

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.3 + phase: 0
(194 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
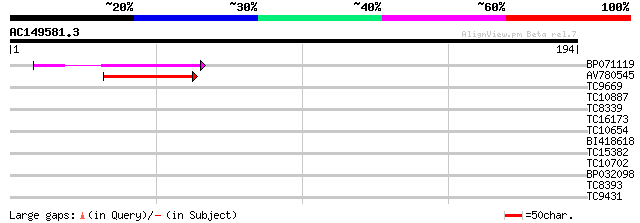
Score E
Sequences producing significant alignments: (bits) Value
BP071119 47 3e-06
AV780545 39 6e-04
TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%) 31 0.12
TC10887 similar to UP|Q84JJ4 (Q84JJ4) Flavonoid 3'-hydroxylase (... 28 1.4
TC8339 homologue to UP|Q84L89 (Q84L89) L-asparaginase, complete 27 1.8
TC16173 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Ar... 26 3.9
TC10654 similar to UP|Q8S0P2 (Q8S0P2) Ribosomal protein L28-like... 26 3.9
BI418618 26 5.2
TC15382 similar to UP|MSK2_MEDSA (P51138) Glycogen synthase kina... 26 5.2
TC10702 26 5.2
BP032098 25 6.7
TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenyla... 25 8.8
TC9431 weakly similar to UP|Q84UI1 (Q84UI1) Allergen len c 3.02 ... 25 8.8
>BP071119
Length = 412
Score = 46.6 bits (109), Expect = 3e-06
Identities = 26/59 (44%), Positives = 32/59 (54%)
Frame = +2
Query: 9 AGLRWFWEPVEGFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDM 67
AGLRW + +V ++I A ERWH ETSSFHL GEMTI L+D+
Sbjct: 266 AGLRWVMKCTS------------PSVDQSIISAFVERWHPETSSFHLPWGEMTITLEDV 406
>AV780545
Length = 555
Score = 38.9 bits (89), Expect = 6e-04
Identities = 16/32 (50%), Positives = 24/32 (75%)
Frame = +1
Query: 33 TVTHAMICAMCERWHTETSSFHLLVGEMTIPL 64
++ + +I A+ ERW ET++FHL VGEMT+ L
Sbjct: 460 SLDNPLISALVERWRRETNTFHLNVGEMTVTL 555
>TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%)
Length = 472
Score = 31.2 bits (69), Expect = 0.12
Identities = 24/69 (34%), Positives = 35/69 (49%), Gaps = 5/69 (7%)
Frame = -2
Query: 52 SFHL-LVGEMTIPLDDMHNLLHIPI---HGRILDHDEAMNQDCAIDL-MIRLLGMSEVDV 106
SFHL +V M + ++H LLH+ + HG + HDE M +D + + R L V
Sbjct: 366 SFHLWVVMVMKVGCSNLH-LLHVRVECFHGHLQSHDEEMEEDSCVSIENQRNLWEESQHV 190
Query: 107 RVEVRSESA 115
+E S SA
Sbjct: 189 DLECNSSSA 163
>TC10887 similar to UP|Q84JJ4 (Q84JJ4) Flavonoid 3'-hydroxylase (Fragment),
partial (43%)
Length = 703
Score = 27.7 bits (60), Expect = 1.4
Identities = 12/43 (27%), Positives = 23/43 (52%)
Frame = -1
Query: 15 WEPVEGFDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLV 57
++ ++GF ++YT + H + W+T TSSF L++
Sbjct: 670 FKQMQGFHTQ*ILYTPKFEIHHISPFRIFTNWYTTTSSFLLII 542
>TC8339 homologue to UP|Q84L89 (Q84L89) L-asparaginase, complete
Length = 1467
Score = 27.3 bits (59), Expect = 1.8
Identities = 10/24 (41%), Positives = 18/24 (74%)
Frame = -2
Query: 145 KLQERGRKRAIGKWSWGGMALAYL 168
KL ++G+K+ + WGG++LA+L
Sbjct: 1448 KLVKKGKKQKATRLFWGGLSLAFL 1377
>TC16173 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;}, partial (37%)
Length = 601
Score = 26.2 bits (56), Expect = 3.9
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = +3
Query: 11 LRWFWEPVEGFDLHD 25
++WFWE V+GF D
Sbjct: 93 IQWFWEVVQGFSKED 137
>TC10654 similar to UP|Q8S0P2 (Q8S0P2) Ribosomal protein L28-like, complete
Length = 510
Score = 26.2 bits (56), Expect = 3.9
Identities = 21/62 (33%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Frame = -1
Query: 68 HNLLHIPIH-GRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRV 126
H +LH H ++L HD +NQ C + L++ LG S+ + RS + GH RV
Sbjct: 366 HLILHCLSHPAKLLLHDRLVNQSCRLGLLL-*LGRSQQHRLIFARSLN-GHSLLVGQTRV 193
Query: 127 YE 128
E
Sbjct: 192 LE 187
>BI418618
Length = 601
Score = 25.8 bits (55), Expect = 5.2
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = +1
Query: 117 HISYPTLKRVYEHHLTEAR 135
HI++ TLK ++ HH TE R
Sbjct: 160 HITHQTLKDLWTHHDTETR 216
>TC15382 similar to UP|MSK2_MEDSA (P51138) Glycogen synthase kinase-3
homolog MsK-2 , partial (17%)
Length = 552
Score = 25.8 bits (55), Expect = 5.2
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = -2
Query: 145 KLQERGRKRAIGKWSWG 161
KL+ GRKR GKW +G
Sbjct: 119 KLKSGGRKRPFGKWVFG 69
>TC10702
Length = 593
Score = 25.8 bits (55), Expect = 5.2
Identities = 9/32 (28%), Positives = 18/32 (56%)
Frame = +3
Query: 11 LRWFWEPVEGFDLHDLIYTGYSTVTHAMICAM 42
+ +FW P+E F L+ Y ++ V ++C +
Sbjct: 384 INFFWHPIELFQLNFCRYMLFACVLRHVVCCV 479
>BP032098
Length = 508
Score = 25.4 bits (54), Expect = 6.7
Identities = 9/19 (47%), Positives = 16/19 (83%)
Frame = +3
Query: 121 PTLKRVYEHHLTEARRLEE 139
PT+K + H++T+A++LEE
Sbjct: 39 PTVKYIGGHNITQAKKLEE 95
>TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenylated Rab
receptor 2 {Arabidopsis thaliana;}, partial (78%)
Length = 1012
Score = 25.0 bits (53), Expect = 8.8
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = -2
Query: 138 EEPQTREKLQERGRKRAIGKWS 159
+EP+ R + QERG +R I +WS
Sbjct: 702 KEPRRRRRQQERGGRRGI-RWS 640
>TC9431 weakly similar to UP|Q84UI1 (Q84UI1) Allergen len c 3.02
(Fragment), partial (67%)
Length = 1328
Score = 25.0 bits (53), Expect = 8.8
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +2
Query: 134 ARRLEEPQTREKLQERGRKRAIGKW 158
+RRLE+P RE+++ER + KW
Sbjct: 230 SRRLEDPDERERVRERTERAK--KW 298
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,410,610
Number of Sequences: 28460
Number of extensions: 44881
Number of successful extensions: 307
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 307
length of query: 194
length of database: 4,897,600
effective HSP length: 85
effective length of query: 109
effective length of database: 2,478,500
effective search space: 270156500
effective search space used: 270156500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC149581.3


