

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.1 - phase: 0
(411 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
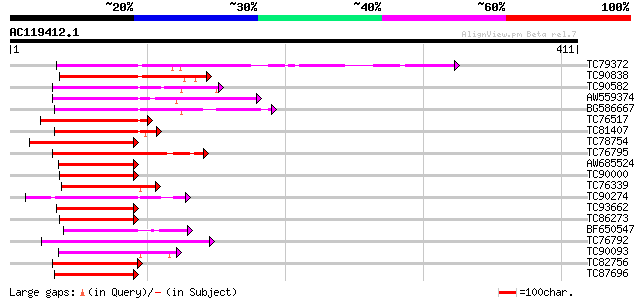
Score E
Sequences producing significant alignments: (bits) Value
TC79372 homologue to GP|2281647|gb|AAC49777.1| AP2 domain contai... 123 1e-28
TC90838 homologue to GP|2281647|gb|AAC49777.1| AP2 domain contai... 121 6e-28
TC90582 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_18... 89 4e-18
AW559374 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabid... 87 1e-17
BG586667 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabid... 84 8e-17
TC76517 similar to GP|11994632|dbj|BAB02769. AP2 domain transcri... 84 1e-16
TC81407 homologue to GP|18650662|gb|AAL75809.1 ethylene response... 82 3e-16
TC78754 homologue to GP|4099921|gb|AAD09248.1| EREBP-3 homolog {... 81 8e-16
TC76795 similar to GP|18496063|emb|CAD21849. ethylene responsive... 81 8e-16
AW685524 homologue to GP|15982737|gb| AT4g23750/F9D16_220 {Arabi... 80 1e-15
TC90000 homologue to GP|9294548|dbj|BAB02811.1 ethylene responsi... 80 1e-15
TC76339 homologue to GP|4099914|gb|AAD00708.1| ethylene-responsi... 80 2e-15
TC90274 similar to GP|10177219|dbj|BAB10294. contains similarity... 79 2e-15
TC93662 similar to GP|10177219|dbj|BAB10294. contains similarity... 79 2e-15
TC86273 similar to SP|O80340|ERF4_ARATH Ethylene responsive elem... 79 2e-15
BF650547 similar to GP|9758388|dbj| AP2 domain transcription fac... 79 4e-15
TC76792 similar to GP|3264767|gb|AAC24587.1| AP2 domain containi... 79 4e-15
TC90093 similar to PIR|T47955|T47955 hypothetical protein F15G16... 78 7e-15
TC82756 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_18... 77 2e-14
TC87696 similar to GP|9759176|dbj|BAB09791.1 contains similarity... 76 2e-14
>TC79372 homologue to GP|2281647|gb|AAC49777.1| AP2 domain containing
protein RAP2.11 {Arabidopsis thaliana}, partial (27%)
Length = 1086
Score = 123 bits (309), Expect = 1e-28
Identities = 106/308 (34%), Positives = 145/308 (46%), Gaps = 16/308 (5%)
Frame = +3
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
R +FVGVRQRPSGR+VAEIKDT Q IR+WLGTF+TAEEAARAYDEAA LLRG+ TRTNF
Sbjct: 12 RDKFVGVRQRPSGRYVAEIKDTTQNIRMWLGTFETAEEAARAYDEAATLLRGSKTRTNF- 188
Query: 95 PCSQSSTSPALSSKITNLLLQR---LKERNM-------------NNSCSSFSSSKYTSLL 138
+ S L+S+I +LL R K+++M N++ S S+S T+ +
Sbjct: 189 -VTHVSYDSPLASRIRHLLNNRKKGTKQQDMNGISSTTSHADTTNDTTSDGSTSSTTNCI 365
Query: 139 GSPEERREQQRANDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNELINDT 198
G+ A+ A G S I + S+N +ND
Sbjct: 366 GTASGAINSTSASGVTSTSTNISTSASGVASTSTDIS------------TNSSNTNVNDK 509
Query: 199 AQFDYISSSFESCLTENNGCKEDEMNMESKSNNVTETSSGDSNSEDGEEEVNNDVSVDVP 258
++ +SSS + + + N ++ S N E+SS SN
Sbjct: 510 SE-SLLSSS--TTMQKPNLFEDAYRPDMSNLTNEYESSSYKSNVS--------------- 635
Query: 259 DFRFLDDIAPTSYYSPFEIAEEMEEQMVENESFCDEPSMIRAVMKRMKYERKFSASLYAF 318
D P PF+ +M + CD + +RMK ER+ SASLYA
