCTAAAALNGEQKKQNISDIKAGIDEAEALIRKMDLEAR 382
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
S+QPNIKGVLLAKLREYKSDLNN KSE+KK+VSGN+NPSARD LLESGMAD MT SADQR
Sbjct: 383 SLQPNIKGVLLAKLREYKSDLNNNKSELKKLVSGNMNPSARDMLLESGMADAMTVSADQR 562
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM STERLNK+ +R+KDSRRTMLETEELGVSILQDLHSQRQSLLHAH+TLHGVDDN+
Sbjct: 563 DRLMFSTERLNKSSDRIKDSRRTMLETEELGVSILQDLHSQRQSLLHAHDTLHGVDDNVS 742
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI++NMS+RMN+NK II IV VLV+ I+ LYFKL K
Sbjct: 743 KSKKILSNMSKRMNRNKCIISTIVAVLVLVIMLTLYFKLSK 865
>TC84475 similar to GP|19571103|dbj|BAB86528. putative vesicle transport
v-SNARE protein {Oryza sativa (japonica
cultivar-group)}, partial (85%)
Length = 762
Score = 164 bits (414), Expect(2) = 4e-76
Identities = 78/113 (69%), Positives = 101/113 (89%)
Frame = +3
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS VF+GYERQYCELSA LS++CTAA+AL+GEQKKQK+SE+K+G+++A+ LIRKMDLEAR
Sbjct: 114 MSEVFDGYERQYCELSAILSRQCTAASALDGEQKKQKLSEIKAGLEDADTLIRKMDLEAR 293
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTM 113
S+QP++K LLAKLREYK+DLNNLK+EVK++ S N + +ARDELLE G D++
Sbjct: 294 SLQPSMKATLLAKLREYKTDLNNLKNEVKRVASANASITARDELLELGRIDSL 452
Score = 138 bits (347), Expect(2) = 4e-76
Identities = 66/99 (66%), Positives = 85/99 (85%)
Frame = +2
Query: 116 SADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGV 175
S DQ+ RL+ +TERLN++ +R+ +SR+T+LETEELGVSILQDLH QRQSLLHAH +LHGV
Sbjct: 464 SNDQKGRLLMTTERLNQSTDRLTNSRKTLLETEELGVSILQDLHQQRQSLLHAHKSLHGV 643
Query: 176 DDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAI 214
DDNI KSKKI+ MS+RM++NKWI+G ++ LV+AII I
Sbjct: 644 DDNISKSKKILAAMSKRMSRNKWIVGSLMAALVLAIILI 760
>TC80921 similar to GP|9295720|gb|AAF87026.1| T24P13.5 {Arabidopsis
thaliana}, partial (79%)
Length = 666
Score = 250 bits (638), Expect = 3e-67
Identities = 123/190 (64%), Positives = 159/190 (82%), Gaps = 1/190 (0%)
Frame = +1
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKK-QKVSEVKSGIDEAEALIRKMDLEA 59
MS VFEGYERQYC+LSANLS+KCT+ + L+ +++K QK SE+K+G+D+A+ LIRKMDLEA
Sbjct: 97 MSQVFEGYERQYCDLSANLSRKCTSTSLLSDQEEKLQKFSEIKTGLDDADVLIRKMDLEA 276
Query: 60 RSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQ 119
RS+QP++K +LLAKLREYKSDLNNLK E K++ S + +ARD+LLE+G ADT ASADQ
Sbjct: 277 RSLQPSVKAMLLAKLREYKSDLNNLKKEFKRLTSPTADQAARDDLLETGRADTHLASADQ 456
Query: 120 RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNI 179
R RL S ERLN++ +R+++S RT+LETEELGVSILQDLH QR++LL +H LHGVD+ I
Sbjct: 457 RERLTMSVERLNQSSDRIRESHRTVLETEELGVSILQDLHQQRETLLSSHKRLHGVDNAI 636
Query: 180 GKSKKIMTNM 189
KSKK++T M
Sbjct: 637 DKSKKVLTXM 666
