

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.2 + phase: 0
(332 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
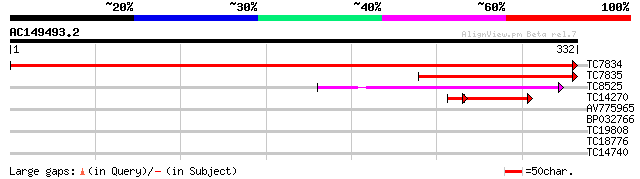
TC7834 homologue to UP|AAS18241 (AAS18241) Cytosolic malate dehy... 632 0.0
TC7835 homologue to UP|Q8GZN2 (Q8GZN2) Malate dehydrogenase , p... 176 4e-45
TC8525 similar to UP|MDHP_PEA (P21528) Malate dehydrogenase [NAD... 97 5e-21
TC14270 similar to UP|Q9FJU0 (Q9FJU0) Cytosolic malate dehydroge... 58 1e-10
AV775965 37 0.006
BP032766 28 2.1
TC19808 homologue to UP|Q9LH76 (Q9LH76) Genomic DNA, chromosome ... 27 6.2
TC18776 similar to UP|Q9FJJ1 (Q9FJJ1) Genomic DNA, chromosome 5,... 26 8.1
TC14740 similar to UP|Q8W584 (Q8W584) AT4g17230/dl4650c, partial... 26 8.1
>TC7834 homologue to UP|AAS18241 (AAS18241) Cytosolic malate dehydrogenase
, complete
Length = 1572
Score = 632 bits (1631), Expect = 0.0
Identities = 319/332 (96%), Positives = 326/332 (98%)
Frame = +3
Query: 1 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDI PAAESLNGVKMELVD
Sbjct: 96 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIPPAAESLNGVKMELVD 275
Query: 61 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVM+KNVSIYKSQASALEKH
Sbjct: 276 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMTKNVSIYKSQASALEKH 455
Query: 121 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 180
AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERL+V VSDVK
Sbjct: 456 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLSVPVSDVK 635
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
NVIIWGNHSSTQYPDVNHATV TPAGEKPVRELVSDD WLNGEFI+TVQQRGAAIIKARK
Sbjct: 636 NVIIWGNHSSTQYPDVNHATVKTPAGEKPVRELVSDDTWLNGEFITTVQQRGAAIIKARK 815
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 300
LSSALSAAS ACDHIRDWVLGTP+GT+VSMGVYSDGSYNVP+GLIYSFPVTCANGEWKIV
Sbjct: 816 LSSALSAASGACDHIRDWVLGTPEGTWVSMGVYSDGSYNVPAGLIYSFPVTCANGEWKIV 995
Query: 301 QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
QGLSIDEFSRKKLDLTAEEL+EEK LAYSCLS
Sbjct: 996 QGLSIDEFSRKKLDLTAEELTEEKALAYSCLS 1091
>TC7835 homologue to UP|Q8GZN2 (Q8GZN2) Malate dehydrogenase , partial
(28%)
Length = 566
Score = 176 bits (447), Expect = 4e-45
Identities = 86/93 (92%), Positives = 90/93 (96%)
Frame = +2
Query: 240 KLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKI 299
KLSSALSAASAACDHIRDWVLGTP+GT+VSMGVYSDGSYNVP+GLIYSFP+T NGEWKI
Sbjct: 2 KLSSALSAASAACDHIRDWVLGTPEGTWVSMGVYSDGSYNVPAGLIYSFPITSQNGEWKI 181
Query: 300 VQGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
VQGLSIDEFSRKKLDLTAEELSEEK LAYSCLS
Sbjct: 182 VQGLSIDEFSRKKLDLTAEELSEEKALAYSCLS 280
>TC8525 similar to UP|MDHP_PEA (P21528) Malate dehydrogenase [NADP],
chloroplast precursor (NADP-MDH) , partial (38%)
Length = 963
Score = 96.7 bits (239), Expect = 5e-21
Identities = 54/146 (36%), Positives = 87/146 (58%), Gaps = 2/146 (1%)
Frame = +1
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
N+ IWGNHS+TQ PD +A ++ PV ++V D WL EF VQ+RG +I+
Sbjct: 4 NMTIWGNHSTTQVPDFLNARIDGV----PVTDVVKDLKWLKEEFTEKVQKRGGVLIQKWG 171
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGS-YNVPSGLIYSFPV-TCANGEWK 298
SSA S A + D +R V TP+G + S GVYS G+ Y + +++S P + +G+++
Sbjct: 172 RSSAASTAVSIVDAMRSLVTPTPEGDWFSSGVYSSGNPYGIAEDIVFSMPCRSKGDGDYE 351
