

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.10 + phase: 2 /partial
(349 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
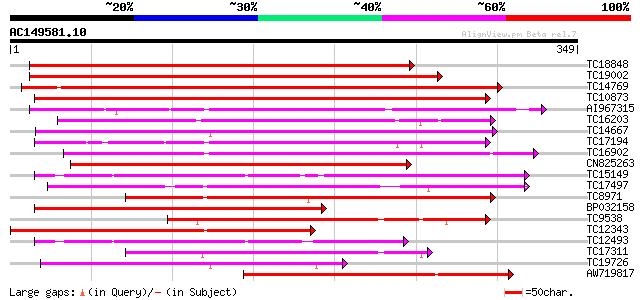
Score E
Sequences producing significant alignments: (bits) Value
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 369 e-103
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 363 e-101
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 234 1e-62
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 230 2e-61
AI967315 211 2e-55
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 205 1e-53
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 199 6e-52
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 198 1e-51
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 191 2e-49
CN825263 185 9e-48
TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pro... 181 2e-46
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 173 4e-44
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 169 6e-43
BP032158 167 2e-42
TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 160 4e-40
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 157 3e-39
TC12493 similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (35%) 155 7e-39
TC17311 similar to UP|Q8K2Y2 (Q8K2Y2) Receptor-interacting serin... 152 6e-38
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 143 4e-35
AW719817 141 1e-34
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 369 bits (946), Expect = e-103
Identities = 173/237 (72%), Positives = 201/237 (83%)
Frame = +2
Query: 13 WERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVE 72
W +T KEL AT F DNK+GEGGFGSVYWG+TS G++IAVK+LK M +KAEMEFAVE
Sbjct: 257 WRIFTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFAVE 436
Query: 73 VEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSIT 132
VEVLGRVRHKNLLGLRG+ G D+RLIVYDYM N SLL+HLHGQ A + L+W +RM I
Sbjct: 437 VEVLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIA 616
Query: 133 VGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKG 192
+G+AEG+ YLHHE PHIIHRDIKASNVLL+++F+ VADFGFAKLIP GVSH+TTRVKG
Sbjct: 617 IGSAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMTTRVKG 796
Query: 193 TLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYV 249
TLGYLAPEYAMWGKVSESCDVYSFGILLLE+++ +KPIEKLPGG+KR I +W P +
Sbjct: 797 TLGYLAPEYAMWGKVSESCDVYSFGILLLELVTGRKPIEKLPGGVKRTITEWAEPLI 967
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 363 bits (932), Expect = e-101
Identities = 175/254 (68%), Positives = 204/254 (79%)
Frame = +1
Query: 13 WERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVE 72
W ++LKEL ATNNF+ DNK+GEGGFGSVYWGQ G +IAVKRLK + KA+MEFAVE
Sbjct: 19 WRVFSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVE 198
Query: 73 VEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSIT 132
VE+L RVRHKNLL LRG+ A G ERLIVYDYM N SLL+HLHGQ +S+CLLDW RRM+I
Sbjct: 199 VEILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIA 378
Query: 133 VGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKG 192
+G+AEG+ YLHH+A PHIIHRDIKASNVLLD++FQA+VADFGFAKLIP G +H+TTRVKG
Sbjct: 379 IGSAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTTRVKG 558
Query: 193 TLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKG 252
TLGYLAPEYAM GK +E CDV+SFGILLLE+ S KKP+EKL +KR I W P
