

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119408.1 + phase: 0 /pseudo
(600 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
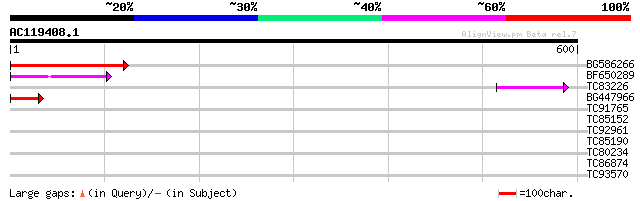
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 106 2e-23
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 64 1e-10
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 46 4e-05
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 42 5e-04
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 33 0.24
TC85152 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 32 0.91
TC92961 similar to GP|22135986|gb|AAM91575.1 unknown protein {Ar... 32 0.91
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 32 0.91
TC80234 similar to GP|12325097|gb|AAG52506.1 unknown protein; 31... 31 1.5
TC86874 homologue to SP|Q42808|TF2D_SOYBN Transcription initiati... 28 7.7
TC93570 similar to GP|21553436|gb|AAM62529.1 unknown {Arabidopsi... 28 7.7
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 106 bits (265), Expect = 2e-23
Identities = 52/125 (41%), Positives = 79/125 (62%), Gaps = 1/125 (0%)
Frame = -3
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
++ R++ +L +I + QS FVPGR I+DN LI + HY+R++ + + +K D+ KA
Sbjct: 718 LSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYLRQSGAKKHVSMAVKTDMTKA 539
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD + WNF+ + LT +GF I+ IM C+ V++S LING P P RG+RQGDP
Sbjct: 538 YDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLINGGPQGRVLPSRGLRQGDPLS 359
Query: 121 PLIFL 125
P +F+
Sbjct: 358 PYLFI 344
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 64.3 bits (155), Expect = 1e-10
Identities = 32/106 (30%), Positives = 59/106 (55%)
Frame = +3
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ +R++ +L ++ ++ QS FV GR+I DN ++ E +R + +K+D+ KA
Sbjct: 303 LTSRMQGVLNSVVSENQSAFVKGRVIFDNIILSHELVKSY--SRKGISPRCMVKIDLXKA 476
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEP 107
YDS +W FI+ + +GFP +N +M + +++ NG T P
Sbjct: 477 YDSXEWPFIKHLMLELGFPYKFVNWVMAXLTTASYTFNXNGDLTXP 614
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 46.2 bits (108), Expect = 4e-05
Identities = 34/76 (44%), Positives = 38/76 (49%)
Frame = +1
Query: 516 IATLRLIYGKNCGPYTLFPYTK*FCGEF*ITLYL*EVNWGKGASFATLFVQDVTQKLSLW 575
+ +R YGK G TLFP T+ GE * TL EV KG TL V D T KL L
Sbjct: 97 LVVMRPSYGKRFGHCTLFPDTRCCFGES*TTLSQLEVL*EKGEFSVTLSVLDATPKLKLS 276
Query: 576 STFS*HVPKSLEFGLA 591
T+S EFGLA
Sbjct: 277 LTYSCLALSQKEFGLA 324
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 42.4 bits (98), Expect = 5e-04
Identities = 20/34 (58%), Positives = 26/34 (75%)
Frame = +2
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVF 35
IANR+K LP++ + QS FV GRLITDN+LI +
Sbjct: 536 IANRVKQTLPDVIDVEQSAFVQGRLITDNALIAW 637
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 33.5 bits (75), Expect = 0.24
Identities = 14/41 (34%), Positives = 23/41 (55%), Gaps = 1/41 (2%)
Frame = +2
Query: 89 LCIRCVTFSILINGQPTEPFSPQRGIRQGDPF-PLIFLYYV 128
+C+ + +L+N +P P RG++QGD P IF+ V
Sbjct: 101 MCVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICV 223
>TC85152 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (18%)
Length = 775
Score = 31.6 bits (70), Expect = 0.91
Identities = 29/94 (30%), Positives = 47/94 (49%)
Frame = +2
