

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149471.21 - phase: 0
(281 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
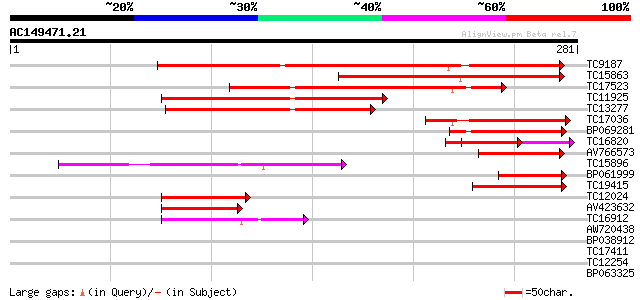
Score E
Sequences producing significant alignments: (bits) Value
TC9187 similar to UP|Q8LAN3 (Q8LAN3) Prolyl 4-hydroxylase alpha ... 190 2e-49
TC15863 135 6e-33
TC17523 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alph... 134 1e-32
TC11925 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alph... 118 1e-27
TC13277 similar to UP|Q9LSI6 (Q9LSI6) Prolyl 4-hydroxylase alpha... 106 5e-24
TC17036 similar to UP|Q8VZD7 (Q8VZD7) AT3g28480/MFJ20_16, partia... 74 4e-14
BP069281 73 5e-14
TC16820 similar to UP|Q8L8T9 (Q8L8T9) Prolyl 4-hydroxylase alpha... 72 1e-13
AV766573 68 2e-12
TC15896 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, ... 59 7e-10
BP061999 57 5e-09
TC19415 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, ... 55 1e-08
TC12024 55 2e-08
AV423632 52 1e-07
TC16912 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, ... 48 2e-06
AW720438 35 0.014
BP038912 29 1.0
TC17411 similar to UP|Q8LDG6 (Q8LDG6) Protein kinase, partial (33%) 27 3.9
TC12254 similar to UP|Q94JV6 (Q94JV6) At1g35190/T32G9_27, partia... 27 3.9
BP063325 27 3.9
>TC9187 similar to UP|Q8LAN3 (Q8LAN3) Prolyl 4-hydroxylase alpha
subunit-like protein, partial (89%)
Length = 1198
Score = 190 bits (483), Expect = 2e-49
Identities = 102/213 (47%), Positives = 135/213 (62%), Gaps = 11/213 (5%)
Frame = +2
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +SW PR + FL+ ECD+L +A LK S V D +G S+VRTSSGMF+
Sbjct: 167 KVKQVSWKPRAFVYEGFLTGLECDHLISLAKSELKRSAVADNLSGDSKLSEVRTSSGMFI 346
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S ++K P++ IE +IS ++ +P ENGE MQVLRYE Q Y PH+DYF+D N+ RGG
Sbjct: 347 S--KKKDPIVAGIEDKISAWTFLPKENGEDMQVLRYEHGQKYDPHYDYFTDKVNIVRGGH 520
Query: 194 RIATMLMYLGDNVEGGETHFPSA-----------GSDECSCGGKLTKGLCVKPVKGNAVL 242
R+AT+L+YL + GGET FP A SD C KG+ VKP +G+A+L
Sbjct: 521 RMATVLLYLTNVTRGGETVFPVAEEPPRRRGLETNSDLSECA---KKGIAVKPRRGDALL 691
Query: 243 FWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
F+S+ DPDS+H GCPV+ GEKWSATKW+
Sbjct: 692 FFSLHTTATPDPDSLHAGCPVIEGEKWSATKWI 790
>TC15863
Length = 607
Score = 135 bits (341), Expect = 6e-33
Identities = 67/118 (56%), Positives = 80/118 (67%), Gaps = 6/118 (5%)
Frame = +2
Query: 164 MQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECS- 222
+QVL YE Q Y PH+DYF D FN K GGQRIAT+LMYL D EGGET FP+A ++ S
Sbjct: 5 LQVLHYEVGQKYEPHYDYFLDEFNTKNGGQRIATVLMYLSDVEEGGETIFPAAKANFSSV 184
Query: 223 -----CGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
KGL VKP +G+A+LFWS+ D DP S+HGGCPV+ G KWS+TKWM
Sbjct: 185 PWYNDLSVCAKKGLSVKPKRGDALLFWSIRPDATLDPSSLHGGCPVIRGNKWSSTKWM 358