Sbjct: 559 TLGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAK 738
Query: 253 VFKHIADPKLKGNF 266
F ADP+L G +
Sbjct: 739 KFTEFADPRLNGEY 780
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 234 bits (598), Expect = 1e-62
Identities = 123/297 (41%), Positives = 185/297 (61%), Gaps = 1/297 (0%)
Frame = +3
Query: 8 IRDYPWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEM 67
I+ + +TL+ + AT + IGEGGFGSVY G + G E+AVK + + +
Sbjct: 2109 IKSVSIQAFTLEYIEVATERYK--TLIGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTR 2282
Query: 68 EFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPR 127
EF E+ +L ++H+NL+ L G+ D++++VY +MSN SL L+G+ A +LDWP
Sbjct: 2283 EFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPT 2462
Query: 128 RMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIP-AGVSHL 186
R+SI +GAA GLAYLH +IHRDIK+SN+LLD AKVADFGF+K P G S++
Sbjct: 2463 RLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYV 2642
Query: 187 TTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVT 246
+ V+GT GYL PEY ++SE DV+SFG++LLEI+S ++P+ + +V+W T
Sbjct: 2643 SLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWAT 2822
Query: 247 PYVQKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDGV 303
PY++ I DP +KG + E + V+ +A++C + RPSM+ +V L+D +
Sbjct: 2823 PYIRGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVRELEDAL 2993
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 230 bits (587), Expect = 2e-61
Identities = 122/283 (43%), Positives = 173/283 (61%), Gaps = 2/283 (0%)
Frame = +1
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEV 75
+ +EL AT NF + N IGEGGFG VY G+ + G +AVK+L + EF +EV +
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLM 594
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGA 135
L + H NL+ L G+ GD+RL+VY+YM SL HL L+W RM + VGA
Sbjct: 595 LSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGA 774
Query: 136 AEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAG-VSHLTTRVKGTL 194
A GL YLH A+P +I+RD+K++N+LLD EF K++DFG AKL P G +H++TRV GT
Sbjct: 775 ARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVSTRVMGTY 954
Query: 195 GYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYV-QKGV 253
GY APEYAM GK++ D+YSFG++LLE+++ ++ I+ ++++V W PY +
Sbjct: 955 GYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDRRR 1134
Query: 254 FKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVV 296
F H+ DP L+G F L I I C P RP + ++V
Sbjct: 1135FGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIV 1263
>AI967315
Length = 1308
Score = 211 bits (536), Expect = 2e-55
Identities = 122/323 (37%), Positives = 184/323 (56%), Gaps = 5/323 (1%)
Frame = +1
Query: 13 WERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAK---AEMEF 69
W+ ++ +EL ATN F +N +G+GG+ VY G+ G EIAVKRL T T + E EF
Sbjct: 103 WKCFSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRL-TRTCRDERKEKEF 279
Query: 70 AVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRM 129
E+ +G V H N++ L G L V++ + S+ + +H + + LDW R
Sbjct: 280 LTEIGTIGHVCHSNVMPLLGCCIDNGLYL-VFELSTVGSVASLIHDEKMAP--LDWKTRY 450
Query: 130 SITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPA-GVSHLTT 188
I +G A GL YLH IIHRDIKASN+LL +F+ +++DFG AK +P+ H
Sbjct: 451 KIVLGTARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIA 630