>CA923022 similar to GP|19571103|dbj putative vesicle transport v-SNARE
protein {Oryza sativa (japonica cultivar-group)},
partial (21%)
Length = 324
Score = 70.9 bits (172), Expect = 4e-13
Identities = 34/50 (68%), Positives = 41/50 (82%)
Frame = -3
Query: 172 LHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
LHGVDDN+ KSKKI++NMS+RMN+NK II IV VLV+ I+ LYFKL K
Sbjct: 238 LHGVDDNVSKSKKILSNMSKRMNRNKCIISTIVAVLVLVIMLTLYFKLSK 89
>TC80848 similar to GP|9758926|dbj|BAB09463.1 golgi SNARE protein
{Arabidopsis thaliana}, partial (24%)
Length = 934
Score = 45.8 bits (107), Expect = 1e-05
Identities = 43/185 (23%), Positives = 83/185 (44%), Gaps = 7/185 (3%)
Frame = +1
Query: 41 VKSGIDEAEALIRKMDLEARSMQPNIKGVLL-AKLREYKSDLNNLKSEVKKIVSGN---- 95
V I + ++L +M +RS+ + L K+ + + +L+ ++K S N
Sbjct: 232 VSKDITQIQSLCVEMSQLSRSIGAKSQRDLWNRKVEQIAEEAESLRESLEKYNSRNHKRM 411
Query: 96 LNPSARDELLE--SGMADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVS 153
+ R+ELL +G + D T+ M S V++S R + LG +
Sbjct: 412 MEAKEREELLGRVNGDPSHVLRIFDDDTQAMLS----------VRNSARELENANALGET 561
Query: 154 ILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIA 213
IL +H QR+ L AH V + G S +++ + RR ++WI +++ V+ + A
Sbjct: 562 ILSSIHGQRERLKSAHRKALDVLNTAGISNRVLRLIERRNRVDQWIKYAGMILTVIFLFA 741
Query: 214 ILYFK 218
+ ++
Sbjct: 742 FVLWR 756
>AJ504216 similar to GP|9294029|dbj| vesicle transport v-SNARE (vesicle
soluble NSF attachment protein receptor) protein,
partial (13%)
Length = 138
Score = 45.1 bits (105), Expect = 2e-05
Identities = 23/35 (65%), Positives = 27/35 (76%)
Frame = +3
Query: 6 EGYERQYCELSANLSKKCTAATALNGEQKKQKVSE 40
EGYERQYCEL + +K + +LNGEQKKQKVSE
Sbjct: 36 EGYERQYCELLL-ICQKNVLSYSLNGEQKKQKVSE 137
>TC90027 similar to GP|4206789|gb|AAD11809.1| syntaxin-related protein
At-SYR1 {Arabidopsis thaliana}, partial (82%)
Length = 1362
Score = 41.2 bits (95), Expect = 3e-04
Identities = 26/130 (20%), Positives = 61/130 (46%), Gaps = 5/130 (3%)
Frame = +3
Query: 91 IVSGNLNPSARDELLESGMADTMTASADQ---RTRLMTSTERLNKTGERVKDSRRTMLET 147
+ N + D L+ +G ++T A Q R R++ + + + + VKD +++L
Sbjct: 558 VTGENPDDKTLDLLISTGESETFLQKAIQEQGRGRILDTINEIQERHDAVKDLEKSLLAL 737
Query: 148 EE--LGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVL 205
+ L ++++ ++ + +H + G + ++ T + N KW CI+L
Sbjct: 738 HQVFLDMTVMVQFQGEQLDDIESHVARASSFVHTG-TDQLQTARKHQKNTRKWACYCIIL 914
Query: 206 VLVVAIIAIL 215
+L++ +I +L
Sbjct: 915 LLIIVLIVVL 944
>TC85174 homologue to SP|P30075|CHS4_MEDSA Chalcone synthase 4 (EC 2.3.1.74)
(Naringenin-chalcone synthase 4) (CHS12-1). [Alfalfa],
partial (82%)
Length = 1072
Score = 30.4 bits (67), Expect = 0.56
Identities = 22/67 (32%), Positives = 36/67 (52%)
Frame = +1
