

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119408.6 - phase: 0
(535 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
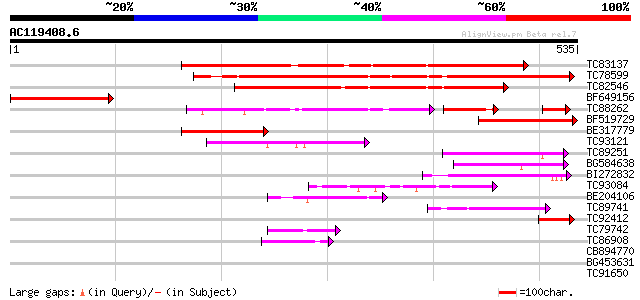
Score E
Sequences producing significant alignments: (bits) Value
TC83137 similar to GP|9758523|dbj|BAB08970.1 contains similarity... 341 4e-94
TC78599 similar to GP|9758008|dbj|BAB08605.1 contains similarity... 307 6e-84
TC82546 similar to PIR|T04446|T04446 hypothetical protein F4D11.... 304 6e-83
BF649156 198 4e-51
TC88262 homologue to GP|15293111|gb|AAK93666.1 unknown protein {... 128 3e-37
BF519729 similar to PIR|T04446|T044 hypothetical protein F4D11.1... 137 1e-32
BE317779 similar to GP|19347795|gb| unknown protein {Arabidopsis... 115 3e-26
TC93121 weakly similar to GP|8953375|emb|CAB96648.1 putative pro... 108 5e-24
TC89251 similar to PIR|H84680|H84680 hypothetical protein At2g28... 82 5e-16
BG584638 similar to PIR|T47480|T47 hypothetical protein F18N11.1... 64 2e-10
BI272832 similar to GP|18650643|gb At1g21480/F24J8_23 {Arabidops... 62 7e-10
TC93084 similar to PIR|H86466|H86466 protein F23M19.7 [imported]... 52 7e-07
BE204106 homologue to GP|15809826|gb AT5g61840/mac9_140 {Arabido... 48 8e-06
TC89741 similar to PIR|T06757|T06757 hypothetical protein F15B8.... 47 1e-05
TC92412 weakly similar to GP|15146187|gb|AAK83577.1 AT5g33290/F1... 45 9e-05
TC79742 similar to GP|10177241|dbj|BAB10615. gb|AAC98455.1~gene_... 45 9e-05
TC86908 similar to PIR|G96708|G96708 hypothetical protein T26J14... 43 3e-04
CB894770 homologue to GP|4512698|gb| unknown protein {Arabidopsi... 34 0.12
BG453631 homologue to GP|21106839|gb| disulfide oxidoreductase {... 31 1.0
TC91650 similar to GP|8809635|dbj|BAA97186.1 gb|AAD21751.1~gene_... 30 1.8
>TC83137 similar to GP|9758523|dbj|BAB08970.1 contains similarity to
limonene cyclase~gene_id:K15O15.2 {Arabidopsis
thaliana}, partial (50%)
Length = 994
Score = 341 bits (875), Expect = 4e-94
Identities = 173/327 (52%), Positives = 225/327 (67%)
Frame = +2
Query: 163 LVYARKEIDHVTSVNEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSL 222
L+ A+ EI++ + + L+AP++R VS F RSYELME LKVYIYR+G +PIFH P +
Sbjct: 38 LLAAKIEIENANVLLKSSGLHAPVYREVSKFSRSYELMERKLKVYIYREGEKPIFHQPKM 217
Query: 223 KGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLR 282
+GIYASEGWFMKLM+ NK+F+ KDP++AHLFYLP+S++ + L D K + +L
Sbjct: 218 RGIYASEGWFMKLMEGNKRFIVKDPKKAHLFYLPFSSQMLRANL-----SDNKKMEQYLD 382
Query: 283 DYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIE 342