Sbjct: 636 -----WDFGPIFDNFPFDQWLDMTN---NDGLLCDMVDKGVSEFERMKVERQISASLYAI 791
Query: 319 NGIPECLK 326
NG+ E +K
Sbjct: 792 NGVQEYMK 815
>TC90838 homologue to GP|2281647|gb|AAC49777.1| AP2 domain containing
protein RAP2.11 {Arabidopsis thaliana}, partial (22%)
Length = 521
Score = 121 bits (303), Expect = 6e-28
Identities = 68/116 (58%), Positives = 84/116 (71%), Gaps = 6/116 (5%)
Frame = +2
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPC 96
+FVGVRQRPSGRWVAEIKDT QKIR+WLGTF+TAEEAARAYDEAACLLRG+NTRTNF
Sbjct: 161 KFVGVRQRPSGRWVAEIKDTTQKIRMWLGTFETAEEAARAYDEAACLLRGSNTRTNF--I 334
Query: 97 SQSSTSPALSSKITNLLLQRLKERNMNNS--CSSFSSSK----YTSLLGSPEERRE 146
+ S L+S+I NLL R ++ + S+ S+SK TS + S ++ +E
Sbjct: 335 THVSLDSPLASRIRNLLNNRKGDKKQEDGAVASAPSNSKTTISNTSTITSNDDNKE 502
>TC90582 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_180
{Arabidopsis thaliana}, partial (29%)
Length = 979
Score = 88.6 bits (218), Expect = 4e-18
Identities = 55/131 (41%), Positives = 73/131 (54%), Gaps = 7/131 (5%)
Frame = +2
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
+R +K++ GVRQRP G+W AEI+D + +RVWLGTF TAEEAARAYD AA RG +
Sbjct: 323 KRAKKKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDNAAIEFRGPRAKL 502
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKERNMN-----NSCSSFSSSKYTSLLGSPEERRE 146
NF P S + ++ + L+ +K+ NMN + F S+K S E
Sbjct: 503 NF-PVVDESLKNVVDPEVV-VPLEDIKDENMNQEIQIETTMGFESNKNCDFWDSIGEADF 676
Query: 147 QQ--RANDHDR 155
QQ R D DR
Sbjct: 677 QQLMRFMDFDR 709
>AW559374 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabidopsis
thaliana}, partial (27%)
Length = 732
Score = 87.0 bits (214), Expect = 1e-17
Identities = 56/153 (36%), Positives = 82/153 (52%), Gaps = 2/153 (1%)
Frame = +3
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
+R +K++ GVRQRP G+W AEI+D + RVWLGTF TAEEAARAYD AA RG +
Sbjct: 282 KRAKKKYRGVRQRPWGKWAAEIRDPRRATRVWLGTFSTAEEAARAYDNAAIEFRGPRAKL 461
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKE--RNMNNSCSSFSSSKYTSLLGSPEERREQQR 149
NF P S S++ L+ +K+ N N + + S+ G+ + R
Sbjct: 462 NF-PMVDDSLK---QSEVVAPDLENVKDENENENENMNQEMQSETNIGFGNNMDCDFWDR 629
Query: 150 ANDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLN 182
+ D Q++ + +D G+ S + + F FLN
Sbjct: 630 LGETDFQQLMRFMDFGGDSSDSRTGNTFN*FLN 728
>BG586667 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabidopsis
thaliana}, partial (28%)
Length = 740
Score = 84.3 bits (207), Expect = 8e-17
Identities = 57/166 (34%), Positives = 83/166 (49%), Gaps = 5/166 (3%)
Frame = +3
Query: 33 RVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
R +K++ GVRQRP G+W AEI+D + +RVWLGTF TAEEAARAYD AA RG + N
Sbjct: 114 RAKKKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDNAAIEFRGPRAKLN 293
Query: 93 FWPCSQSSTSPALSSKITNLLLQRLKERNMN-----NSCSSFSSSKYTSLLGSPEERREQ 147
F P S ++ K+ NMN + + F ++K L S
Sbjct: 294 F-PLVDESLKHVEEPEVIVHSKHVTKDENMNQEMQIETMTGFENNKDCDFLDS------- 449
