

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.4 + phase: 0 /pseudo
(548 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
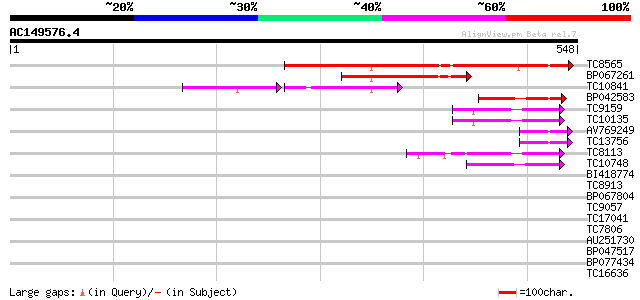
Sequences producing significant alignments: (bits) Value
TC8565 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, pa... 268 2e-72
BP067261 132 2e-31
TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1,... 86 6e-24
BP042583 74 5e-14
TC9159 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose tr... 55 3e-08
TC10135 50 9e-07
AV769249 47 6e-06
TC13756 similar to UP|Q9ARI9 (Q9ARI9) Phosphate transporter 1, p... 47 8e-06
TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter li... 45 3e-05
TC10748 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter,... 43 1e-04
BI418774 33 0.16
TC8913 similar to UP|Q7XYW2 (Q7XYW2) Seed specific protein Bn15D... 30 0.77
BP067804 30 0.77
TC9057 29 1.7
TC17041 homologue to GB|AAL25568.1|16648753|AY058152 AT5g16150/T... 29 1.7
TC7806 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1,... 29 2.2
AU251730 28 2.9
BP047517 28 2.9
BP077434 28 3.8
TC16636 homologue to UP|Q9LGB7 (Q9LGB7) ESTs C73688(E20155), par... 28 5.0
>TC8565 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, partial (97%)
Length = 1913
Score = 268 bits (684), Expect = 2e-72
Identities = 143/287 (49%), Positives = 184/287 (63%), Gaps = 7/287 (2%)
Frame = +2
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N QAA DM KVL V + E+ + + N + L S++F RHG L
Sbjct: 818 YTALVAKNAKQAASDMSKVLQVELEAEEEKVEKILESENQQFGLFSKQFASRHGMHLLGT 997
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
+ WFLLDI FYSQ LFQ +I+ ++ + + E + +AR Q ++A+CST+PGY+
Sbjct: 998 TTTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYW 1177
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV FID +GR IQ+MGFFFM V FAL PY HWTK +N GF+ +Y L FFFAN
Sbjct: 1178 FTVAFIDYMGRFAIQLMGFFFMTVFMFALALPY-DHWTKKDNRF--GFVAMYALTFFFAN 1348
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGYPK 498
FGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+AS + + GYP
Sbjct: 1349 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYASQSKDPAKTDPGYPT 1528
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEE 545
GIG++ SLI+LG + +GM T F E+ G+SLEE E E
Sbjct: 1529 GIGVRNSLIMLGVINFLGMVFT-FLVPESKGKSLEELSGETEEDGVE 1666
Score = 31.6 bits (70), Expect = 0.35
Identities = 53/181 (29%), Positives = 71/181 (38%), Gaps = 4/181 (2%)
Frame = +1
Query: 33 QED*RFYQPLTQQKHNTTISRPSS*QEWDFSLTLMTFFPSHSSQRC*EEYTTPT-TKKSV 91
+E * L +HN T S EWDFSL M F S YTT T + +
Sbjct: 100 REQ*ECLMHLMWPRHNGTTSLQL*LLEWDFSLMHMISFAFPLSPSYWGGYTTLT*PNQGL 279
Query: 92 KSTRSWSQPL---SPLHS*EPPSVSSYSDASET*KAGATSTVSPYY*C*QAL*LLASPSV 148
+ +PL S P S SS + G + +S +*C A L SP V
Sbjct: 280 VFFQLMCKPL*LVSRFAVP*PASFSSVGSVTRWEGKGFMA*LS*SW*C--APLLQGSPLV 453
Query: 149 QQRELVFYYLLVSLDSS*GLELVVIILYLLPLCLSLLIKGQEVLL*LRFFLCKGLVYWLV 208
++ + VS S LV L+ C + K +E LR LCKG W+V
Sbjct: 454 IAQK-ASWPRFVSSGSGLVSGLVATTLFRPQSCPNTRTKRREARSLLRCLLCKGSESWVV 630
Query: 209 Q 209
+
Sbjct: 631 E 633
>BP067261
Length = 460
Score = 132 bits (331), Expect = 2e-31
Identities = 65/128 (50%), Positives = 85/128 (65%), Gaps = 2/128 (1%)