Query: 189 RVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPY 248
++GT G+LAPEY M G V E DV++FG+ LLE+IS +KP++ G + + W P
Sbjct: 631 PIEGTFGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVD----GSHQSLHTWAKPI 798
Query: 249 VQKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDG-VSKRK 307
+ K + + DP+L+G +D+ Q V A C +S RP+M EV+E +++G + K +
Sbjct: 799 LSKWEIEKLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVMEEGEMDKER 978
Query: 308 KEIPNLSNNKGHDEENDENYEEF 330
++P +EE D+ EEF
Sbjct: 979 WKMPK-------EEEEDQQGEEF 1026
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 205 bits (521), Expect = 1e-53
Identities = 115/279 (41%), Positives = 165/279 (58%), Gaps = 9/279 (3%)
Frame = +2
Query: 30 QDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTM-TAKAEMEFAVEVEVLGRVRHKNLLGLR 88
++N IG+GG G VY G G ++A+KRL + + + F E+E LG++RH+N++ L
Sbjct: 2198 EENIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLL 2377
Query: 89 GFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAEGLAYLHHEANP 148
G+ + D L++Y+YM N SL LHG A L W R I V AA GL Y+HH+ +P
Sbjct: 2378 GYVSNKDTNLLLYEYMPNGSLGEWLHG--AKGGHLRWEMRYKIAVEAARGLCYMHHDCSP 2551
Query: 149 HIIHRDIKASNVLLDTEFQAKVADFGFAK-LIPAGVSHLTTRVKGTLGYLAPEYAMWGKV 207
IIHRD+K++N+LLD +F+A VADFG AK L G S + + G+ GY+APEYA KV
Sbjct: 2552 LIIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKV 2731
Query: 208 SESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQK-------GVFKHIADP 260
E DVYSFG++LLE+I +KP+ + G+ DIV WV + + + + DP
Sbjct: 2732 DEKSDVYSFGVVLLELIIGRKPVGEFGDGV--DIVGWVNKTMSELSQPSDTALVLAVVDP 2905
Query: 261 KLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWL 299
+L G + L + + IA+ C RP+M EVV L
Sbjct: 2906 RLSG-YPLTSVIHMFNIAMMCVKEMGPARPTMREVVHML 3019
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 199 bits (506), Expect = 6e-52
Identities = 108/291 (37%), Positives = 171/291 (58%), Gaps = 7/291 (2%)
Frame = +2
Query: 17 TLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTA-KAEMEFAVEVEV 75
+L EL T+NF IGEG +G VY+ + G +AVK+L + + EF +V +
Sbjct: 326 SLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQVSM 505
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCL-----LDWPRRMS 130
+ R+++ N + L G+ G+ R++ Y++ + SL LHG+ LDW +R+
Sbjct: 506 VSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVR 685
Query: 131 ITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHL-TTR 189
I V AA GL YLH + P IIHRDI++SNVL+ +++AK+ADF + P + L +TR
Sbjct: 686 IAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTR 865
Query: 190 VKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYV 249
V GT GY APEYAM G++++ DVYSFG++LLE+++ +KP++ ++ +V W TP +
Sbjct: 866 VLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRL 1045
Query: 250 QKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLK 300
+ K DPKLKG + + + + +A C + RP+M VV+ L+
Sbjct: 1046SEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQ 1198
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 198 bits (503), Expect = 1e-51
Identities = 111/287 (38%), Positives = 167/287 (57%), Gaps = 6/287 (2%)
Frame = +1
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEV 75
++ +EL +ATNNF DNKIG+GGFG+VY+ + +G + A+K+ M +A EF E++V
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAEL-RGKKTAIKK---MDVQASTEFLCELKV 1218