>TC17523 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alpha
subunit-like protein, partial (55%)
Length = 448
Score = 134 bits (338), Expect = 1e-32
Identities = 78/145 (53%), Positives = 94/145 (64%), Gaps = 8/145 (5%)
Frame = +1
Query: 110 STVVDANTGKGIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRY 169
STVVD+ TGK S VRTSSG FL K ++ IEK+I+ ++ IP+E+GE +QVL Y
Sbjct: 10 STVVDSETGKSKDSRVRTSSGTFLPRGRDK--IVRNIEKKIADFTFIPVEHGEGLQVLHY 183
Query: 170 EKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGS--------DEC 221
E Q Y PH+DYF D FN K GGQRIAT+LMYL D GGET FPSA +E
Sbjct: 184 EVGQKYEPHYDYFLDEFNTKNGGQRIATVLMYLTDVEXGGETVFPSAKGNFXSVPWWNEL 363
Query: 222 SCGGKLTKGLCVKPVKGNAVLFWSM 246
S GK KGL + P G+A+LFWSM
Sbjct: 364 SDCGK--KGLSINPXXGDALLFWSM 432
>TC11925 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alpha
subunit-like protein, partial (43%)
Length = 613
Score = 118 bits (295), Expect = 1e-27
Identities = 60/112 (53%), Positives = 76/112 (67%)
Frame = +1
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
E++SW PR L HNFL+ EEC+YL +A P ++ STVVD+ TGK S RTSSG FL+
Sbjct: 280 ELISWEPRAFLYHNFLTKEECEYLINIAKPHMRKSTVVDSETGKSKDSRARTSSGTFLAR 459
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFN 187
K ++ IEKRI+ Y+ IPIE+GE +QVL YE Q Y H+DYF D FN
Sbjct: 460 GRDK--IVRNIEKRIADYTFIPIEHGEGLQVLHYEVGQKYDSHYDYFMDEFN 609
>TC13277 similar to UP|Q9LSI6 (Q9LSI6) Prolyl 4-hydroxylase alpha
subunit-like protein, partial (34%)
Length = 582
Score = 106 bits (264), Expect = 5e-24
Identities = 53/104 (50%), Positives = 73/104 (69%)
Frame = +1
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSWSPR L FLS +ECD+L +A +L++S V D +GK IKS+VRTSSGMFL+ +
Sbjct: 277 LSWSPRAFLHTGFLSDKECDHLVNLAKDKLEVSMVADNESGKSIKSEVRTSSGMFLNKAQ 456
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDY 181
+ ++ IE RI+ ++ +PIENGE +Q+L YE Q Y PH+DY
Sbjct: 457 DE--VVADIEARIATWTFLPIENGESIQILHYENGQKYEPHYDY 582
>TC17036 similar to UP|Q8VZD7 (Q8VZD7) AT3g28480/MFJ20_16, partial (21%)
Length = 697
Score = 73.6 bits (179), Expect = 4e-14
Identities = 39/82 (47%), Positives = 51/82 (61%), Gaps = 10/82 (12%)
Frame = +2
Query: 207 EGGETHFPSAGS----------DECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDS 256
+GGET FP A S EC+ KG VKP KG+A+LF+S+ L+ +D +S
Sbjct: 8 KGGETIFPHAESKLSQPKDESWSECA-----HKGYAVKPRKGDALLFFSLHLNATTDSNS 172
Query: 257 VHGGCPVLAGEKWSATKWMRQS 278
+HG CPV+ GEKWSATKW+ S
Sbjct: 173 LHGSCPVIEGEKWSATKWIHVS 238
>BP069281
Length = 468
Score = 73.2 bits (178), Expect = 5e-14
Identities = 34/58 (58%), Positives = 42/58 (71%)
Frame = -3
Query: 219 DECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
+E S GK KGL +KP +G+A+LFWSM D +D S+HGGCPV G+KWS TKWMR
Sbjct: 448 NELSDCGK--KGLSIKPKRGDALLFWSMKPDTTADTSSLHGGCPVTKGDKWSCTKWMR 281