YVN IA KY FWNRT G+DHFLVACH+W + +N +++LCNA++++ I
Sbjct: 383 KYVNIIAGKYRFWNRTGGADHFLVACHDWASRIT---RQPMKNCIRSLCNANVAKGFQI- 550
Query: 343 GRDVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMK 402
G+D +LP T I + PLR + G S R ILAFFAGSMHG +RP LLK+W K DMK
Sbjct: 551 GKDTTLPATYIHSVMNPLRKIAGKHPSERTILAFFAGSMHGYLRPILLKHW-ENKEPDMK 727
Query: 403 IYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLS 462
I+ + K Y+ +M SSKYC+C G+EV SPRIVEAI+ ECVPVII+DN+V P
Sbjct: 728 IFGAMARDAEGKRIYMDYMNSSKYCICARGYEVYSPRIVEAIFSECVPVIISDNYVPPFF 907
Query: 463 EVLDWSAFSVVVAEKDIPRLKDILLSI 489
EVL W AFSV V E+D+P L ILLSI
Sbjct: 908 EVLKWEAFSVFVRERDVPNLXSILLSI 988
>TC78599 similar to GP|9758008|dbj|BAB08605.1 contains similarity to
limonene cyclase~gene_id:MED24.9 {Arabidopsis thaliana},
partial (85%)
Length = 1857
Score = 307 bits (787), Expect = 6e-84
Identities = 162/361 (44%), Positives = 234/361 (63%), Gaps = 1/361 (0%)
Frame = +3
Query: 174 TSVNEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSLKGIYAS-EGWF 232
T+V++ + LFRN ET+ + + R GS F L +Y +G F
Sbjct: 558 TNVSQS*FISQELFRNG----------ETIQSICV*RKGSLR-FSIMDLAKVYTPWKGNF 704
Query: 233 MKLMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKY 292
+ ++ N +F T+DPE+AH+F+LP+S +M +YV SHD P+ + DY+N ++ KY
Sbjct: 705 IHAIEMNDRFRTRDPEKAHVFFLPFSVAKMVQFVYVRDSHDFSPIRKTITDYINVVSEKY 884
Query: 293 PFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETT 352
PFWNR+ G+DHF+++CH+WGP T L +N+++ALCNA+ SE F +DVS+PET
Sbjct: 885 PFWNRSLGADHFMLSCHDWGPETSKSVPNLYKNSIRALCNANTSEG-FKPAKDVSIPETN 1061
Query: 353 IRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMKIYKPLPLRVS 412
++ +GG S R +LAFFAG +HG VRP LL++W K ED++++K LP
Sbjct: 1062LQTGTIH-GIVGGPSPSKRSVLAFFAGGVHGPVRPVLLEHWE-HKDEDLQVHKYLP---- 1223
Query: 413 KKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSV 472
K ++Y ++ SK+CLCP G+EV SPR+VEAIY CVPV+I+D++V P S+VL+W +FSV
Sbjct: 1224KGVSYYDMLRKSKFCLCPSGYEVASPRVVEAIYTGCVPVLISDHYVPPFSDVLNWKSFSV 1403
Query: 473 VVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKL 532
V+ DIP LK IL SI R+Y+ MQ V V++HF + P R+D+FHMILHSIWL +L
Sbjct: 1404EVSVNDIPNLKKILTSISPRQYIRMQRRVGQVRRHFEVHSPPTRFDVFHMILHSIWLRRL 1583
Query: 533 N 533
N
Sbjct: 1584N 1586
Score = 31.6 bits (70), Expect = 0.80
Identities = 29/99 (29%), Positives = 44/99 (44%), Gaps = 13/99 (13%)
Frame = +1
Query: 126 KDRNLANTST-----TVTTSFPRGRVPSGKQTDIR---LITPTEA-LVYARKEIDHVTSV 176
KD+ N S T +TS ++ K R ++ TEA L+ AR I +
Sbjct: 343 KDKESLNASQILPNITTSTSMNESQILLQKPNLPRKFSILDRTEAGLLQARAAIRKARNE 522