Query: 148 QRANDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNNE 193
+ D Q+ + ++ G+ S + + F F+N ST+N +
Sbjct: 450 --IGEPDFQQFMKFMEFSGDSSGSRTGNPFN*FMN-----STTNEQ 566
>TC76517 similar to GP|11994632|dbj|BAB02769. AP2 domain transcription
factor RAP2.3 {Arabidopsis thaliana}, partial (33%)
Length = 1290
Score = 83.6 bits (205), Expect = 1e-16
Identities = 43/81 (53%), Positives = 54/81 (66%)
Frame = +3
Query: 23 KEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAAC 82
K S GG R + + G+RQRP G+W AEI+D Q +RVWLGTF TAEEAARAYD AA
Sbjct: 531 KGKKSTGGKRARKNVYRGIRQRPWGKWAAEIRDPQQGVRVWLGTFSTAEEAARAYDTAAK 710
Query: 83 LLRGANTRTNFWPCSQSSTSP 103
+RG + NF P + + T+P
Sbjct: 711 RIRGDKAKLNF-PDTPAVTAP 770
>TC81407 homologue to GP|18650662|gb|AAL75809.1 ethylene response factor 1
{Lycopersicon esculentum}, partial (59%)
Length = 770
Score = 82.4 bits (202), Expect = 3e-16
Identities = 45/82 (54%), Positives = 59/82 (71%), Gaps = 4/82 (4%)
Frame = +1
Query: 33 RVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
R ++R+ GVRQR G WV+EI+ + K R+WLGTF+TAE+AARAYDEAA L+ G RTN
Sbjct: 58 RPQQRYRGVRQRHWGSWVSEIRHPLLKTRIWLGTFETAEDAARAYDEAARLMCGPKARTN 237
Query: 93 FWPCS----QSSTSPALSSKIT 110
F P + QSS+S LS+ +T
Sbjct: 238 F-PYNPNGPQSSSSKLLSATLT 300
>TC78754 homologue to GP|4099921|gb|AAD09248.1| EREBP-3 homolog
{Stylosanthes hamata}, partial (46%)
Length = 1038
Score = 80.9 bits (198), Expect = 8e-16
Identities = 44/80 (55%), Positives = 53/80 (66%), Gaps = 1/80 (1%)
Frame = +3
Query: 15 SMVWDDMMKEAASLGGARRVRK-RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEA 73
S+V M + GG V++ F GVR+RP GR+ AEI+D +K RVWLGTFDTAEEA
Sbjct: 81 SIVIISMAPKVVKNGGVNGVKEAHFRGVRKRPWGRYAAEIRDPGKKSRVWLGTFDTAEEA 260
Query: 74 ARAYDEAACLLRGANTRTNF 93
ARAYD AA RG +TNF
Sbjct: 261 ARAYDSAARQFRGPKAKTNF 320
>TC76795 similar to GP|18496063|emb|CAD21849. ethylene responsive element
binding protein {Fagus sylvatica}, partial (32%)
Length = 966
Score = 80.9 bits (198), Expect = 8e-16
Identities = 45/114 (39%), Positives = 70/114 (60%), Gaps = 1/114 (0%)
Frame = +3
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R+ + ++ G+RQRP G+W AEI+D + +RVWLGTF+TAEEAARAYD A +RG +
Sbjct: 348 RKRKNQYRGIRQRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAARAYDAEARRIRGKKAKV 527
Query: 92 NFWPCSQSSTSPALSSKITNLLLQRLKERNMNN-SCSSFSSSKYTSLLGSPEER 144
NF + +++S L + LL ++N+N+ C + + Y S + E+R
Sbjct: 528 NFPEEAPNASSKRLKTNSETQLL----DKNLNSFKCENIEN--YYSPVNQVEQR 671
>AW685524 homologue to GP|15982737|gb| AT4g23750/F9D16_220 {Arabidopsis
thaliana}, partial (21%)
Length = 666
Score = 80.5 bits (197), Expect = 1e-15
Identities = 36/58 (62%), Positives = 45/58 (77%)
Frame = +2
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
K+F GVRQRP G+W AEI+D +++R+WLGTF+TAEEAA YD AA LRG + TNF
Sbjct: 455 KKFRGVRQRPWGKWAAEIRDPARRVRLWLGTFETAEEAAMVYDNAAIKLRGPDALTNF 628
>TC90000 homologue to GP|9294548|dbj|BAB02811.1 ethylene responsive element
