

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.1 + phase: 0
(254 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
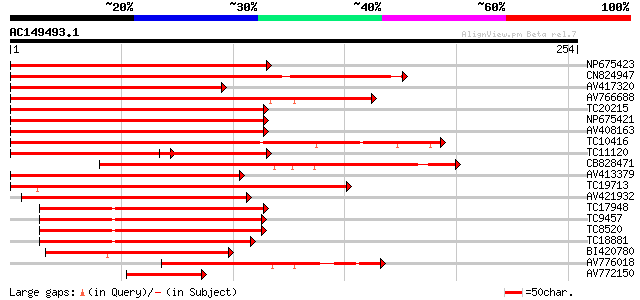
Score E
Sequences producing significant alignments: (bits) Value
NP675423 transcription factor MYB101 [Lotus japonicus] 193 2e-50
CN824947 189 5e-49
AV417320 185 6e-48
AV766688 183 2e-47
TC20215 similar to AAS10104 (AAS10104) MYB transcription factor,... 183 2e-47
NP675421 transcription factor MYB103 [Lotus japonicus] 182 4e-47
AV408163 182 4e-47
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 182 5e-47
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 122 9e-46
CB828471 161 1e-40
AV413379 160 3e-40
TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%) 147 1e-36
AV421932 145 7e-36
TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose... 108 9e-25
TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 103 4e-23
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 102 6e-23
TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose... 100 3e-22
BI420780 75 8e-15
AV776018 70 5e-13
AV772150 59 6e-10
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 193 bits (491), Expect = 2e-50
Identities = 87/117 (74%), Positives = 97/117 (82%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R PCC++ GLKKGPWT EED IL YI KHGH +WRALPK AGL RCGKSCRLRW NY
Sbjct: 1 MGRTPCCDEIGLKKGPWTPEEDAILVDYIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNY 180
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLL 117
LR DIKRG F+ EEEQ+II +H +LGN+WSAIA LPGRTDNEIKN W+THLKK+LL
Sbjct: 181 LRPDIKRGKFTEEEEQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKKLL 351
>CN824947
Length = 738
Score = 189 bits (479), Expect = 5e-49
Identities = 92/178 (51%), Positives = 118/178 (65%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M RAPCCEK GLK+G WT EEDE+LT YI G +WR+LPK+AGLLRCGKSCRLRWINY
Sbjct: 151 MGRAPCCEKVGLKRGRWTAEEDELLTKYIQASGEGSWRSLPKNAGLLRCGKSCRLRWINY 330
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR D+KRGN S+EEE +I+K+H GNRWS IA+ +PGRTDNEIKN W++HL ++L+ T
Sbjct: 331 LRGDLKRGNISDEEESLIVKLHASFGNRWSLIASHMPGRTDNEIKNYWNSHLSRKLVYTF 510
Query: 121 NNQPHSNTKKRVSKQKIKISDSNANSNSMETTSSNCTFSSDFSSQGKNLDNSIICEDP 178
S TK V + + +M+ + ++D +Q L+ EDP
Sbjct: 511 RG---STTKSIVDTPPKRRRCGRTSRWAMKKNKTYSLINNDLQNQNNELEK----EDP 663
>AV417320
Length = 369
Score = 185 bits (470), Expect = 6e-48
Identities = 85/97 (87%), Positives = 89/97 (91%)
Frame = +3
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
MVRAPCCEK GLKKGPWT EED+IL SYI KHGH NWRALPK AGLLRCGKSCRLRWINY
Sbjct: 78 MVRAPCCEKIGLKKGPWTSEEDQILISYIQKHGHGNWRALPKHAGLLRCGKSCRLRWINY 257
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLP 97
LR DIKRGNF+ EEE++IIKMHELLGNRWSAIAAKLP
Sbjct: 258 LRPDIKRGNFTAEEEELIIKMHELLGNRWSAIAAKLP 368
>AV766688
Length = 608
Score = 183 bits (465), Expect = 2e-47
