

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.7 + phase: 0
(392 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
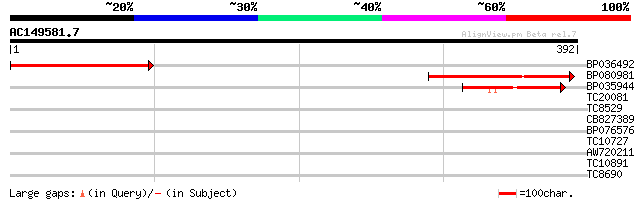
Sequences producing significant alignments: (bits) Value
BP036492 198 1e-51
BP080981 168 1e-42
BP035944 49 1e-06
TC20081 similar to UP|AAQ73157 (AAQ73157) LysM domain-containing... 28 2.6
TC8529 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-interac... 28 3.4
CB827389 28 3.4
BP076576 27 5.9
TC10727 weakly similar to UP|Q15287 (Q15287) RNA-binding protein... 27 5.9
AW720211 27 5.9
TC10891 similar to UP|Q941Q3 (Q941Q3) Floral homeotic protein HU... 27 7.7
TC8690 similar to UP|Q8RVX1 (Q8RVX1) Pectin methylesterase precu... 27 7.7
>BP036492
Length = 552
Score = 198 bits (503), Expect = 1e-51
Identities = 90/99 (90%), Positives = 94/99 (94%)
Frame = +3
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
MILTAKHHDGFCLWPSKYTKHSVISS WQ GKGDVV EFVNAA+ +GIDVGIYLSPWDRH
Sbjct: 255 MILTAKHHDGFCLWPSKYTKHSVISSPWQGGKGDVVGEFVNAASAQGIDVGIYLSPWDRH 434
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDP 99
DSRYGHDLLYNEYY+AQLQELL KYQ+VREIWFDGAKDP
Sbjct: 435 DSRYGHDLLYNEYYMAQLQELLNKYQNVREIWFDGAKDP 551
>BP080981
Length = 427
Score = 168 bits (426), Expect = 1e-42
Identities = 81/101 (80%), Positives = 90/101 (88%)
Frame = -3
Query: 290 EDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEIYVDGKSIIQGTTIGYKRLH 349
E+ + DHWIEIWG DGSLRFNVIRI EAIGLGQRI+R+EIYVDGK II+GTT+GYKRLH
Sbjct: 419 EEGENNDHWIEIWGKDGSLRFNVIRIPEAIGLGQRIKRHEIYVDGKLIIKGTTVGYKRLH 240
Query: 350 RLDGDVVHARVVRIRFIKARGVPLISSIGLHFDPFWHSRFT 390
RLDGD V+A+VVRIR KAR VPLISSIGLHFDP+WHS FT
Sbjct: 239 RLDGD-VNAQVVRIRIKKARAVPLISSIGLHFDPYWHSSFT 120
>BP035944
Length = 534
Score = 48.9 bits (115), Expect = 1e-06
Identities = 28/76 (36%), Positives = 49/76 (63%), Gaps = 5/76 (6%)
Frame = -2
Query: 314 RIQEAIGLGQRIERYEI---YVDG--KSIIQGTTIGYKRLHRLDGDVVHARVVRIRFIKA 368
++QE I +GQR+ + + + DG K +++GTTIGY+RL L + ++ +++ K+
Sbjct: 533 QVQEPIHMGQRVIEFHLEALHQDGVWKRVVKGTTIGYQRL--LLFPKIKSQFLKLIVDKS 360
Query: 369 RGVPLISSIGLHFDPF 384
R PLIS +G++ DPF
Sbjct: 359 RADPLISYLGIYIDPF 312
>TC20081 similar to UP|AAQ73157 (AAQ73157) LysM domain-containing
receptor-like kinase 6, partial (9%)
Length = 506
Score = 28.1 bits (61), Expect = 2.6
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = -1
Query: 272 GPENMLDSDHLWSYWTP 288
GP+NM+++ W YW+P
Sbjct: 206 GPKNMMENAKYWLYWSP 156
>TC8529 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-interacting
protein 1, partial (44%)
Length = 650
Score = 27.7 bits (60), Expect = 3.4
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +1
Query: 333 DGKSIIQGTTIGYKRLHRLDGDVVHARV 360
DGKS G GY RLH D D + ++
Sbjct: 358 DGKSFSSGGEDGYVRLHHFDPDYFNIKI 441
>CB827389
Length = 497
Score = 27.7 bits (60), Expect = 3.4
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = -2
Query: 284 SYWTPREDDKEKDHWIEIWGND 305
S W+P EDD+ ++ W +WG D
Sbjct: 94 SSWSPGEDDESEESW-RLWGLD 32
>BP076576
Length = 427
Score = 26.9 bits (58), Expect = 5.9
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -2
Query: 6 KHHDGFCLWPSKYTKHSVISSKW 28
+ +G WP K+T +VIS KW
Sbjct: 258 RQEEGERRWPEKFTVAAVISEKW 190
>TC10727 weakly similar to UP|Q15287 (Q15287) RNA-binding protein
(RNA-binding protein S1, serine-rich domain), partial
(15%)
Length = 932
Score = 26.9 bits (58), Expect = 5.9
Identities = 13/24 (54%), Positives = 15/24 (62%), Gaps = 3/24 (12%)
Frame = +1
Query: 274 ENMLDSD---HLWSYWTPREDDKE 294
EN L SD LWSYW+ ED+ E
Sbjct: 526 ENKLYSDLILGLWSYWSSEEDEVE 597
>AW720211
Length = 465
Score = 26.9 bits (58), Expect = 5.9
Identities = 10/13 (76%), Positives = 12/13 (91%)
Frame = +2
Query: 1 MILTAKHHDGFCL 13
++LTAKHHDGF L
Sbjct: 419 VLLTAKHHDGFLL 457
>TC10891 similar to UP|Q941Q3 (Q941Q3) Floral homeotic protein HUA1, partial
(15%)
Length = 581
Score = 26.6 bits (57), Expect = 7.7
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = -3
Query: 1 MILTAKHHDGFCLWPSKYTKH 21
M+L+ + DG C++P YT+H
Sbjct: 192 MVLSGQLGDGTCIFPQIYTRH 130
>TC8690 similar to UP|Q8RVX1 (Q8RVX1) Pectin methylesterase precursor
(Pectinesterase) , partial (59%)
Length = 1082
Score = 26.6 bits (57), Expect = 7.7
Identities = 29/111 (26%), Positives = 46/111 (41%), Gaps = 7/111 (6%)
Frame = +3
Query: 258 VKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDKEKDHWIE------IWGNDGSLRFN 311
V V + +GG E + LD D + W R+D + + + + DGS RF
Sbjct: 672 VSVLAPKGGNEQII--DEPLDGD--FPSWVTRKDRRLLESSVGDVNANVVVAKDGSGRFK 839
Query: 312 VIRIQEAIGLGQRIERYEIYVDGKSIIQGTTIGYKRLH-RLDGDVVHARVV 361
+ A RY IYV + + IG K+ + L GD + A ++
Sbjct: 840 TVAEAVASAPDSGKTRYVIYVKKGTYKENIEIGKKKTNVMLTGDGMDATII 992
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,617,437
Number of Sequences: 28460
Number of extensions: 104033
Number of successful extensions: 548
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 540
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 546
length of query: 392
length of database: 4,897,600
effective HSP length: 92
effective length of query: 300
effective length of database: 2,279,280
effective search space: 683784000
effective search space used: 683784000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149581.7