Query: 177 NEDPDL----YAPLFRNVSVFKRSYELMETVLKVYIYRD 211
N D+ P++ N + F RSY ME KV++Y +
Sbjct: 523 NRTQDIDYVPTGPMYHNPNSFHRSYLEMEKQFKVFVYEE 639
>TC82546 similar to PIR|T04446|T04446 hypothetical protein F4D11.10 -
Arabidopsis thaliana, partial (35%)
Length = 760
Score = 304 bits (778), Expect = 6e-83
Identities = 152/258 (58%), Positives = 188/258 (71%)
Frame = +1
Query: 213 SRPIFHNPSLKGIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSH 272
+RPI H+P L GIYASEGWFMKLM+ NK+FVTK+P++AHLFYLP+S+R +E LYV SH
Sbjct: 4 ARPISHSPFLTGIYASEGWFMKLMEANKRFVTKNPKKAHLFYLPFSSRMLEEALYVKNSH 183
Query: 273 DLKPLSIFLRDYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCN 332
K L +L DYV+ IAA++ FWNRT G+DHFLV CH+W P +E + N +++LCN
Sbjct: 184 SHKNLIQYLHDYVDMIAARHSFWNRTGGADHFLVGCHDWAP---SETKLRLANCIRSLCN 354
Query: 333 ADLSERIFIEGRDVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKY 392
AD+ E F+ G+D SLPET +R + P R LGGN S + LAFFAGSMHG VRP LLK+
Sbjct: 355 ADVKEG-FVFGKDASLPETYVRNAQIPTRDLGGNSFSKKTTLAFFAGSMHGYVRPILLKH 531
Query: 393 WGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVI 452
W K DMKI+ LP YI +MKSSKYC+C G+EVNSPR+VEAI+YECVPVI
Sbjct: 532 W-ENKDPDMKIFGKLP-NSKGNSNYIHYMKSSKYCICAKGYEVNSPRVVEAIFYECVPVI 705
Query: 453 IADNFVLPLSEVLDWSAF 470
I+DNFV EVLDW +F
Sbjct: 706 ISDNFVPAFFEVLDWESF 759
>BF649156
Length = 509
Score = 198 bits (504), Expect = 4e-51
Identities = 97/98 (98%), Positives = 97/98 (98%)
Frame = +3
Query: 1 MKKVFHLTVIILMRELSWRCFRHKIDWRRLFLLVSIFTASGFVFQMIVHTSVISPNKHFH 60
MKKVFHLTVIILMRELSWRCFRHKIDWRRLFLLVSIFTASGFVFQMIVHTSVISPNKHFH
Sbjct: 198 MKKVFHLTVIILMRELSWRCFRHKIDWRRLFLLVSIFTASGFVFQMIVHTSVISPNKHFH 377
Query: 61 FPPGAADSYKSSNSTIQSNEFIKETILQNVHLTLPNSA 98
FPPGAADSYKSSNSTIQSNEFIKETILQNVHLTL NSA
Sbjct: 378 FPPGAADSYKSSNSTIQSNEFIKETILQNVHLTLANSA 491
>TC88262 homologue to GP|15293111|gb|AAK93666.1 unknown protein {Arabidopsis
thaliana}, partial (66%)
Length = 1367
Score = 128 bits (321), Expect(2) = 3e-37
Identities = 81/239 (33%), Positives = 128/239 (52%), Gaps = 5/239 (2%)
Frame = +2
Query: 168 KEIDHVTSVNEDP--DLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNP--SLK 223
K +D + ED D + ++ + VFK ++ ME KVYIY DG F+ L
Sbjct: 293 KVVDAGKNEEEDDGGDEFGDVYHSPRVFKLNFAEMEKKFKVYIYPDGDSKTFYQTPRKLT 472
Query: 224 GIYASEGWFMKLMQENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSIFLRD 283
G YASEG+F + ++E++ F T DP+ AHLF++P S +M S+D ++I +++
Sbjct: 473 GKYASEGYFFQNIRESR-FRTLDPDEAHLFFIPISCHKMRGK---GTSYD--NMTIIVQN 634
