

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
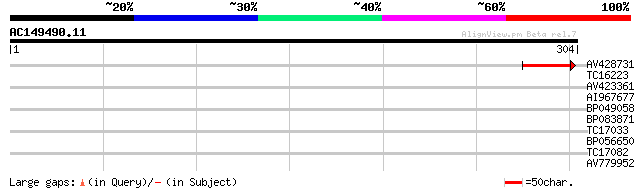
Query= AC149490.11 + phase: 0
(304 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV428731 42 2e-04
TC16223 weakly similar to PIR|T04762|T04762 chitinase homolog T1... 38 0.002
AV423361 33 0.078
AI967677 29 0.86
BP049058 29 0.86
BP083871 28 1.9
TC17033 28 1.9
BP056650 27 4.3
TC17082 27 4.3
AV779952 26 9.5
>AV428731
Length = 151
Score = 41.6 bits (96), Expect = 2e-04
Identities = 18/28 (64%), Positives = 23/28 (81%)
Frame = -2
Query: 276 GIFVYSADDSKADGFRYEKEAQEVLASP 303
GIFV+SADDS +GF YEK++Q +LA P
Sbjct: 150 GIFVWSADDSMGNGFPYEKQSQALLAXP 67
>TC16223 weakly similar to PIR|T04762|T04762 chitinase homolog T16H5.170 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 570
Score = 37.7 bits (86), Expect = 0.002
Identities = 41/151 (27%), Positives = 67/151 (44%), Gaps = 6/151 (3%)
Frame = +2
Query: 11 ILLILTTSLFASTILAANSNLFREYIGAQFKGVKFSDVPINPNVNFHFILSFA-IDYDTS 69
+ L++ L+ S+ AA G + G S INP+ H +FA +D T+
Sbjct: 41 LTLLVVLQLYFSSAEAAIKG------GYWYSGSGLSVSDINPSHFTHLFCAFADLDLQTN 202
Query: 70 SPP-SPTNG-NFNIFWDTQNLSPSEVSFIKNQNPNVKVALSLGGDTVQGDPVNFIPSSID 127
+ SP N F+ F T ++ +NP+VK LS+GG NF S +
Sbjct: 203 TLTISPANAAQFSTFTQT----------LQAKNPSVKTLLSIGGGGGHDLAANF--SKMA 346
Query: 128 SWVSNA---VSSLTSIIKTYNLDGIDVDYEH 155
S S+ + S + + N G+D+D+E+
Sbjct: 347 SQASSRKTFIDSSIQLARKNNFHGLDLDWEY 439
>AV423361
Length = 454
Score = 32.7 bits (73), Expect = 0.078
Identities = 15/49 (30%), Positives = 30/49 (60%), Gaps = 3/49 (6%)
Frame = -2
Query: 62 FAIDYDTS---SPPSPTNGNFNIFWDTQNLSPSEVSFIKNQNPNVKVAL 107
FA+ +D S P+P+N N+N+ + Q+ +P ++ I ++N NV + +
Sbjct: 183 FAVYWDRFGDFSNPNPSNPNWNLSMEPQSTNPMQIQ*ISHKNSNVLITI 37
>AI967677
Length = 407
Score = 29.3 bits (64), Expect = 0.86
Identities = 17/65 (26%), Positives = 33/65 (50%), Gaps = 1/65 (1%)
Frame = +1
Query: 92 EVSFIKNQNPNVKVALSLGGDTV-QGDPVNFIPSSIDSWVSNAVSSLTSIIKTYNLDGID 150
++ +K + ++K LS GG T Q NF ++ + V S +++ DG+D
Sbjct: 190 QLYLLKQRKRSLKTILSFGGWTYSQAGHFNFATNATSR--ATFVKSAIQLLEDNGFDGLD 363
Query: 151 VDYEH 155
+D+E+
Sbjct: 364 IDWEY 378
>BP049058
Length = 534
Score = 29.3 bits (64), Expect = 0.86
Identities = 18/68 (26%), Positives = 35/68 (51%), Gaps = 3/68 (4%)
Frame = +1
Query: 43 VKFSDVPINP--NVNFHFILSFAIDYDTSSPPSPTNGNFNIFWD-TQNLSPSEVSFIKNQ 99
V+FSD P N + +++ + YD + +FN FWD + +PS +S + NQ
Sbjct: 316 VQFSDHPTIYFYNTSLTLVITGKLMYDMALLSKEAITDFNFFWDGNHSYNPSSLSNLGNQ 495
Query: 100 NPNVKVAL 107
+ + +++
Sbjct: 496 SIAMSISI 519
>BP083871
Length = 373
Score = 28.1 bits (61), Expect = 1.9
Identities = 17/47 (36%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = -1
Query: 15 LTTSLFASTILAANSNLFREYIGAQF-KGVKFSDVPINPNVNFHFIL 60
L + +F STIL ++ L+ A KGV D+PIN V+ F++
Sbjct: 154 LYSRIFGSTILESSITLYFFLTPALCCKGVSIMDIPINWKVHLRFLV 14
>TC17033
Length = 950
Score = 28.1 bits (61), Expect = 1.9
Identities = 17/51 (33%), Positives = 25/51 (48%), Gaps = 4/51 (7%)
Frame = +2
Query: 48 VPINPNVNFHFILSFAIDYD----TSSPPSPTNGNFNIFWDTQNLSPSEVS 94
V NP NF LS DYD + P P+ G F++F D ++ ++S
Sbjct: 569 VAFNPQGNFDLSLS---DYDDAPEVTPPMPPSEGRFDVFIDNDAITRLDLS 712
>BP056650
Length = 496
Score = 26.9 bits (58), Expect = 4.3
Identities = 16/52 (30%), Positives = 25/52 (47%), Gaps = 3/52 (5%)
Frame = +3
Query: 99 QNPNVK---VALSLGGDTVQGDPVNFIPSSIDSWVSNAVSSLTSIIKTYNLD 147
+NP ++ +AL D + DP+NF +WV + V + I T LD
Sbjct: 312 ENPRIRNAYIALQAWKDAILSDPLNF----TTNWVGSNVCNYNGIYCTQALD 455
>TC17082
Length = 554
Score = 26.9 bits (58), Expect = 4.3
Identities = 22/70 (31%), Positives = 32/70 (45%), Gaps = 6/70 (8%)
Frame = +1
Query: 98 NQNPNVKVALSLGGDTVQGDPVNFIPSSIDSWVSNAVSSL-----TSIIKTYNLDGIDVD 152
+Q +V +S GD + IPS+ W+ VSSL S+ + + GID D
Sbjct: 31 SQKLSVIKVVSFAGDGFVPTCLGNIPSTGKLWMVTGVSSLPGSNYPSVARVRVISGIDGD 210
Query: 153 YEHFI-GDPN 161
E + GD N
Sbjct: 211 QEPVVLGDDN 240
>AV779952
Length = 557
Score = 25.8 bits (55), Expect = 9.5
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +2
Query: 94 SFIKNQNPNVKVALSLGGDTVQGDPV 119
S I N + N+++ LSLGGD ++G V
Sbjct: 20 SGITN*DCNLRIGLSLGGDFIKGRQV 97
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,275,874
Number of Sequences: 28460
Number of extensions: 70011
Number of successful extensions: 373
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 373
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 373
length of query: 304
length of database: 4,897,600
effective HSP length: 90
effective length of query: 214
effective length of database: 2,336,200
effective search space: 499946800
effective search space used: 499946800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149490.11