Identities = 92/176 (52%), Positives = 117/176 (66%), Gaps = 12/176 (6%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R P C+K GL+KG WT EED L +Y+ ++G NWR LPK AGL RCGKSCRLRW+NY
Sbjct: 77 MARTPSCDKSGLRKGTWTAEEDRKLIAYVTRYGCWNWRQLPKFAGLARCGKSCRLRWLNY 256
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKR----- 115
LR DIKRG+F+ EEE+III+MH+ LGNRWS IAA+LPGRTDNE+KN WHT LK+R
Sbjct: 257 LRPDIKRGHFTQEEEEIIIRMHKNLGNRWSIIAAELPGRTDNEVKNHWHTSLKRRVQQNE 436
Query: 116 ---LLNTKNNQPHS----NTKKRVSKQKIKISDSNANSNSMETTSSNCTFSSDFSS 164
+ N+KN P S + + +ISDS + ++S SSD ++
Sbjct: 437 EAKVSNSKNEGPTSGNGIGSLQVTPPATSQISDSTGALSPFSSSSEFSCISSDLNT 604
>TC20215 similar to AAS10104 (AAS10104) MYB transcription factor, partial
(35%)
Length = 464
Score = 183 bits (465), Expect = 2e-47
Identities = 78/116 (67%), Positives = 100/116 (85%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M RAPCCEK GLK+G W+ EED+ILT +I ++G +W++LPK+AGLLRCGKSCRLRWINY
Sbjct: 29 MGRAPCCEKVGLKRGRWSEEEDKILTDFIKQNGEGSWKSLPKNAGLLRCGKSCRLRWINY 208
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
LR D+KRGN ++EEE+II+K+H LGNRWS IA LPGRTDNEIKN W++HL++++
Sbjct: 209 LREDVKRGNITSEEEEIIVKLHAALGNRWSVIAGHLPGRTDNEIKNYWNSHLRRKI 376
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 182 bits (463), Expect = 4e-47
Identities = 79/116 (68%), Positives = 97/116 (83%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M+R P ++ G+KKGPW+ EED++L YI KHGH +WRALP+ AGL RCGKSCRLRW NY
Sbjct: 1 MMRTPSIDESGMKKGPWSQEEDKVLVDYIQKHGHGSWRALPQLAGLNRCGKSCRLRWTNY 180
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
LR DIKRG FS+EEEQ+II +H LGN+W+ IA+ LPGRTDNEIKN+W+THLKK+L
Sbjct: 181 LRPDIKRGKFSDEEEQLIINLHASLGNKWATIASHLPGRTDNEIKNLWNTHLKKKL 348
>AV408163
Length = 425
Score = 182 bits (463), Expect = 4e-47
Identities = 79/116 (68%), Positives = 96/116 (82%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R+PCCEK + KG WT EED+ L SYI HG WR+LPK AGLLRCGKSCRLRWINY
Sbjct: 77 MGRSPCCEKAHINKGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 256
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
LR D+KRGNF+ EE+++IIK+H LLGN+WS IA +LPGRTDNEIKN W+TH++++L
Sbjct: 257 LRPDLKRGNFTEEEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKL 424
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 182 bits (462), Expect = 5e-47
Identities = 99/214 (46%), Positives = 134/214 (62%), Gaps = 19/214 (8%)
Frame = +2
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R P C+K G++KG WT EED+ L +Y+ ++G NWR LPK AGL RCGKSCRLRW+NY
Sbjct: 92 MARTPSCDKSGMRKGTWTAEEDKKLIAYVTRYGCWNWRQLPKFAGLARCGKSCRLRWLNY 271
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLNTK 120
LR +IKRGNF+ EEE+III+MH+ LGNRWS IAA+LPGRTD+E+KN WHT L KR ++ +
Sbjct: 272 LRPNIKRGNFTQEEEEIIIRMHKKLGNRWSTIAAELPGRTDSEVKNHWHTTL-KRTVHEQ 448
Query: 121 NNQPHSNTKKRVSKQK-------------IKISDSNANSNSMETTSSNCTFSSDFSSQGK 167