>TC16820 similar to UP|Q8L8T9 (Q8L8T9) Prolyl 4-hydroxylase alpha
subunit-like protein, partial (7%)
Length = 571
Score = 71.6 bits (174), Expect = 1e-13
Identities = 31/38 (81%), Positives = 33/38 (86%)
Frame = +2
Query: 217 GSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDP 254
GS +CSCGGK +GL VKP KGNAVLFWSMGLDGQSDP
Sbjct: 251 GSGQCSCGGKTVEGLSVKPTKGNAVLFWSMGLDGQSDP 364
Score = 58.9 bits (141), Expect = 9e-10
Identities = 30/56 (53%), Positives = 32/56 (56%)
Frame = +3
Query: 225 GKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQSVH 280
GKL+KG +K G P SVHGGC VLAGEKWSATKWMRQ H
Sbjct: 276 GKLSKGYL*NQLKEMQCFSGVWGWMDSQIPFSVHGGCEVLAGEKWSATKWMRQKAH 443
>AV766573
Length = 338
Score = 67.8 bits (164), Expect = 2e-12
Identities = 27/43 (62%), Positives = 34/43 (78%)
Frame = -3
Query: 233 VKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
VKP +G+A+LFWS+ D DP S+HGGCPV+ G KWS+TKWM
Sbjct: 336 VKPKRGDALLFWSIRPDATLDPSSLHGGCPVIRGNKWSSTKWM 208
>TC15896 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, partial
(45%)
Length = 613
Score = 59.3 bits (142), Expect = 7e-10
Identities = 48/145 (33%), Positives = 72/145 (49%), Gaps = 2/145 (1%)
Frame = +3
Query: 25 LSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNNNDKEAQILRLGYVKPEVLSWSPRI 84
L Q+ R L E T T LLP G T N + + +VLSW+P
Sbjct: 210 LRQIQRPRTTRLLENLTEKETESHLLPAGETGDNF----------ITTIPFQVLSWNPHA 359
Query: 85 ILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSD--VRTSSGMFLSHEERKYPM 142
+ NF + E+C+ + A LK ST+V G+ +S +RTSSG+F+S E K +
Sbjct: 360 LYFPNFATAEQCESIIETAKEGLKPSTLV-LRVGETDESTTGIRTSSGVFISAFEDKTGV 536
Query: 143 IHAIEKRISVYSQIPIENGELMQVL 167
+ IE++I+ ++IP +GE VL
Sbjct: 537 LDVIEEKIARATKIPRTHGEAFNVL 611
>BP061999
Length = 564
Score = 56.6 bits (135), Expect = 5e-09
Identities = 22/34 (64%), Positives = 26/34 (75%)
Frame = -2
Query: 243 FWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
FWSM D +D S+HGGCPV+ G+KWS TKWMR
Sbjct: 563 FWSMKPDTTADTSSLHGGCPVIKGDKWSCTKWMR 462
>TC19415 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, partial
(24%)
Length = 443
Score = 55.5 bits (132), Expect = 1e-08
Identities = 24/47 (51%), Positives = 33/47 (70%)
Frame = +3
Query: 230 GLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
GL V+P G+ +LF+S+ +G DP S+ G CPV+ GEKW A KW+R
Sbjct: 51 GLRVRPRPGDGLLFYSLFPNGTIDPTSLPGSCPVIKGEKWVAPKWIR 191
>TC12024
Length = 469
Score = 54.7 bits (130), Expect = 2e-08
Identities = 23/44 (52%), Positives = 31/44 (70%)
Frame = +1
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGK 119
E+LSW PR + HNFLS EC+Y+ +A P + S+VVD+ TGK
Sbjct: 334 EILSWEPRAFIFHNFLSKVECEYMINLAKPHMAKSSVVDSQTGK 465
>AV423632
Length = 484
Score = 51.6 bits (122), Expect = 1e-07
Identities = 22/40 (55%), Positives = 29/40 (72%)