binding factor-like protein {Arabidopsis thaliana},
partial (25%)
Length = 458
Score = 80.1 bits (196), Expect = 1e-15
Identities = 38/57 (66%), Positives = 46/57 (80%)
Frame = +2
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
R+ GVR+RP GR+ AEI+D ++K RVWLGTFDTAE+AARAYD AA LRG +TNF
Sbjct: 263 RYRGVRKRPWGRFAAEIRDPLKKARVWLGTFDTAEQAARAYDTAARNLRGPKAKTNF 433
>TC76339 homologue to GP|4099914|gb|AAD00708.1| ethylene-responsive element
binding protein homolog {Stylosanthes hamata}, partial
(37%)
Length = 1110
Score = 79.7 bits (195), Expect = 2e-15
Identities = 43/74 (58%), Positives = 50/74 (67%), Gaps = 2/74 (2%)
Frame = +2
Query: 38 FVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF--WP 95
F GVR+RP GR+ AEI+D +K RVWLGTFDTAEEAARAYD AA RG +TNF P
Sbjct: 143 FRGVRKRPWGRYAAEIRDPGKKSRVWLGTFDTAEEAARAYDNAARQFRGPKAKTNFPPPP 322
Query: 96 CSQSSTSPALSSKI 109
SP+ SS +
Sbjct: 323 SDVKEDSPSQSSTV 364
>TC90274 similar to GP|10177219|dbj|BAB10294. contains similarity to AP2
domain transcription factor~gene_id:MPF21.9 {Arabidopsis
thaliana}, partial (32%)
Length = 750
Score = 79.3 bits (194), Expect = 2e-15
Identities = 49/120 (40%), Positives = 69/120 (56%)
Frame = +3
Query: 12 EEGSMVWDDMMKEAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAE 71
E G+ V++ E G ++++ GVRQRP G+W AEI+D + RVWLGTF+TAE
Sbjct: 93 EMGNAVYEYKRTENVVREG--EAKRKYRGVRQRPWGKWAAEIRDPFKAARVWLGTFETAE 266
Query: 72 EAARAYDEAACLLRGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSS 131
AARAYDEAA RG + NF P + + +P ++ T + N++NS SS S
Sbjct: 267 AAARAYDEAALRFRGNKAKLNF-PENVTLRNPPPATTTT--------QWNVSNSPSSIVS 419
>TC93662 similar to GP|10177219|dbj|BAB10294. contains similarity to AP2
domain transcription factor~gene_id:MPF21.9 {Arabidopsis
thaliana}, partial (31%)
Length = 555
Score = 79.3 bits (194), Expect = 2e-15
Identities = 37/59 (62%), Positives = 44/59 (73%)
Frame = +3
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
R+R+ GVRQRP G+W AEI+D + RVWLGTF+TAE AARAYDEAA RG + NF
Sbjct: 249 RRRYRGVRQRPWGKWAAEIRDPHKAARVWLGTFETAEAAARAYDEAALRFRGNRAKLNF 425
>TC86273 similar to SP|O80340|ERF4_ARATH Ethylene responsive element binding
factor 4 (AtERF4). [Mouse-ear cress] {Arabidopsis
thaliana}, partial (43%)
Length = 1171
Score = 79.3 bits (194), Expect = 2e-15
Identities = 38/57 (66%), Positives = 46/57 (80%)
Frame = +3
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
R+ GVR+RP GR+ AEI+D +K RVWLGTFDTAEEAARAYD+AA RG+ +TNF
Sbjct: 177 RYRGVRKRPWGRYAAEIRDPGKKTRVWLGTFDTAEEAARAYDKAAREFRGSKAKTNF 347
>BF650547 similar to GP|9758388|dbj| AP2 domain transcription factor-like
{Arabidopsis thaliana}, partial (24%)
Length = 418
Score = 78.6 bits (192), Expect = 4e-15
Identities = 47/93 (50%), Positives = 55/93 (58%)
Frame = -1
Query: 40 GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCSQS 99
GVRQRP G+W AEI+D + RVWLGTF+TAE AA AYDEAA +G + NF
Sbjct: 379 GVRQRPWGKWAAEIRDPKKGARVWLGTFETAEAAATAYDEAALRFKGNKAKLNF------ 218