N + TK SK + + + + S S+ S T SS+FS
Sbjct: 449 NTVTNEETKVSNSKNERDPVPKSGIASLLVTPPPATSISGSIGALSPFST-SSEFSCTTS 625
Query: 168 NLDNS-----IICEDPEDSFVTMPQ-IDESFWSE 195
+L S ++ ED D + ++E+FWS+
Sbjct: 626 DLTTSSSSEMLVFEDDFDFLDDFTENVNENFWSD 727
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 122 bits (305), Expect(2) = 9e-46
Identities = 54/74 (72%), Positives = 60/74 (80%)
Frame = +1
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R+PCC++ GLKKGPWT EED+ L +I KHGH +WRALPK AGL RCGKSCRLRW NY
Sbjct: 67 MGRSPCCDENGLKKGPWTPEEDQKLVEHIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNY 246
Query: 61 LRTDIKRGNFSNEE 74
LR DIKRG FS EE
Sbjct: 247 LRPDIKRGKFSPEE 288
Score = 77.8 bits (190), Expect(2) = 9e-46
Identities = 32/50 (64%), Positives = 41/50 (82%)
Frame = +2
Query: 68 GNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLL 117
GNF ++EQ I+ +H +LGN+WS IA LPGRTDNEIKN W+THLKK+L+
Sbjct: 269 GNFLQKKEQTILHLHSILGNKWSTIATHLPGRTDNEIKNFWNTHLKKKLI 418
>CB828471
Length = 525
Score = 161 bits (407), Expect = 1e-40
Identities = 90/177 (50%), Positives = 112/177 (62%), Gaps = 15/177 (8%)
Frame = +1
Query: 41 PKDAGLLRCGKSCRLRWINYLRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRT 100
PK AGLLRCGKSCRLRW NYLR DIKRGNFS EEE II +HELLGNRWSAIAA+LPGRT
Sbjct: 1 PKQAGLLRCGKSCRLRWANYLRPDIKRGNFSREEEDEIINLHELLGNRWSAIAARLPGRT 180
Query: 101 DNEIKNMWHTHLKKRLL-NTKNNQPH-----SNTKKRVSKQ---------KIKISDSNAN 145
DNEIKN+WHTHLKKRL TK Q + S + K K+ + S SN++
Sbjct: 181 DNEIKNVWHTHLKKRLQPQTKQKQTNLDLDASKSNKDAKKEHQEDEGPRSPQQCSSSNSD 360
Query: 146 SNSMETTSSNCTFSSDFSSQGKNLDNSIICEDPEDSFVTMPQIDESFWSETITDDET 202
+ S T SS+ S++ N +++ + E+S +DE FWSE ++ D +
Sbjct: 361 NMSSLTNSSSIAPSTNHDDMSINDVHNVNFDSLENSLA----LDEDFWSEVLSSDNS 519
>AV413379
Length = 416
Score = 160 bits (404), Expect = 3e-40
Identities = 71/105 (67%), Positives = 84/105 (79%)
Frame = +3
Query: 1 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINY 60
M R+PCCEK KG WT EED+ L +YI HG WR+LPK AGLLRCGKSCRLRWINY
Sbjct: 102 MGRSPCCEKAHTNKGAWTKEEDDRLIAYIRAHGEGCWRSLPKSAGLLRCGKSCRLRWINY 281
Query: 61 LRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIK 105
LR D+KRGNF+ +E+++IIK+H LLGN+WS IA +L GRTDNEIK
Sbjct: 282 LRPDLKRGNFTPQEDELIIKLHSLLGNKWSLIAGRLAGRTDNEIK 416
>TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%)
Length = 532
Score = 147 bits (372), Expect = 1e-36
Identities = 70/155 (45%), Positives = 96/155 (61%), Gaps = 2/155 (1%)
Frame = +3
Query: 1 MVRAPCCEKKG--LKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWI 58
M + PC ++KGPWT+EED IL +YI HG W L K AGL R GKSCRLRW+
Sbjct: 66 MDKKPCNSSHDPEVRKGPWTIEEDLILINYIANHGEGVWNTLAKSAGLKRTGKSCRLRWL 245
Query: 59 NYLRTDIKRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRLLN 118
NYLR D++RGN + EE+ +I+++H GNRWS IA LPGRTDNEIKN W T ++K +
Sbjct: 246 NYLRPDVRRGNITPEEQLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKHIKQ 425