Frame = +1
Query: 321 LFACSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTI 378
L S+ WFLLDI FYSQ LFQ +I+ ++ + + E + +AR Q ++A+CST+
Sbjct: 85 LLGTSSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTV 264
Query: 379 PGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAF 438
PGY+FTV ID +GR IQ+MGFFFM V FAL P Y HWT+ +N GF+ +Y L F
Sbjct: 265 PGYWFTVALIDYMGRFAIQLMGFFFMTVFMFALAIP-YDHWTEKDN--RIGFVAMYSLTF 435
Query: 439 FFANFGPN 446
FFANFGPN
Sbjct: 436 FFANFGPN 459
>TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, partial
(59%)
Length = 991
Score = 85.9 bits (211), Expect(2) = 6e-24
Identities = 49/116 (42%), Positives = 67/116 (57%), Gaps = 2/116 (1%)
Frame = +3
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACS 325
YTALV +N QAAKDM KVL V +I E A + L S+EF+ RHG L +
Sbjct: 642 YTALVAKNTEQAAKDMSKVLQV---EIQAEPKGDQAQANTFALFSKEFMRRHGLHLLGTA 812
Query: 326 ANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIP 379
+ WFLLDI FYSQ LFQ +I+ ++ + +E + +AR Q ++A+CS P
Sbjct: 813 STWFLLDIAFYSQNLFQKDIFSAIGWIPPAKTMNALEEVYRIARAQTLIALCSYCP 980
Score = 42.0 bits (97), Expect(2) = 6e-24
Identities = 38/100 (38%), Positives = 49/100 (49%), Gaps = 5/100 (5%)
Frame = +1
Query: 168 LELVVIILYLLPLCLSLLIKGQEVLL*LRFFLCKGLVYWLVQLLLWWFAWY-----FAAV 222
LELV +LLPLCLS+ I+ LR LC+ L + V L F + +
Sbjct: 334 LELVETTPFLLPLCLSIRIRRLAAPSSLRSLLCRVLEFLEVVYLQS*FLLHSRPSLMLRL 513
Query: 223 RSRQRLLMCRRRLTLHGG*Y**LVQFQLH*LIIGV**CQK 262
RL+ +LT+ GG* * L QLH*L G * C+K
Sbjct: 514 MRLIRLVQLFLKLTIFGG*L*WLEHSQLH*LTTGG*RCRK 633
>BP042583
Length = 459
Score = 74.3 bits (181), Expect = 5e-14
Identities = 41/85 (48%), Positives = 53/85 (62%)
Frame = -3
Query: 454 AELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVC 513
AE+FP R RSTCHGIS A GK GA++GS GFL+A + IG++ LI+LG
Sbjct: 457 AEIFPPRLRSTCHGISAAAGKAGAMVGSFGFLYAQN---------AIGLRNVLILLGVAN 305
Query: 514 IVGMFVTYFFTKETMGRSLEENEDE 538
++G+F T F E G+SLEE E
Sbjct: 304 LLGLFFT-FLVPEPNGKSLEEISGE 233
>TC9159 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose transporter
HXT4 (Low-affinity glucose transporter LGT1), partial
(5%)
Length = 728
Score = 55.1 bits (131), Expect = 3e-08
Identities = 33/110 (30%), Positives = 53/110 (48%), Gaps = 2/110 (1%)
Frame = +3
Query: 429 GFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW 486
GF+ + GLA + F P T ++V +E++P R+R C GI+ V ++ S FL
Sbjct: 111 GFLALGGLALYIIFFSPGMGTVPWVVNSEIYPLRYRGICGGIASTTVWVSNLVVSQSFL- 287
Query: 487 ASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+ IG + ++ G + I+G+F F ET G +EE E
Sbjct: 288 --------SLTQTIGTAWTFMMFGIIAIIGIFFVLIFVPETKGVPMEEVE 413
>TC10135
Length = 1501
Score = 50.1 bits (118), Expect = 9e-07
Identities = 30/110 (27%), Positives = 51/110 (46%), Gaps = 2/110 (1%)
Frame = +2
Query: 429 GFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW 486
G+ + GLA + F P T +++ +E++P R+R C GI+ + +I S FL
Sbjct: 884 GWAALVGLALYIITFSPGMGTVPWVINSEIYPFRYRGVCGGIASTTVWISNLIVSQSFL- 1060
Query: 487 ASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+ IG + ++ G + +V +F F ET G +EE E
Sbjct: 1061--------SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 1186
>AV769249