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGA 135
L V H NL+ L G+ G +VY+++ N +L +LHG L W R+ I + A
Sbjct: 1219 LTHVHHLNLVRLIGYCVEGS-LFLVYEHIDNGNLGQYLHGSGKEP--LPWSSRVQIALDA 1389
Query: 136 AEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTLG 195
A GL Y+H P IHRD+K++N+L+D + KVADFG KLI G S L TR+ GT G
Sbjct: 1390 ARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQTRLVGTFG 1569
Query: 196 YLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGI--KRDIVQWVTPYVQKG- 252
Y+ PEYA +G +S DVY+FG++L E+ISAK + K + + +V + K
Sbjct: 1570 YMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNKSD 1749
Query: 253 ---VFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVV 296
+ + DP+L N+ ++ + + + CT +P RPSM +V
Sbjct: 1750 PCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLV 1890
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 191 bits (485), Expect = 2e-49
Identities = 105/293 (35%), Positives = 165/293 (55%), Gaps = 1/293 (0%)
Frame = +3
Query: 34 IGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEVLGRVRHKNLLGLRGFYAG 93
+G GGFG VY+G+ G ++A+KR ++ + EF E+E+L ++RH++L+ L G+
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEE 194
Query: 94 GDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAEGLAYLHHEANPHIIHR 153
E ++VYD+M+ +L HL+ L W +R+ I +GAA GL YLH A IIHR
Sbjct: 195 NTEMILVYDHMAYGTLREHLYKTQKPP--LPWKQRLEICIGAARGLHYLHTGAKYTIIHR 368
Query: 154 DIKASNVLLDTEFQAKVADFGFAKLIPA-GVSHLTTRVKGTLGYLAPEYAMWGKVSESCD 212
D+K +N+LLD ++ AKV+DFG +K P +H++T VKG+ GYL PEY ++++ D
Sbjct: 369 DVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSD 548
Query: 213 VYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKHIADPKLKGNFDLEQLK 272
VYSFG++L EI+ A+ + + + +W + KG+ I DP LKG E K
Sbjct: 549 VYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFK 728
Query: 273 SVIMIAVRCTDSSPDKRPSMIEVVEWLKDGVSKRKKEIPNLSNNKGHDEENDE 325
A++C +RPSM +V+ W + + ++ N G DE
Sbjct: 729 KFAETAMKCVSDQGIERPSMGDVL-WNLEFALQLQESAEESGNGFGGIHGEDE 884
>CN825263
Length = 663
Score = 185 bits (470), Expect = 9e-48
Identities = 94/211 (44%), Positives = 133/211 (62%), Gaps = 1/211 (0%)
Frame = +1
Query: 38 GFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEVLGRVRHKNLLGLRGFYAGGDER 97
GFG VY G + G ++AVK LK + EF EVE+L R+ H+NL+ L G R
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 98 LIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAEGLAYLHHEANPHIIHRDIKA 157
++Y+ + N S+ +HLHG LDW RM I +GAA GLAYLH ++NP +IHRD K+
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 158 SNVLLDTEFQAKVADFGFAK-LIPAGVSHLTTRVKGTLGYLAPEYAMWGKVSESCDVYSF 216
SN+LL+ +F KV+DFG A+ + G H++T V GT GYLAPEYAM G + DVYS+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 217 GILLLEIISAKKPIEKLPGGIKRDIVQWVTP 247
G++LLE+++ KP++ + ++V W P
Sbjct: 541 GVVLLELLTGTKPVDLSQPPGQENLVTWARP 633
>TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (45%)
Length = 1489
Score = 181 bits (459), Expect = 2e-46
Identities = 116/308 (37%), Positives = 178/308 (57%), Gaps = 3/308 (0%)
Frame = +3
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEV 75
+ L++LLRA+ +G+G FG+ Y G+ +AVKRLK +T +E EF ++E
Sbjct: 255 FDLEDLLRASAEV-----LGKGTFGTAYKAVLETGLVVAVKRLKDVTI-SEKEFKDKIET 416
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQL-ASDCLLDWPRRMSITVG 134