Query: 299 IVQGLSIDEFSRKKLDLTAEELSEEK 324
+V+ + ID++ R+++ + EL EK
Sbjct: 352 LVKDVLIDDYLRQRIAKSEAELLAEK 429
>TC14270 similar to UP|Q9FJU0 (Q9FJU0) Cytosolic malate dehydrogenase ,
partial (12%)
Length = 826
Score = 57.8 bits (138), Expect(2) = 1e-10
Identities = 27/40 (67%), Positives = 28/40 (69%)
Frame = +1
Query: 267 FVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQGLSID 306
FVSMGV SDGSY + GLIY FPVTC G W IV G S D
Sbjct: 235 FVSMGVCSDGSYGIQHGLIYYFPVTCEKGVWNIVPGASND 354
Score = 24.3 bits (51), Expect(2) = 1e-10
Identities = 8/12 (66%), Positives = 11/12 (91%)
Frame = +3
Query: 257 DWVLGTPQGTFV 268
+WVLG P+GT+V
Sbjct: 201 EWVLGAPKGTWV 236
>AV775965
Length = 375
Score = 36.6 bits (83), Expect = 0.006
Identities = 18/57 (31%), Positives = 36/57 (62%), Gaps = 2/57 (3%)
Frame = -3
Query: 259 VLGTPQGTFVSMGVYSDGS-YNVPSGLIYSFPV-TCANGEWKIVQGLSIDEFSRKKL 313
V TP+ + S GVYS G+ Y + +++S P + +G++++VQ + ID++ R ++
Sbjct: 370 VTPTPERDWFSSGVYSRGNPYGIAEDIVFSMPCRSKGDGDYELVQDVLIDDYLRPRI 200
>BP032766
Length = 535
Score = 28.1 bits (61), Expect = 2.1
Identities = 24/89 (26%), Positives = 41/89 (45%), Gaps = 7/89 (7%)
Frame = +1
Query: 1 MAKDPVRVLVTGAAGQIG-YALVPMIARG------VMLGPDQPVILHMLDIAPAAESLNG 53
M+ + V VTGAAG IG + ++ +I RG V + + H+L++ A L+
Sbjct: 55 MSSESETVCVTGAAGFIGSWLVMRLIERGYTVRATVRDPANMKKVKHLLELPDAKTKLSL 234
Query: 54 VKMELVDAAFPLLKGVVATTDVVEACTGV 82
K +L + + + + CTGV
Sbjct: 235 WKADLAEEG--------SFDEAIRGCTGV 297
>TC19808 homologue to UP|Q9LH76 (Q9LH76) Genomic DNA, chromosome 3, BAC
clone: T21E2 (AT3g14790/T21E2_4), partial (19%)
Length = 515
Score = 26.6 bits (57), Expect = 6.2
Identities = 19/72 (26%), Positives = 32/72 (44%)
Frame = +2
Query: 5 PVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFP 64
P +L+TGAAG I + + R PD ++ +LD +L + F
Sbjct: 158 PKNILITGAAGFIASHVANRLVRTY---PDYKIV--VLDKLDYCSNLKNLLPSKSSPNFK 322
Query: 65 LLKGVVATTDVV 76
+KG + + D+V
Sbjct: 323 FVKGDIGSADLV 358
>TC18776 similar to UP|Q9FJJ1 (Q9FJJ1) Genomic DNA, chromosome 5, TAC
clone:K19B1, partial (8%)
Length = 555
Score = 26.2 bits (56), Expect = 8.1
Identities = 16/42 (38%), Positives = 20/42 (47%)
Frame = +2
Query: 242 SSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSG 283
+SAL+ S+AC +RDW G VS G PSG
Sbjct: 173 ASALTIRSSACPGLRDW--GNVASHLVSWSGRCFGRIGEPSG 292
>TC14740 similar to UP|Q8W584 (Q8W584) AT4g17230/dl4650c, partial (57%)
Length = 1406
Score = 26.2 bits (56), Expect = 8.1
Identities = 10/35 (28%), Positives = 22/35 (62%)
Frame = -2
Query: 291 TCANGEWKIVQGLSIDEFSRKKLDLTAEELSEEKN 325
TC++ W I +G +++ +++ + LTA+ +E N
Sbjct: 1348 TCSSLHWNI*RGKELNKKTKQNVKLTAKPTKQEYN 1244
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,942,474
Number of Sequences: 28460
Number of extensions: 61453
Number of successful extensions: 269
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 268
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 268
length of query: 332
length of database: 4,897,600
effective HSP length: 91
effective length of query: 241
effective length of database: 2,307,740
effective search space: 556165340
effective search space used: 556165340
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149493.2