Query: 119 TKNNQPHSNTKKRVSKQKIKISDSNANSNSMETTS 153
++ Q ++ + + S + + M+T S
Sbjct: 426 AESLQQQQSSSEINDHHQTSTSQMSTMAEPMDTYS 530
>AV421932
Length = 402
Score = 145 bits (366), Expect = 7e-36
Identities = 63/103 (61%), Positives = 80/103 (77%)
Frame = +3
Query: 6 CCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLRTDI 65
CC ++ +K+G W+ EEDE L YI HG+ W +P+ AGL RCGKSCRLRWINYLR DI
Sbjct: 93 CCNQQKVKRGLWSPEEDEKLIRYITTHGYGCWSEVPEKAGLQRCGKSCRLRWINYLRPDI 272
Query: 66 KRGNFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMW 108
+RG F+ EEE++II +H ++GNRW+ IA+ LPGRTDNEIKN W
Sbjct: 273 RRGRFTPEEEKLIISLHGVVGNRWAHIASHLPGRTDNEIKNYW 401
>TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (50%)
Length = 487
Score = 108 bits (270), Expect = 9e-25
Identities = 51/103 (49%), Positives = 67/103 (64%)
Frame = +2
Query: 14 KGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLRTDIKRGNFSNE 73
KGPW+ EED ILT + +HG NW + + R GKSCRLRW N L ++ FS +
Sbjct: 170 KGPWSAEEDRILTRLVERHGARNWSLISRYIKG-RSGKSCRLRWCNQLSPTVEHRPFSVQ 346
Query: 74 EEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
E++ II H GNRW+AIA LPGRTDN +KN W++ LK+R+
Sbjct: 347 EDETIIAAHARFGNRWAAIARLLPGRTDNAVKNHWNSTLKRRV 475
>TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (43%)
Length = 860
Score = 103 bits (256), Expect = 4e-23
Identities = 49/102 (48%), Positives = 63/102 (61%)
Frame = +3
Query: 14 KGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLRTDIKRGNFSNE 73
KGPW+ EEDE L + KHG NW + K R GKSCRLRW N L ++ F+ E
Sbjct: 78 KGPWSPEEDEALQRLVEKHGPRNWSLISKSIPG-RSGKSCRLRWCNQLSPQVEHRAFTAE 254
Query: 74 EEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKR 115
E+ II+ H GN+W+ IA L GRTDN IKN W++ LK++
Sbjct: 255 EDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK 380
Score = 30.8 bits (68), Expect = 0.24
Identities = 16/56 (28%), Positives = 30/56 (53%), Gaps = 1/56 (1%)
Frame = +3
Query: 62 RTDIKRGNFSNEEEQIIIKMHELLGNR-WSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
+ D +G +S EE++ + ++ E G R WS I+ +PGR+ + W L ++
Sbjct: 63 KMDRIKGPWSPEEDEALQRLVEKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQV 230
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 102 bits (254), Expect = 6e-23
Identities = 48/102 (47%), Positives = 63/102 (61%)
Frame = +1
Query: 14 KGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLRTDIKRGNFSNE 73
KGPW+ EEDE L + +HG NW + K R GKSCRLRW N L ++ F+ E
Sbjct: 184 KGPWSPEEDEALQKLVERHGPRNWSLISKSIPG-RSGKSCRLRWCNQLSPQVEHRAFTPE 360
Query: 74 EEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKR 115
E+ II+ H GN+W+ IA L GRTDN IKN W++ LK++
Sbjct: 361 EDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK 486
Score = 31.6 bits (70), Expect = 0.14
Identities = 16/51 (31%), Positives = 28/51 (54%), Gaps = 1/51 (1%)
Frame = +1
Query: 67 RGNFSNEEEQIIIKMHELLGNR-WSAIAAKLPGRTDNEIKNMWHTHLKKRL 116