+G + H++L+ LR +Y DE+L+VYDYM SL LHG A L+W R I +G
Sbjct: 417 VGAMDHQSLVPLRAYYFSRDEKLLVYDYMPMGSLSALLHGNKGAGRTPLNWEIRSGIALG 596
Query: 135 AAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTL 194
AA G+ YLH + P++ H +IKASN+LL ++AKV+DFG A L+ G S RV
Sbjct: 597 AARGIEYLHSQ-GPNVSHGNIKASNILLTKSYEAKVSDFGLAHLV--GPSSTPNRV---A 758
Query: 195 GYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVF 254
GY APE KVS+ DVYSFG+LLLE+++ K P L D+ +WV V++
Sbjct: 759 GYRAPEVTDPRKVSQKADVYSFGVLLLELLTGKAPTHALLNEEGVDLPRWVQSLVREEWT 938
Query: 255 KHIADPKLKGNFDL-EQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDGV-SKRKKEIPN 312
+ D +L ++ E++ ++ +AV C PD+RPS+ EV +++ S K++
Sbjct: 939 SEVFDLELLRYQNVEEEMVQLLQLAVDCAAPYPDRRPSIAEVTRSIEELCRSNLKEDQDQ 1118
Query: 313 LSNNKGHD 320
+ +++ HD
Sbjct: 1119IPHDQIHD 1142
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 173 bits (438), Expect = 4e-44
Identities = 111/310 (35%), Positives = 158/310 (50%), Gaps = 13/310 (4%)
Frame = +1
Query: 24 ATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEVLGRVRHKN 83
ATN+F NK+GEGGFG VY G + G EIAVKRL + + EF E++++ R++H+N
Sbjct: 1660 ATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKLIARLQHRN 1839
Query: 84 LLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGAAEGLAYLH 143
L+ L G DE + +N + L + L+DW +R+ I G A GL YLH
Sbjct: 1840 LVKLFGCSVHQDE-----NSHANKKMKILLDSTRSK--LVDWNKRLQIIDGIARGLLYLH 1998
Query: 144 HEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKL-IPAGVSHLTTRVKGTLGYLAPEYA 202
++ IIHRD+K SN+LLD E K++DFG A++ I V T RV GT GY+ PEYA
Sbjct: 1999 QDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEYA 2178
Query: 203 MWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKH------ 256
+ G S DV+SFG+++LEIIS KK I ++ P+ + H
Sbjct: 2179 VHGSFSIKSDVFSFGVIVLEIISGKK------------IGRFYDPHHHLNLLSHAWRLWI 2322
Query: 257 ------IADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDGVSKRKKEI 310
+ D L ++ I +A+ C P+ RP M+ +V L K +
Sbjct: 2323 EERPLELVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIVLMLNGEKELPKPRL 2502
Query: 311 PNLSNNKGHD 320
P K HD
Sbjct: 2503 PAFYTGK-HD 2529
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 169 bits (428), Expect = 6e-43
Identities = 90/230 (39%), Positives = 140/230 (60%), Gaps = 2/230 (0%)
Frame = +3
Query: 72 EVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSI 131
EVE+L ++ H+NL+ L GF G+ER+++ +Y+ N +L HL G +LD+ +R+ I
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGK--ILDFNQRLEI 179
Query: 132 TVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAG--VSHLTTR 189
+ A GL YLH A IIHRD+K+SN+LL +AKVADFGFA+L P +H++T+
Sbjct: 180 AIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTK 359
Query: 190 VKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYV 249
VKGT+GYL PEY +++ DVYSFGILLLEI++ ++P+E +R ++W
Sbjct: 360 VKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY 539
Query: 250 QKGVFKHIADPKLKGNFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWL 299
+G + DP ++ + + L ++ ++ +C RP+M V E L
Sbjct: 540 NEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 689
>BP032158
Length = 555
Score = 167 bits (423), Expect = 2e-42
Identities = 85/180 (47%), Positives = 120/180 (66%)
Frame = +2
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEV 75