Query: 133 LV*SQNVKMKV*FMVSPLPLMPQLYLIYFMLMIVSYFVGPNPRRLVLL*TFFRFIKRLLD 192
L+ N M++* ++ PLPL+P LI +L+I+ L+LL + RL
Sbjct: 194 LLRKNNRLMEL*LLLLPLPLLPLPPLIRKLLLIL----------LLLL------LMRLRS 325
Query: 193 RKLIWINQRWRSAQTSLMIYRKYFKTIYLLRSAL 226
R+ +W+ RWR L++ + F +LLR L
Sbjct: 326 RRKLWL-IRWR---MKLLLMKARFLNRFLLRKKL 415
>TC92961 similar to GP|22135986|gb|AAM91575.1 unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 842
Score = 31.6 bits (70), Expect = 0.91
Identities = 16/54 (29%), Positives = 30/54 (54%)
Frame = +1
Query: 145 FMVSPLPLMPQLYLIYFMLMIVSYFVGPNPRRLVLL*TFFRFIKRLLDRKLIWI 198
F +SP+ LMP L+L++ L+ + + +LVLL + ++ L R+ W+
Sbjct: 358 FGISPILLMPNLFLLFIELLFMGLMLRYLGNQLVLLQQMIQIMEILYRRQKQWL 519
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 31.6 bits (70), Expect = 0.91
Identities = 29/94 (30%), Positives = 47/94 (49%)
Frame = +1
Query: 133 LV*SQNVKMKV*FMVSPLPLMPQLYLIYFMLMIVSYFVGPNPRRLVLL*TFFRFIKRLLD 192
L+ N M++* ++ PLPL+P LI +L+I+ L+LL + RL
Sbjct: 190 LLRKNNRLMEL*LLLLPLPLLPLPPLIRKLLLIL----------LLLL------LMRLRS 321
Query: 193 RKLIWINQRWRSAQTSLMIYRKYFKTIYLLRSAL 226
R+ +W+ RWR L++ + F +LLR L
Sbjct: 322 RRKLWL-IRWR---MKLLLMKARFLNRFLLRKKL 411
>TC80234 similar to GP|12325097|gb|AAG52506.1 unknown protein; 31958-32770
{Arabidopsis thaliana}, partial (65%)
Length = 1374
Score = 30.8 bits (68), Expect = 1.5
Identities = 28/114 (24%), Positives = 51/114 (44%), Gaps = 10/114 (8%)
Frame = +2
Query: 120 FPLIFLYYVLRFFLV*SQNVKMKV*FMVSPLPLMPQLYLIYFMLMIVSYFVGPNPRRLV- 178
FPL+ ++ ++ FF++ V + + F+++ + +P IV YF P +RLV
Sbjct: 908 FPLVIIFLLIAFFVIPCSIVILIIYFLLTIIFAIPS--------FIVLYFAYPTIQRLVR 1063
Query: 179 ---------LL*TFFRFIKRLLDRKLIWINQRWRSAQTSLMIYRKYFKTIYLLR 223
LL + L ++K +WI +++ +R YF LLR
Sbjct: 1064 EITS*DEDLLLQSVLVV*SNLSEKKNLWIE------SSTMW*FRCYF*VFILLR 1207
>TC86874 homologue to SP|Q42808|TF2D_SOYBN Transcription initiation factor
TFIID (TATA-box factor) (TATA sequence-binding protein)
(TBP)., complete
Length = 1595
Score = 28.5 bits (62), Expect = 7.7
Identities = 15/58 (25%), Positives = 30/58 (50%)
Frame = +3
Query: 175 RRLVLL*TFFRFIKRLLDRKLIWINQRWRSAQTSLMIYRKYFKTIYLLRSALPSINIW 232
++ L+ TF + +L + +W+ + W+ A +L IY+ + LL L ++IW
Sbjct: 237 KKKFLIFTFKNSLYKLRVNRSLWLTKDWKGA--NLWIYQSTLLELCLLFKILCQLSIW 404
>TC93570 similar to GP|21553436|gb|AAM62529.1 unknown {Arabidopsis
thaliana}, partial (57%)
Length = 759
Score = 28.5 bits (62), Expect = 7.7
Identities = 16/36 (44%), Positives = 24/36 (66%)
Frame = +2
Query: 459 IHISCLLTKIK*NVSLLLIWPMMMRLCGILIKMANI 494
IHI+CL+ K+ * V LI M+ LCG+LI++ +
Sbjct: 299 IHIACLVLKLG*RV-WGLIRVRMIGLCGLLIRVIGV 403
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.344 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,361,693
Number of Sequences: 36976
Number of extensions: 314855
Number of successful extensions: 3058
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1752
Number of HSP's successfully gapped in prelim test: 136
Number of HSP's that attempted gapping in prelim test: 1268
Number of HSP's gapped (non-prelim): 1958
length of query: 600
length of database: 9,014,727
effective HSP length: 102
effective length of query: 498
effective length of database: 5,243,175
effective search space: 2611101150
effective search space used: 2611101150
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC119408.1