Length = 485
Score = 47.4 bits (111), Expect = 6e-06
Identities = 25/52 (48%), Positives = 31/52 (59%)
Frame = -1
Query: 493 EEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAE 544
+ GYP GIG+K SL++LG V I+G F T F E G+SLEE E E
Sbjct: 473 DAGYPAGIGVKNSLLLLGVVNILGFFCT-FLVPEAKGKSLEEMSGENEEEGE 321
>TC13756 similar to UP|Q9ARI9 (Q9ARI9) Phosphate transporter 1, partial
(11%)
Length = 446
Score = 47.0 bits (110), Expect = 8e-06
Identities = 25/52 (48%), Positives = 31/52 (59%)
Frame = +3
Query: 493 EEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAE 544
+ GYP GIG+K SL++LG V I+G T F E G+SLEE EQ E
Sbjct: 33 DAGYPAGIGVKNSLLLLGVVNILGFLFT-FLVPEAKGKSLEEMSGEQEEEVE 185
>TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter like protein,
partial (93%)
Length = 1865
Score = 45.1 bits (105), Expect = 3e-05
Identities = 42/161 (26%), Positives = 69/161 (42%), Gaps = 8/161 (4%)
Frame = +2
Query: 384 TVYFIDRVGR---VKIQMMGFFF-MAVSFFALGFPYYSH----WTKGENHDNKGFMVIYG 435
+ + +DRVGR ++I + G F + + F+L YS W + + Y
Sbjct: 1058 STFLLDRVGRRRLLQISVAGMIFGLTILGFSLTMVEYSSEKLVWAL-----SLSIVATYT 1222
Query: 436 LAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEG 495
FF N G T++ +E+FP R R+ + I AV + S+ F+
Sbjct: 1223 YVAFF-NVGLAPVTWVYSSEIFPLRLRAQGNSIGVAVNRGMNAAISMSFI---------S 1372
Query: 496 YPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
K I + + + G+ +V YF ET G++LEE E
Sbjct: 1373 IYKAITIGGAFFLFAGMSVVAWVFFYFCLPETKGKALEEME 1495
>TC10748 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(21%)
Length = 550
Score = 43.1 bits (100), Expect = 1e-04
Identities = 27/95 (28%), Positives = 45/95 (46%)
Frame = +2
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIG 501
+ G T++ +E+FP R R+ + V +V + + S+ FL S KGI
Sbjct: 29 SIGAGPITWVYSSEIFPLRLRAQGCAMGVVVNRVTSGVISMTFLSLS---------KGIT 181
Query: 502 MKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+ + + GG+ I G Y ET G++LE+ E
Sbjct: 182 IGGAFFLFGGIAICGWIFFYTMLPETRGKTLEDME 286
>BI418774
Length = 554
Score = 32.7 bits (73), Expect = 0.16
Identities = 24/106 (22%), Positives = 46/106 (42%), Gaps = 2/106 (1%)
Frame = +3
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGF-PYYSHWTKGENHDNKGFMVIYGLAFF- 439
F + + +DRVGR + ++ + V+ L F + +K E F ++ F
Sbjct: 231 FISAFLLDRVGRRILLLISSGGVVVTLLGLSFCSFIMQQSKEEPLWGISFTIVAIYIFVA 410
Query: 440 FANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFL 485
F G T++ +E+FP R R+ G+ V ++ + F+
Sbjct: 411 FVAIGIGPVTWVYSSEIFPLRLRAQGLGVCVLVNRLANVAVLTSFI 548
>TC8913 similar to UP|Q7XYW2 (Q7XYW2) Seed specific protein Bn15D17A,
partial (53%)
Length = 615
Score = 30.4 bits (67), Expect = 0.77
Identities = 20/52 (38%), Positives = 24/52 (45%)
Frame = +2
Query: 436 LAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWA 487
LA F+ G P L PA FRS + G + V I GSVG LW+
Sbjct: 227 LAKFYGRAGLMNLVNSGPEHLRPAIFRSLLYEACGRI--VNPIYGSVGLLWS 376
>BP067804
Length = 412
Score = 30.4 bits (67), Expect = 0.77
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = -2
Query: 505 SLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+ +L G+ +V YFF ET GRSLE+ E
Sbjct: 405 TFFMLAGINLVAWSFYYFFLPETKGRSLEDME 310
>TC9057
Length = 603