++L+++ ATNNF NKIGEGGFG VY G S+G IAVK+L + + + EF E+ +
Sbjct: 14 FSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIGM 193
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITVGA 135
+ ++H NL+ L G G++ L+VY+YM N+SL L G L+W RM I VG
Sbjct: 194 ISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVGI 373
Query: 136 AEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTLG 195
A+GLAYLH E+ I+HRDIKA+NVLLD + +AK++DFG AKL +H++TR+ GT+G
Sbjct: 374 AKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHISTRIAGTIG 553
>TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 1067
Score = 160 bits (404), Expect = 4e-40
Identities = 87/212 (41%), Positives = 129/212 (60%), Gaps = 13/212 (6%)
Frame = +2
Query: 98 LIVYDYMSNHSLLTHLH---------GQLASDCLLDWPRRMSITVGAAEGLAYLHHEANP 148
L+VY+Y+ NHSL LH G + +LDWP+R+ I +GAA+GL+Y+HH+ +P
Sbjct: 50 LLVYEYLENHSLDKWLHLKPKSSSVSGVVQQYTVLDWPKRLKIAIGAAQGLSYMHHDCSP 229
Query: 149 HIIHRDIKASNVLLDTEFQAKVADFGFAK-LIPAGVSHLTTRVKGTLGYLAPEYAMWGKV 207
I+HRD+K SN+LLD +F AKVADFG A+ LI G ++ + V GT GY+APEY ++
Sbjct: 230 PIVHRDVKTSNILLDKQFNAKVADFGLARMLIKPGELNIMSTVIGTFGYIAPEYVQTTRI 409
Query: 208 SESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKHIADPKLKGNFD 267
SE DVYSFG++LLE+ + K E G + +W ++ G ++ D K +
Sbjct: 410 SEKVDVYSFGVVLLELTTGK---EANYGDQHSSLAEWAWRHILIG--SNVXDLLXKDVME 574
Query: 268 ---LEQLKSVIMIAVRCTDSSPDKRPSMIEVV 296
++++ SV + V CT + P RPSM EV+
Sbjct: 575 ASYIDEMCSVFKLGVMCTATLPATRPSMKEVL 670
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 157 bits (396), Expect = 3e-39
Identities = 81/188 (43%), Positives = 119/188 (63%)
Frame = +3
Query: 1 NVDDKKNIRDYPWERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKT 60
N DD +NI + ++ + L+ AT NFH NK+GEGGFG V+ G+ + G EIAVK+L
Sbjct: 162 NEDDIQNIAAKEHKTFSYETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKKLSR 341
Query: 61 MTAKAEMEFAVEVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASD 120
+ + +F E ++L RV+H+N++ L G+ A G E+L+VY+Y+ SL L +
Sbjct: 342 RSNQGRTQFINEAKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQKKE 521
Query: 121 CLLDWPRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIP 180
LDW RR I G A GL YLH +++ IIHRDIKA+N+LLD ++ K+ADFG A++ P
Sbjct: 522 -QLDWKRRFDIISGVARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARIFP 698
Query: 181 AGVSHLTT 188
+H+ T
Sbjct: 699 EDQTHVNT 722
>TC12493 similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (35%)
Length = 837
Score = 155 bits (393), Expect = 7e-39
Identities = 97/232 (41%), Positives = 135/232 (57%), Gaps = 2/232 (0%)
Frame = +3
Query: 16 YTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEVEV 75
++L ELLRA+ +G+G FG+ Y G +AVKRLK +TA EMEF ++E
Sbjct: 177 FSLDELLRASAEV-----LGKGTFGTTYKATMEMGRSVAVKRLKDVTA-TEMEFREKIEE 338
Query: 76 LGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQL-ASDCLLDWPRRMSITVG 134
+G++ H+NL+ LRG+Y DE+L+VYDYM SL LH A L+W R +I +G
Sbjct: 339 VGKLVHENLVPLRGYYFSRDEKLVVYDYMPMGSLSALLHANNGAGRTPLNWETRSAIALG 518
Query: 135 AAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKL-IPAGVSHLTTRVKGT 193
AA G+AYLH + P H +IK+SN+LL F+ +V+D G A L +P T+
Sbjct: 519 AAHGIAYLHSQ-GPTSSHGNIKSSNILLTKSFEPRVSDLGLAYLALP------TSTPNRI 677
Query: 194 LGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWV 245
GY APE KVS+ DVYSFGI+LLE+++ K P D+ +WV
