

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.6 - phase: 0
(597 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
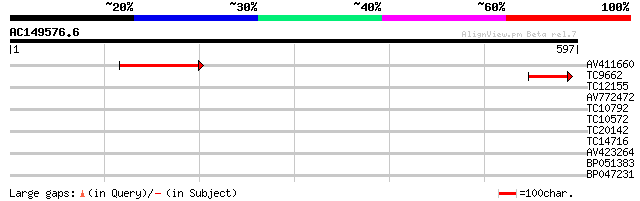
Score E
Sequences producing significant alignments: (bits) Value
AV411660 182 1e-46
TC9662 79 3e-15
TC12155 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, par... 31 0.65
AV772472 30 1.5
TC10792 29 2.5
TC10572 28 3.2
TC20142 28 4.2
TC14716 similar to UP|RNP2_HUMAN (Q14498) RNA-binding region con... 28 4.2
AV423264 28 4.2
BP051383 28 5.5
BP047231 28 5.5
>AV411660
Length = 270
Score = 182 bits (463), Expect = 1e-46
Identities = 86/89 (96%), Positives = 87/89 (97%)
Frame = +2
Query: 116 RFSLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVE 175
RFSLGTAIGFRIRGGVLTDIPAILVFVA KVHRQWLNHVQCLPAALEGPGGVWCDVDVVE
Sbjct: 2 RFSLGTAIGFRIRGGVLTDIPAILVFVARKVHRQWLNHVQCLPAALEGPGGVWCDVDVVE 181
Query: 176 FSYYGAPAPTPKEQLYTELADGLRGSDSC 204
FSYYGAPA TPKEQLYT+LADGLRGSDSC
Sbjct: 182 FSYYGAPAQTPKEQLYTDLADGLRGSDSC 268
>TC9662
Length = 638
Score = 78.6 bits (192), Expect = 3e-15
Identities = 38/46 (82%), Positives = 41/46 (88%)
Frame = +2
Query: 547 VHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSS 592
V Q + KED++FKSLSELRN PDEDNFVSL LGEPEAKRRKHSNSS
Sbjct: 2 VRQRYPKEDLEFKSLSELRNGPDEDNFVSLPLGEPEAKRRKHSNSS 139
>TC12155 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, partial (8%)
Length = 483
Score = 30.8 bits (68), Expect = 0.65
Identities = 12/35 (34%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Frame = -3
Query: 488 LEESFEPFCLNMEHVPVEE-PSTIVKPSLRPCEFH 521
+E +PFCL ++ + PS +++P ++PC FH
Sbjct: 145 METLLQPFCLKLQTSFYKGIPSRLIQPLIQPCSFH 41
>AV772472
Length = 423
Score = 29.6 bits (65), Expect = 1.5
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = +3
Query: 147 HRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPA 183
H++W++H Q ++ GP G CD F++ GA A
Sbjct: 312 HQRWVSHSQQQSSSPAGPSGSSCDFFSDAFAFSGAGA 422
>TC10792
Length = 480
Score = 28.9 bits (63), Expect = 2.5
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Frame = +3
Query: 174 VEFSYYGAPAPTPKEQLYTELADGLRGSD--SCVGSGSQV 211
VE++ G P+E L+ EL GL GSD +C+ + S V
Sbjct: 195 VEWAVSGGMRALPREALFPELGAGLSGSDAIACLEAPSSV 314
>TC10572
Length = 526
Score = 28.5 bits (62), Expect = 3.2
Identities = 18/56 (32%), Positives = 31/56 (55%), Gaps = 1/56 (1%)
Frame = +2
Query: 7 GLSAHHSGSTQSEESALDLERNYYGHPS-SSPLHMQTFAVGVQHSEGNAAYFSWPT 61
G H++ +++ + L N +G P + PLH+ + AVGV+H N + SWP+
Sbjct: 92 GSHIHNARTSKGRPGKIRL--NDWG*PIVNGPLHLSS-AVGVRHKLANHCH*SWPS 250
>TC20142
Length = 595
Score = 28.1 bits (61), Expect = 4.2
Identities = 19/71 (26%), Positives = 31/71 (42%)
Frame = -3
Query: 504 VEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAGKSPVHQSFLKEDMQFKSLSE 563
V EP+ P + C ++ + E+ R S A + +HQ ++D+ S E
Sbjct: 443 VTEPNN*GSPLRQLC*YNAQQEVWIKHYPYKTQPRLSTASEX*IHQELDEKDIPASSARE 264
Query: 564 LRNEPDEDNFV 574
L N+P D V
Sbjct: 263 LENKPPRDQRV 231
>TC14716 similar to UP|RNP2_HUMAN (Q14498) RNA-binding region containing
protein 2 (Hepatocellular carcinoma protein 1) (Splicing
factor HCC1), partial (4%)
Length = 700
Score = 28.1 bits (61), Expect = 4.2
Identities = 8/25 (32%), Positives = 16/25 (64%)
Frame = -3
Query: 147 HRQWLNHVQCLPAALEGPGGVWCDV 171
H +H+ C P++++GP G +C +
Sbjct: 305 HEACSDHLPCKPSSMQGPWGTYCSL 231
>AV423264
Length = 262
Score = 28.1 bits (61), Expect = 4.2
Identities = 12/47 (25%), Positives = 23/47 (48%), Gaps = 18/47 (38%)
Frame = +2
Query: 502 VPVEEPSTIVK------------------PSLRPCEFHIRNEIETVP 530
+P++EP+ I+ P L+PC+FH + ++T+P
Sbjct: 65 LPLQEPNNIITNNILQGSQSQISNWGTHIPLLKPCQFHYHSMVKTIP 205
>BP051383
Length = 366
Score = 27.7 bits (60), Expect = 5.5
Identities = 16/34 (47%), Positives = 20/34 (58%), Gaps = 4/34 (11%)
Frame = +1
Query: 418 RGRLKLRV----GQPPENWTSGVDLGRLLDLLEL 447
+G L LRV GQ P W+SG+ G DLL+L
Sbjct: 151 KGHLGLRVIQEQGQGPGAWSSGLSKG*TRDLLQL 252
>BP047231
Length = 476
Score = 27.7 bits (60), Expect = 5.5
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = -1
Query: 176 FSYYGAPAPTPKEQLYTELADGLRGSDSC 204
FS+ GAP P + + ++ D RGS++C
Sbjct: 356 FSFTGAPRPELENSIRGKIIDQCRGSNAC 270
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,042,756
Number of Sequences: 28460
Number of extensions: 137951
Number of successful extensions: 659
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 656
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 659
length of query: 597
length of database: 4,897,600
effective HSP length: 95
effective length of query: 502
effective length of database: 2,193,900
effective search space: 1101337800
effective search space used: 1101337800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149576.6