Score = 29.3 bits (64), Expect = 1.7
Identities = 16/51 (31%), Positives = 24/51 (46%)
Frame = +2
Query: 407 SFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGPNTTTFIVPAELF 457
SFF+ FP W++ +H +V + FF FG +F P+ LF
Sbjct: 218 SFFSFLFPLKQRWSRSGSHTRLS-IVHFQFVFFLILFGFTFFSFGFPSSLF 367
>TC17041 homologue to GB|AAL25568.1|16648753|AY058152 AT5g16150/T21H19_70
{Arabidopsis thaliana;} , partial (18%)
Length = 632
Score = 29.3 bits (64), Expect = 1.7
Identities = 25/105 (23%), Positives = 45/105 (42%)
Frame = +2
Query: 432 VIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKE 491
V+Y L+F + G ++ E+F +R R+ +S + + + + FL +K
Sbjct: 5 VLYVLSF---SLGAGPVPALLLPEIFASRIRAKAVSLSLGMHWISNFVIGLYFLSVVNK- 172
Query: 492 KEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
G+ + + VC++ + ET GRSLEE E
Sbjct: 173 --------FGISSVYLGFSAVCVLAVLYIAGNVVETKGRSLEEIE 283
>TC7806 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1, complete
Length = 1413
Score = 28.9 bits (63), Expect = 2.2
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = +1
Query: 432 VIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGIS 469
VIYG F N+ +F+ P+ LF F +CH IS
Sbjct: 1219 VIYGCDCVFCQIKNNSESFLDPSFLFSVFFFLSCH*IS 1332
>AU251730
Length = 336
Score = 28.5 bits (62), Expect = 2.9
Identities = 20/55 (36%), Positives = 24/55 (43%)
Frame = +2
Query: 41 PLTQQKHNTTISRPSS*QEWDFSLTLMTFFPSHSSQRC*EEYTTPTTKKSVKSTR 95
PL +HN S PSS WD S T + C EY TT+K V +R
Sbjct: 101 PLILPRHNGITSLPSSSLVWDSSQMHTTCSAYQTLPSCLAEY---TTQKKVHQSR 256
>BP047517
Length = 495
Score = 28.5 bits (62), Expect = 2.9
Identities = 15/51 (29%), Positives = 24/51 (46%)
Frame = -1
Query: 432 VIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSV 482
++ G+ F G FI +ELFP R+ G + ++GAI+ V
Sbjct: 429 MVCGVLGIFGMAGTYNLLFIYTSELFPTVVRNAALGCATQASQMGAILAPV 277
>BP077434
Length = 375
Score = 28.1 bits (61), Expect = 3.8
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = -3
Query: 450 FIVPAELFPARFRSTCHGISGAVGKVGAIIGSVG 483
F+V + ++ HG+S G + AI+G+VG
Sbjct: 370 FLVNLQEIAPQYAGVLHGMSNTAGTLAAIVGTVG 269
>TC16636 homologue to UP|Q9LGB7 (Q9LGB7) ESTs C73688(E20155), partial (12%)
Length = 592
Score = 27.7 bits (60), Expect = 5.0
Identities = 22/75 (29%), Positives = 32/75 (42%), Gaps = 3/75 (4%)
Frame = -3
Query: 43 TQQKHNTTISRPSS*QEWDFSLTLMTFFPSHSSQRC*EEYTTPTTKKSVKSTRS---WSQ 99
T Q TT P + + T FP+ +S TPTT +S K T++ S
Sbjct: 533 THQNDQTTHQTPRN----QYCFHTQTRFPTRTSHNSASGPQTPTTHRSRKLTQAEHGTSP 366
Query: 100 PLSPLHS*EPPSVSS 114
P +P + PPS +
Sbjct: 365 PPAPPPTTSPPSAEA 321
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.342 0.150 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,276,648
Number of Sequences: 28460
Number of extensions: 159679
Number of successful extensions: 1478
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1441
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1465
length of query: 548
length of database: 4,897,600
effective HSP length: 95
effective length of query: 453
effective length of database: 2,193,900
effective search space: 993836700
effective search space used: 993836700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149576.4