Query: 284 YVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERIFIEG 343
YV + +KYP+WNRT G+DHF V CH+ G L +N+++A+C+ FI
Sbjct: 635 YVESLISKYPYWNRTLGADHFFVTCHDVGVRATEGLPLLVKNSIRAVCSPSYDVG-FIPH 811
Query: 344 RDVSLPETTIRAPRRPLRY-LGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDM 401
+DV+LP+ +P GGN R L F+AG + ++R L + W + D+
Sbjct: 812 KDVALPQVL-----QPFALPAGGNDVENRTSLGFWAGHRNSKIRVILARVWENDTELDI 973
Score = 47.8 bits (112), Expect = 1e-05
Identities = 15/27 (55%), Positives = 22/27 (80%)
Frame = +1
Query: 503 MVQKHFLWNPKPIRYDLFHMILHSIWL 529
++QKHF WN P+RYD FHM+++ +WL
Sbjct: 1114 LIQKHFQWNSPPVRYDAFHMVMYDLWL 1194
Score = 45.8 bits (107), Expect(2) = 3e-37
Identities = 20/52 (38%), Positives = 33/52 (63%)
Frame = +3
Query: 410 RVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPL 461
R + + Y + S+K+C+CP G +VNS RI ++I+Y C+P D+ LP+
Sbjct: 993 RATGHLVYQKRFYSTKFCICPGGSQVNSARIADSIHYGCIP----DSETLPV 1136
>BF519729 similar to PIR|T04446|T044 hypothetical protein F4D11.10 -
Arabidopsis thaliana, partial (15%)
Length = 536
Score = 137 bits (345), Expect = 1e-32
Identities = 61/93 (65%), Positives = 77/93 (82%)
Frame = +3
Query: 443 AIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVK 502
AI+YECVPVII+DNFV P EVL+W +FS+++AEKDIP LK ILLS+P KY+ +Q V+
Sbjct: 3 AIFYECVPVIISDNFVPPFFEVLNWDSFSLILAEKDIPNLKQILLSVPEEKYLKLQLGVR 182
Query: 503 MVQKHFLWNPKPIRYDLFHMILHSIWLNKLNQI 535
VQKHFLW+ KP++YDLFHM LHSIW N++ QI
Sbjct: 183 RVQKHFLWHTKPLKYDLFHMTLHSIWYNRVFQI 281
>BE317779 similar to GP|19347795|gb| unknown protein {Arabidopsis thaliana},
partial (14%)
Length = 295
Score = 115 bits (289), Expect = 3e-26
Identities = 52/82 (63%), Positives = 66/82 (80%)
Frame = +1
Query: 163 LVYARKEIDHVTSVNEDPDLYAPLFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSL 222
++ + EI++ ++ D LY+PL+RNVS+F+RSYELME +LKVYIY DG RPIFH P L
Sbjct: 43 IMKVKSEIENAPTIMNDSRLYSPLYRNVSMFRRSYELMEKMLKVYIYADGDRPIFHEPLL 222
Query: 223 KGIYASEGWFMKLMQENKQFVT 244
+GIYASEGWFMKLM+ NKQF T
Sbjct: 223 EGIYASEGWFMKLMEGNKQFTT 288
>TC93121 weakly similar to GP|8953375|emb|CAB96648.1 putative protein
{Arabidopsis thaliana}, partial (27%)
Length = 918
Score = 108 bits (270), Expect = 5e-24
Identities = 53/160 (33%), Positives = 92/160 (57%), Gaps = 6/160 (3%)
Frame = +1
Query: 186 LFRNVSVFKRSYELMETVLKVYIYRDGSRPIFHNPSLKGIYASEGWFMKLMQENKQ--FV 243
++ N F +S++ M KV++Y++G +P+ H+ + Y+ EG F+ M + + F
Sbjct: 325 IYLNPHAFHQSHKEMVKRFKVWVYKEGEQPLVHDGPVNNKYSIEGQFIDEMDTSNKSPFK 504
Query: 244 TKDPERAHLFYLPYSARQMEVTLYVP--GSHDLKP--LSIFLRDYVNKIAAKYPFWNRTH 299