Query: 45 IDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDEL 104
I E E I +M A+++ P+ +G + LRE + LK +V +IVS N+N + D
Sbjct: 787 IPEIEKPIFEMVWTAQTIAPDSEGAIDGHLREAGLTFHLLK-DVPEIVSKNINKALVDAF 963
Query: 105 LESGMAD 111
G++D
Sbjct: 964 QPLGVSD 984
>TC81488 GP|17985051|gb|AAL54093.1 EXOPOLYSACCHARIDE PRODUCTION PROTEIN EXOF
PRECURSOR {Brucella melitensis}, partial (3%)
Length = 921
Score = 30.0 bits (66), Expect = 0.73
Identities = 29/107 (27%), Positives = 52/107 (48%)
Frame = -3
Query: 51 LIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMA 110
LI+++ + + + IK V +EY++ L+ E+K++ S N+ +DE+L
Sbjct: 667 LIQELKMLKQLRKLGIKSVR----QEYRTALSAAIQEMKQLESLNIGAITKDEIL----- 515
Query: 111 DTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQD 157
D ASA R R++ RL K + + + L LG+S +D
Sbjct: 514 DLDLASAPSRLRVLNLKCRLTKLPNWIPNLK--SLVKLRLGLSNFED 380
>BQ156769 homologue to GP|13750761|em ORF2 {Rhodococcus erythropolis},
partial (72%)
Length = 683
Score = 28.9 bits (63), Expect = 1.6
Identities = 9/22 (40%), Positives = 17/22 (76%)
Frame = +2
Query: 164 SLLHAHNTLHGVDDNIGKSKKI 185
S + H +HGVD+++GK+K++
Sbjct: 98 SFVRHHGLVHGVDESVGKAKRV 163
>TC87195 similar to GP|16604629|gb|AAL24107.1 putative bZIP protein
{Arabidopsis thaliana}, partial (38%)
Length = 1412
Score = 27.7 bits (60), Expect = 3.6
Identities = 12/24 (50%), Positives = 18/24 (75%)
Frame = -1
Query: 122 RLMTSTERLNKTGERVKDSRRTML 145
R+ + E+ KTG+R+KDSR+T L
Sbjct: 1115 RIKSLGEQRRKTGQRLKDSRQTCL 1044
>BI311908 similar to GP|10177105|d gene_id:MXC20.14~pir||T13959~similar to
unknown protein {Arabidopsis thaliana}, partial (14%)
Length = 724
Score = 27.3 bits (59), Expect = 4.8
Identities = 31/121 (25%), Positives = 48/121 (39%), Gaps = 13/121 (10%)
Frame = +2
Query: 32 EQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLR--------EYKSD-LN 82
EQK + E S +D +L+RKM + + VL K R EY D L
Sbjct: 350 EQKSRPCEEYASIVDFLNSLVRKMLKKMKKQPLLFVEVLFWKTRRECHYINAEYLLDELG 529
Query: 83 NLKSEVKK----IVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVK 138
+LK+E K G + S +AD + D+ +++ R GE++
Sbjct: 530 HLKNETKNWNDTQGDGEIGSSPGKPWTRRSLADAL--GDDEADVVISHDSRYQNNGEKLD 703
Query: 139 D 139
D
Sbjct: 704 D 706
>TC93700
Length = 504
Score = 27.3 bits (59), Expect = 4.8
Identities = 19/71 (26%), Positives = 27/71 (37%)
Frame = +2
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
T L LNKT + S T+ +E+ S L + LLH + G
Sbjct: 182 THLFEEIRELNKTNDSASSSSVTVSNSEDSWSSDLVLALQSAKRLLHEAKNFSSNSSSDG 361
Query: 181 KSKKIMTNMSR 191
+KKI+ R
Sbjct: 362 AAKKIIFQFQR 394
>TC80977 similar to GP|19310773|gb|AAL85117.1 unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 911