Frame = +2
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDA 115
EV+SW PR + HNFL+ EEC+YL +A P + STVVD+
Sbjct: 362 EVISWEPRAFVYHNFLTKEECEYLIDIAKPNMHKSTVVDS 481
>TC16912 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, partial
(30%)
Length = 518
Score = 47.8 bits (112), Expect = 2e-06
Identities = 29/75 (38%), Positives = 42/75 (55%), Gaps = 2/75 (2%)
Frame = +1
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVV--DANTGKGIKSDVRTSSGMFL 133
+VLSW PR + NF + E+C + GVA LK S + + T + K +RTSSG+F+
Sbjct: 295 QVLSWKPRALYFPNFATAEQCKSIVGVAKAGLKPSALALREGETEQNTKG-IRTSSGVFV 471
Query: 134 SHEERKYPMIHAIEK 148
S E K + IE+
Sbjct: 472 SSSEDKTGTLAIIEE 516
>AW720438
Length = 463
Score = 35.0 bits (79), Expect = 0.014
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = +3
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVA 103
+SW PR+ L FLS +ECDYL +A
Sbjct: 357 ISWQPRVFLYKGFLSDKECDYLISLA 434
>BP038912
Length = 481
Score = 28.9 bits (63), Expect = 1.0
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = -2
Query: 256 SVHGGCPVLAGEKWSATKW 274
S H CPVL G+ WSA K+
Sbjct: 477 SYHARCPVLKGDMWSAIKF 421
>TC17411 similar to UP|Q8LDG6 (Q8LDG6) Protein kinase, partial (33%)
Length = 632
Score = 26.9 bits (58), Expect = 3.9
Identities = 34/90 (37%), Positives = 42/90 (45%), Gaps = 5/90 (5%)
Frame = -2
Query: 55 TYWNNNNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVD 114
T+ N+ E L +KP VLS+S R I H L + YLR + P LK ST V
Sbjct: 424 TFTNHRQLLEVVNSNLPSLKPHVLSFSARQIPHHLLLLFS--*YLRFLHFP-LKCSTSVL 254
Query: 115 A--NTGKGIK---SDVRTSSGMFLSHEERK 139
A T K I D SSG FL + R+
Sbjct: 253 AILQTLKIILVMILDELLSSGGFLPRDRRR 164
>TC12254 similar to UP|Q94JV6 (Q94JV6) At1g35190/T32G9_27, partial (34%)
Length = 486
Score = 26.9 bits (58), Expect = 3.9
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = -1
Query: 158 IENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRG 191
+ + E M++LR +K++ Y P+HD D N G
Sbjct: 213 LPHDEKMKLLRNDKHRGYTPNHDEILDPENQVHG 112
>BP063325
Length = 429
Score = 26.9 bits (58), Expect = 3.9
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = -3
Query: 174 YYRPHHDYFSDTFNLKRGGQRIATMLM 200
Y+R +DY D + R GQR A LM
Sbjct: 280 YHRGRYDYLKDIISKTRSGQRAA*TLM 200
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,299,585
Number of Sequences: 28460
Number of extensions: 73063
Number of successful extensions: 362
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 353
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 353
length of query: 281
length of database: 4,897,600
effective HSP length: 89
effective length of query: 192
effective length of database: 2,364,660
effective search space: 454014720
effective search space used: 454014720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149471.21