PE AH+F+LP+S ++ +Y P D P L + + DY+ +A KYP+WN +
Sbjct: 505 ATHPELAHVFFLPFSVSKVIRYVYKPRKSRSDYNPHRLQLLVEDYIKIVANKYPYWNISQ 684
Query: 300 GSDHFLVACHEWGPYTVTEHEELARNTLKALCNADLSERI 339
G+D FL++CH+WGP + +L ++ ++ALCN RI
Sbjct: 685 GADQFLLSCHDWGPRVSYANPKLFKHFIRALCNGQHFRRI 804
>TC89251 similar to PIR|H84680|H84680 hypothetical protein At2g28110
[imported] - Arabidopsis thaliana, partial (30%)
Length = 923
Score = 82.0 bits (201), Expect = 5e-16
Identities = 46/121 (38%), Positives = 67/121 (55%), Gaps = 2/121 (1%)
Frame = +1
Query: 409 LRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWS 468
LR + Y + S +CLCP+G+ SPR+VE++ CVPVIIAD+ LP S ++W
Sbjct: 79 LRRHRFAGYQSEIARSVFCLCPLGWAPWSPRLVESVALGCVPVIIADSIRLPFSSAVNWP 258
Query: 469 AFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNV--KMVQKHFLWNPKPIRYDLFHMILHS 526
SV VAEKD+ RL +IL + +Q N+ +K L+N + D +LHS
Sbjct: 259 EISVTVAEKDVWRLGEILEKVAATNLSIIQRNLWDPRTRKALLFNSRVHEGDATWQVLHS 438
Query: 527 I 527
+
Sbjct: 439 L 441
>BG584638 similar to PIR|T47480|T47 hypothetical protein F18N11.160 -
Arabidopsis thaliana, partial (23%)
Length = 758
Score = 63.5 bits (153), Expect = 2e-10
Identities = 40/112 (35%), Positives = 57/112 (50%), Gaps = 3/112 (2%)
Frame = -2
Query: 419 QHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIADNFVLPLSEVLDWSAFSVVVAEKD 478
Q M SK+CL G +S R+ +AI CVPVII+D LP +VLD+S FSV V D
Sbjct: 682 QGMSMSKFCLNIAGDTPSSNRLFDAIVSHCVPVIISDEIELPFEDVLDYSEFSVFVRAAD 503
Query: 479 IPR---LKDILLSIPMRKYVAMQNNVKMVQKHFLWNPKPIRYDLFHMILHSI 527
+ L ++L SI ++ M +K + HF + D +MI +
Sbjct: 502 AVKKGYLLNLLHSIKREEWTKMWERLKEITNHFEYQYPSQPGDAVNMIWQEV 347
>BI272832 similar to GP|18650643|gb At1g21480/F24J8_23 {Arabidopsis
thaliana}, partial (42%)
Length = 607
Score = 61.6 bits (148), Expect = 7e-10
Identities = 46/156 (29%), Positives = 73/156 (46%), Gaps = 15/156 (9%)
Frame = +3
Query: 390 LKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECV 449
LK+ GGEK + Y +H+++SK+CL P G + R E+ + ECV
Sbjct: 105 LKFSGGEKLG--------------RKDYFEHLRNSKFCLAPRGESSWTLRFYESFFVECV 242
Query: 450 PVIIADNFVLPLSEVLDWSAFSVVVAEKDI-PRLKDILLSIPMRKYVAMQNNVKMVQKHF 508
PVI++D LP V+D+S S+ I P L L SIP + A+ + V+ +
Sbjct: 243 PVILSDQIELPFQNVIDYSQISIKWPSSRIGPELLQYLESIPDKDIEAIIARGRQVRCMW 422
Query: 509 LW--NPKP------IRYDL------FHMILHSIWLN 530
++ + KP I ++L FH + WL+
Sbjct: 423 VYASDSKPCSAMRGIMWELQRKVRQFHQSAETFWLH 530
>TC93084 similar to PIR|H86466|H86466 protein F23M19.7 [imported] -
Arabidopsis thaliana, partial (46%)
Length = 676