Score = 27.3 bits (59), Expect = 4.8
Identities = 28/127 (22%), Positives = 53/127 (41%), Gaps = 15/127 (11%)
Frame = +3
Query: 38 VSEVKSGIDEAEALIRKMDLEARSMQPNIKGV---LLAKLREYKSDLNNLKSEVKKIVSG 94
V +K + EA ++ +LE+ + G L E ++ + V + S
Sbjct: 216 VQNLKRSVREAALSLQSYNLESSRNSSHSDGSSEHFFVPLSETSFSHSDTEKNVTSLRSK 395
Query: 95 NLNPSARDE-LLESGMADTMTASADQRTRLMTSTERLNKT-----------GERVKDSRR 142
L S D+ LLES +D + D+ + +++ ERL+ + D+RR
Sbjct: 396 RLFVSPMDDPLLESHASDEHGSKFDEFSDMLSDMERLSYSDNVNGFLSYTGSNETSDARR 575
Query: 143 TMLETEE 149
+M + E+
Sbjct: 576 SMFDFED 596
>TC86944 similar to GP|15215806|gb|AAK91448.1 At3g14201 {Arabidopsis
thaliana}, partial (50%)
Length = 2552
Score = 27.3 bits (59), Expect = 4.8
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = -1
Query: 14 ELSANLSKKCTAATALNGEQKKQKVS 39
+LS N+ KKC LNG +KK+ V+
Sbjct: 2438 KLSNNMYKKCAYCIILNGPKKKKTVN 2361
>AW685524 homologue to GP|15982737|gb| AT4g23750/F9D16_220 {Arabidopsis
thaliana}, partial (21%)
Length = 666
Score = 27.3 bits (59), Expect = 4.8
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = -3
Query: 192 RMNKNKWIIGCIVLVLVVAI 211
R + N W+ GCIV+V+ V I
Sbjct: 259 RYSYNSWVYGCIVVVVTVTI 200
>AJ498715 similar to PIR|A96772|A9677 hypothetical protein F1M20.2
[imported] - Arabidopsis thaliana, complete
Length = 612
Score = 26.6 bits (57), Expect = 8.1
Identities = 10/29 (34%), Positives = 19/29 (65%)
Frame = +1
Query: 10 RQYCELSANLSKKCTAATALNGEQKKQKV 38
R Y E S++C ++ A+NG++ K+K+
Sbjct: 61 RSYKEKVRRKSRRCLSSAAVNGKKAKKKI 147
>TC78778 weakly similar to PIR|S71277|S71277 serine/threonine-specific
receptor protein kinase (EC 2.7.1.-) - Arabidopsis
thaliana, partial (23%)
Length = 1808
Score = 26.6 bits (57), Expect = 8.1
Identities = 21/70 (30%), Positives = 33/70 (47%), Gaps = 4/70 (5%)
Frame = +3
Query: 154 ILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLV----LVV 209
+ QD+ Q L T +I + I + S + + NK I+ + + LV+
Sbjct: 411 VCQDVXKSHQVSLGVILTXQSCFRHIYRVDDINLSDSNKSDTNKAIVPIVATLSSVFLVL 590
Query: 210 AIIAILYFKL 219
IIAILY+KL
Sbjct: 591 LIIAILYWKL 620
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.129 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,919,958
Number of Sequences: 36976
Number of extensions: 48387
Number of successful extensions: 267
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 265
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 265
length of query: 221
length of database: 9,014,727
effective HSP length: 92
effective length of query: 129
effective length of database: 5,612,935
effective search space: 724068615
effective search space used: 724068615
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC119412.5