Query: 100 STSPALSSKITNLLLQRLKERNMNNSCSSFSSS 132
P L I+NL RN NN+ SS SSS
Sbjct: 217 ---PRL---ISNLPSDPKPPRNKNNASSSSSSS 137
>TC76792 similar to GP|3264767|gb|AAC24587.1| AP2 domain containing protein
{Prunus armeniaca}, partial (49%)
Length = 1729
Score = 78.6 bits (192), Expect = 4e-15
Identities = 46/125 (36%), Positives = 67/125 (52%)
Frame = +3
Query: 24 EAASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACL 83
+ A R+ + ++ G+RQRP G+W AEI+D + +RVWLGTF+TAEEAARAYD A
Sbjct: 486 QQAEKDSKRKRKNQYRGIRQRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAARAYDAEARR 665
Query: 84 LRGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEE 143
+RG + NF + ++S L LL N F+ S+ SP +
Sbjct: 666 IRGNKAKVNFPEEAPVASSKRLKKNPEMQLLNDNLNSFKPNGNQMFNFSENMENYYSPMD 845
Query: 144 RREQQ 148
+ EQ+
Sbjct: 846 QVEQK 860
>TC90093 similar to PIR|T47955|T47955 hypothetical protein F15G16.20 -
Arabidopsis thaliana, partial (24%)
Length = 1005
Score = 77.8 bits (190), Expect = 7e-15
Identities = 45/102 (44%), Positives = 58/102 (56%), Gaps = 13/102 (12%)
Frame = +3
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF-- 93
K++ GVRQRP G+W AEI+D + +RVWLGTF TAEEAA YD AA LRG + TNF
Sbjct: 690 KKYRGVRQRPWGKWAAEIRDPARGVRVWLGTFQTAEEAAIVYDNAAIKLRGPDALTNFIT 869
Query: 94 ------WPCSQSSTSPALSSKITNLLL-----QRLKERNMNN 124
C SS P +++L L + + +N NN
Sbjct: 870 PPVSAAATCQLSSPPPENEIPLSSLPLPANSGEESQSQNNNN 995
>TC82756 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_180
{Arabidopsis thaliana}, partial (35%)
Length = 630
Score = 76.6 bits (187), Expect = 2e-14
Identities = 36/65 (55%), Positives = 43/65 (65%)
Frame = +3
Query: 32 RRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRT 91
R ++ GVRQRP G+W AEI+D + +RVWLGTF TAEEAARAYD AA RG +
Sbjct: 246 RAKNNKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDNAAIEFRGPRAKL 425
Query: 92 NFWPC 96
N C
Sbjct: 426 NLSTC 440
>TC87696 similar to GP|9759176|dbj|BAB09791.1 contains similarity to
pathogenesis-related genes transcriptional
activator~gene_id:K19E1.9, partial (24%)
Length = 1725
Score = 76.3 bits (186), Expect = 2e-14
Identities = 35/61 (57%), Positives = 43/61 (70%)
Frame = +3
Query: 33 RVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
R F GVRQRP G+W AEI+D I++ R+WLGTF TAEEAA YD A +L G+N TN
Sbjct: 870 RRHSNFRGVRQRPWGKWAAEIRDPIRRKRLWLGTFSTAEEAAAEYDRVAVMLHGSNAVTN 1049
Query: 93 F 93
+
Sbjct: 1050Y 1052
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,938,993
Number of Sequences: 36976
Number of extensions: 156082
Number of successful extensions: 1012
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 961
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 988
length of query: 411
length of database: 9,014,727
effective HSP length: 98
effective length of query: 313
effective length of database: 5,391,079
effective search space: 1687407727
effective search space used: 1687407727
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC119412.1