+G +S EE++ + K+ E G R WS I+ +PGR+ + W L ++
Sbjct: 184 KGPWSPEEDEALQKLVERHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQV 336
>TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (28%)
Length = 346
Score = 100 bits (248), Expect = 3e-22
Identities = 47/97 (48%), Positives = 61/97 (62%)
Frame = +1
Query: 14 KGPWTLEEDEILTSYINKHGHSNWRALPKDAGLLRCGKSCRLRWINYLRTDIKRGNFSNE 73
KGPW+ EEDE LT + HG NW A+ K R GKSCRLRW N L +++ F+ E
Sbjct: 55 KGPWSPEEDEALTRLVQVHGPKNWTAISKSIPG-RSGKSCRLRWCNQLSPEVEHRPFTPE 231
Query: 74 EEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHT 110
E+ II+ H GN+W+ IA L GRTDN +KN W++
Sbjct: 232 EDSTIIRAHARCGNKWATIARILNGRTDNAVKNHWNS 342
Score = 31.2 bits (69), Expect = 0.18
Identities = 18/64 (28%), Positives = 34/64 (53%), Gaps = 1/64 (1%)
Frame = +1
Query: 50 GKSCRLRWINYLRTDIKRGNFSNEEEQIIIKMHELLGNR-WSAIAAKLPGRTDNEIKNMW 108
G SC L + D +G +S EE++ + ++ ++ G + W+AI+ +PGR+ + W
Sbjct: 7 GTSCNLL-SPLSQMDRVKGPWSPEEDEALTRLVQVHGPKNWTAISKSIPGRSGKSCRLRW 183
Query: 109 HTHL 112
L
Sbjct: 184 CNQL 195
>BI420780
Length = 475
Score = 75.5 bits (184), Expect = 8e-15
Identities = 34/86 (39%), Positives = 55/86 (63%), Gaps = 2/86 (2%)
Frame = +1
Query: 17 WTLEEDEILTSYINKHGHSNWRALPK--DAGLLRCGKSCRLRWINYLRTDIKRGNFSNEE 74
W+ EED +L +Y+ ++G W + + + L R KSC RW NYL+ IK+G+ + EE
Sbjct: 211 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 390
Query: 75 EQIIIKMHELLGNRWSAIAAKLPGRT 100
++++I + GN+W IAA++PGRT
Sbjct: 391 QRLVILLQANYGNKWKKIAAEVPGRT 468
>AV776018
Length = 476
Score = 69.7 bits (169), Expect = 5e-13
Identities = 42/107 (39%), Positives = 65/107 (60%), Gaps = 7/107 (6%)
Frame = +1
Query: 69 NFSNEEEQIIIKMHELLGNRWSAIAAKLPGRTDNEIKNMWHTHLKKRL----LNTKNNQP 124
+F EE+ +II++H +LGNRWS A + PGRTDNEIKN+W++ LKK+L ++ ++P
Sbjct: 10 HFHKEEDNLIIELHAVLGNRWSQDATQWPGRTDNEIKNLWNSCLKKKLRQRGIDPVTHKP 189
Query: 125 HS---NTKKRVSKQKIKISDSNANSNSMETTSSNCTFSSDFSSQGKN 168
S N K+ +K + K+ SN + S +F SD +S +N
Sbjct: 190 LSEVENGKEDKNKSQEKV------SNELNLLKSE-SFKSDAASYEQN 309
>AV772150
Length = 535
Score = 59.3 bits (142), Expect = 6e-10
Identities = 24/36 (66%), Positives = 30/36 (82%)
Frame = +3
Query: 53 CRLRWINYLRTDIKRGNFSNEEEQIIIKMHELLGNR 88
CRLRWINYLR D+K+G + +EE +IIK+H LLGNR
Sbjct: 27 CRLRWINYLRPDLKKGQLAEDEEDLIIKLHALLGNR 134
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.130 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,050,241
Number of Sequences: 28460
Number of extensions: 69378
Number of successful extensions: 319
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 308
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 313
length of query: 254
length of database: 4,897,600
effective HSP length: 88
effective length of query: 166
effective length of database: 2,393,120
effective search space: 397257920
effective search space used: 397257920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC149493.1