Score = 51.6 bits (122), Expect = 7e-07
Identities = 59/198 (29%), Positives = 85/198 (42%), Gaps = 20/198 (10%)
Frame = +3
Query: 283 DYVNKIAAKYPFWNRTHGSDHFLVACHEWGPYTVTEHEELARNTL------------KAL 330
D+V K A WNR+ G DH V + V + E+A L L
Sbjct: 123 DFVKKTEA----WNRSGGRDHVFVVTDPVAMWHVKD--EIAPAILLVVDFGGWYRLDSKL 284
Query: 331 CNADLSERIFIEG----RDVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMH---- 382
N SE I +DV +P T + P +L N+ R L +F G+ H
Sbjct: 285 SNCTSSEMIQHTQVSVIKDVIVPYTHLL----PRLHLSENKE--RHNLLYFKGAKHRHRG 446
Query: 383 GRVRPTLLKYWGGEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVE 442
G VR L E M+ + P ++ + IQ M++S++CL P G S R+ +
Sbjct: 447 GLVREKLWDLLNNEPSVIME--EGFPNATGREQS-IQGMRTSEFCLHPAGDTPTSCRLFD 617
Query: 443 AIYYECVPVIIADNFVLP 460
AI C+PVI++DN LP
Sbjct: 618 AIQSLCIPVIVSDNIELP 671
>BE204106 homologue to GP|15809826|gb AT5g61840/mac9_140 {Arabidopsis
thaliana}, partial (44%)
Length = 555
Score = 48.1 bits (113), Expect = 8e-06
Identities = 36/122 (29%), Positives = 55/122 (44%), Gaps = 9/122 (7%)
Frame = +2
Query: 244 TKDPERAHLFYLPYSARQMEVTLYVPGSHDLKPLSI--------FLRDYVNKIAAKYPFW 295
T +PE A FY P V + DL P + +R + I++ +P+W
Sbjct: 119 TLNPEEADWFYTP-----------VYTTCDLTPNGLPLPFKSPRMMRSAIQLISSNWPYW 265
Query: 296 NRTHGSDHFLVACHEWGP-YTVTEHEELARNTLKALCNADLSERIFIEGRDVSLPETTIR 354
NRT G+DHF V H++G + E + + R L L A L + F + V L + +I
Sbjct: 266 NRTEGADHFFVVPHDFGACFHYQEEKAIERGILPLLRRATLVQ-TFGQRNHVCLKDGSIT 442
Query: 355 AP 356
P
Sbjct: 443 IP 448
>TC89741 similar to PIR|T06757|T06757 hypothetical protein F15B8.180 -
Arabidopsis thaliana, partial (32%)
Length = 1233
Score = 47.4 bits (111), Expect = 1e-05
Identities = 32/116 (27%), Positives = 57/116 (48%)
Frame = +1
Query: 395 GEKYEDMKIYKPLPLRVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPVIIA 454
G++Y + I PL + Y + SS +C G + S R+ +++ C+PV+I
Sbjct: 415 GKQYAEDVIMTPL-----RSDNYHADIASSVFCGVLPG-DGWSGRMEDSVLQGCIPVVIQ 576
Query: 455 DNFVLPLSEVLDWSAFSVVVAEKDIPRLKDILLSIPMRKYVAMQNNVKMVQKHFLW 510
D LP VL++ +F+V + E +IP + IL + NV+ + + FL+
Sbjct: 577 DGIFLPYENVLNYDSFAVRIPEDEIPNMIKILRGFNETEINLKLANVQKIWQRFLY 744
>TC92412 weakly similar to GP|15146187|gb|AAK83577.1 AT5g33290/F19N2_10
{Arabidopsis thaliana}, partial (17%)
Length = 540
Score = 44.7 bits (104), Expect = 9e-05
Identities = 19/34 (55%), Positives = 24/34 (69%)
Frame = +1
Query: 500 NVKMVQKHFLWNPKPIRYDLFHMILHSIWLNKLN 533
NV+ V+KHF N +DL HMILHS+WL +LN
Sbjct: 127 NVRRVRKHFEMNRPAKPFDLIHMILHSVWLRRLN 228
>TC79742 similar to GP|10177241|dbj|BAB10615.