Sbjct: 678 SGYRAPEVTDARKVSQKADVYSFGIMLLELLTGKPPTHSSLNEEGVDLPRWV 833
>TC17311 similar to UP|Q8K2Y2 (Q8K2Y2) Receptor-interacting serine-threonine
kinase 3, partial (7%)
Length = 606
Score = 152 bits (385), Expect = 6e-38
Identities = 81/204 (39%), Positives = 123/204 (59%), Gaps = 15/204 (7%)
Frame = +3
Query: 72 EVEVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQL--------ASDCLL 123
EVE+L +RH N++ L+ + + L+VYDY+ N SL LH + + ++
Sbjct: 3 EVEILSNIRHNNIVKLQCCISSEESLLLVYDYLENLSLDRWLHKKSKPPTVSGSVQNNII 182
Query: 124 DWPRRMSITVGAAEGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGV 183
DWPRR+ I +G A+GL Y+HH+ +P ++HRD+K SN+LLD++F AKVADFG A +
Sbjct: 183 DWPRRLHIAIGVAQGLCYMHHDCSPPVVHRDLKPSNILLDSQFNAKVADFGLAMMSVKPE 362
Query: 184 SHLT-TRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIV 242
T + V GT GY+APEYA+ +V+E DVYSFG++LLE+ + K E G +
Sbjct: 363 ELATMSAVAGTFGYIAPEYALTIRVNEKIDVYSFGVILLELTTGK---EANHGDEYSSLA 533
Query: 243 QWVTPYVQKG------VFKHIADP 260
+W ++Q G + + IA+P
Sbjct: 534 EWARRHIQIGSNIEDLLVEEIAEP 605
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 143 bits (361), Expect = 4e-35
Identities = 80/194 (41%), Positives = 116/194 (59%), Gaps = 5/194 (2%)
Frame = +3
Query: 20 ELLRATNNFHQDNKIGEGGFGSVYWGQTSKG-VEIAVKRLKTMTAKAEMEFAVEVEVLGR 78
EL +ATNNF + +K+G+GG+G VY G K +E+AVK K+ +F E+ ++ R
Sbjct: 144 ELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSELIIINR 323
Query: 79 VRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCL--LDWPRRMSITVGAA 136
+RHK+L+ L+G+ L+VYDYM N SL +H+ + L WP R I G A
Sbjct: 324 LRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKIISGVA 503
Query: 137 EGLAYLHHEANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLT--TRVKGTL 194
L YLH+E + ++HRD+KASN++LD+EF AK+ DFG A+ + T V GT+
Sbjct: 504 SALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTELEGVHGTM 683
Query: 195 GYLAPEYAMWGKVS 208
GY+APE GK +
Sbjct: 684 GYIAPECFHTGKAT 725
>AW719817
Length = 499
Score = 141 bits (356), Expect = 1e-34
Identities = 68/166 (40%), Positives = 105/166 (62%)
Frame = +1
Query: 145 EANPHIIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTLGYLAPEYAMW 204
E PHI+HRDIKASN+LLD +F K+ DFG AKL P ++H++TR+ GT GYLAPEYAM
Sbjct: 1 ELVPHIVHRDIKASNILLDRDFNPKIGDFGLAKLFPDDITHISTRIAGTTGYLAPEYAMG 180
Query: 205 GKVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKHIADPKLKG 264
G+++ DVYSFG+L+LE++S K GG + +++W ++G + DP +
Sbjct: 181 GQLTMKADVYSFGVLILEVVSGKGSARTNWGGSNKYLLEWAWQLHEEGKLLELVDPDM-D 357
Query: 265 NFDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDGVSKRKKEI 310
+F E + + +A CT ++ +RP M +VV+ L + +K++
Sbjct: 358 DFPEE*VIRYMKVAFFCTQAAASRRPLMSQVVDMLTKKIRLNEKQL 495
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,561,450
Number of Sequences: 28460
Number of extensions: 70822
Number of successful extensions: 841
Number of sequences better than 10.0: 343
Number of HSP's better than 10.0 without gapping: 664
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 671
length of query: 349
length of database: 4,897,600
effective HSP length: 91
effective length of query: 258
effective length of database: 2,307,740
effective search space: 595396920
effective search space used: 595396920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149581.10