gb|AAC98455.1~gene_id:MRN17.17~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (32%)
Length = 1063
Score = 44.7 bits (104), Expect = 9e-05
Identities = 25/70 (35%), Positives = 35/70 (49%), Gaps = 1/70 (1%)
Frame = +2
Query: 244 TKDPERAHLFYLP-YSARQMEVTLYVPGSHDLKPLSIFLRDYVNKIAAKYPFWNRTHGSD 302
T DP A F++P Y + P + L + V I+ +YPFWNR+ GSD
Sbjct: 626 TFDPYEADFFFVPVYVSCNFSTVNGFPAIGHARSL---ISSAVKLISTEYPFWNRSTGSD 796
Query: 303 HFLVACHEWG 312
H VA H++G
Sbjct: 797 HVFVASHDFG 826
>TC86908 similar to PIR|G96708|G96708 hypothetical protein T26J14.4
[imported] - Arabidopsis thaliana, partial (65%)
Length = 1587
Score = 43.1 bits (100), Expect = 3e-04
Identities = 24/69 (34%), Positives = 39/69 (55%), Gaps = 1/69 (1%)
Frame = +2
Query: 238 ENKQFVTKDPERAHLFYLPYSARQMEVTLYVPGSHDLKP-LSIFLRDYVNKIAAKYPFWN 296
EN T DP +A LFY+P+ +++ +H L+ L++ L D+V + P+WN
Sbjct: 605 ENHVCRTWDPNQAILFYIPFYGGLHASSMFREANHTLRDSLAVDLIDHVLSL----PWWN 772
Query: 297 RTHGSDHFL 305
R +G DHF+
Sbjct: 773 RHNGKDHFI 799
>CB894770 homologue to GP|4512698|gb| unknown protein {Arabidopsis thaliana},
partial (40%)
Length = 720
Score = 34.3 bits (77), Expect = 0.12
Identities = 42/175 (24%), Positives = 67/175 (38%), Gaps = 14/175 (8%)
Frame = +1
Query: 291 KYPFWNRTHGSDHFLVACH-EWGPYTVTEHEELARNTLKALCNADLSERIFIE-----GR 344
K P W+ +G DHFLVA W ++E E N L L A + +E
Sbjct: 79 KRPEWSIMNGKDHFLVAGRITWDFRRLSEEETDWGNKLLFLPAAKNMSMLVVESSPWNAN 258
Query: 345 DVSLPETTIRAPRRPLRYLGGNRASLRPILAFFAGSMHGRVRPTLLKYWGGEKYEDMK-- 402
D +P T P + + +R + + S G RP K G+ E +
Sbjct: 259 DFGIPYPTYFHPAKDADVFNW-QDRMRHLERKWLFSFAGAPRPDNAKSIRGQLIEQCRSS 435
Query: 403 -IYKPLPL-----RVSKKMTYIQHMKSSKYCLCPMGFEVNSPRIVEAIYYECVPV 451
+ K L + + +Q +SS++CL P G +++ C+PV
Sbjct: 436 PVGKLLECDFGESKCHSPSSIMQMFQSSQFCLQPQGDSYTRRSAFDSMLAGCIPV 600
>BG453631 homologue to GP|21106839|gb| disulfide oxidoreductase {Xanthomonas
axonopodis pv. citri str. 306}, partial (5%)
Length = 661
Score = 31.2 bits (69), Expect = 1.0
Identities = 12/30 (40%), Positives = 22/30 (73%), Gaps = 2/30 (6%)
Frame = +2
Query: 7 LTVIILMRELSWR--CFRHKIDWRRLFLLV 34
L +I LMR++SW+ C +KI W+ ++L++
Sbjct: 287 LLIICLMRKMSWKIICLMNKISWKIVWLII 376
>TC91650 similar to GP|8809635|dbj|BAA97186.1
gb|AAD21751.1~gene_id:MMI9.5~similar to unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 754
Score = 30.4 bits (67), Expect = 1.8
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +1
Query: 510 WNPKPIRYDLFHMILHSIWLN 530
WNP P+R +FH L S W N
Sbjct: 616 WNPNPLRGSMFHSTLDSQWEN 678
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,510,493
Number of Sequences: 36976
Number of extensions: 251993
Number of successful extensions: 1462
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1439
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1447
length of query: 535
length of database: 9,014,727
effective HSP length: 101
effective length of query: 434
effective length of database: 5,280,151
effective search space: 2291585534
effective search space used: 2291585534
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC119408.6


