

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
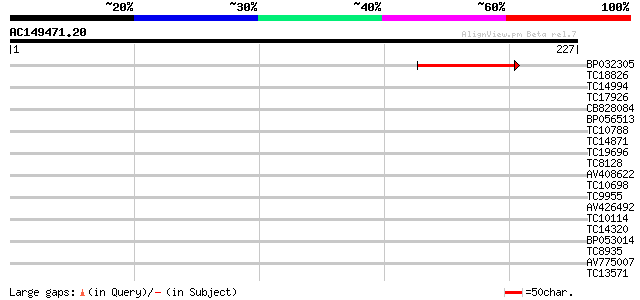
Query= AC149471.20 + phase: 0
(227 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP032305 81 2e-16
TC18826 similar to UP|Q9LUW5 (Q9LUW5) DEAD-box RNA helicase-like... 32 0.12
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 30 0.26
TC17926 similar to UP|IF43_NICPL (P41380) Eukaryotic initiation ... 30 0.26
CB828084 29 0.76
BP056513 29 0.76
TC10788 weakly similar to UP|Q8YG29 (Q8YG29) Cytochrome C-type b... 28 1.00
TC14871 similar to UP|Q85HQ0 (Q85HQ0) NADH dehydrogenase subunit... 28 1.3
TC19696 similar to UP|Q9ZQT8 (Q9ZQT8) WREBP-2 protein, partial (... 27 2.2
TC8128 homologue to UP|TBA3_ELEIN (O22349) Tubulin alpha-3 chain... 27 2.9
AV408622 26 5.0
TC10698 26 5.0
TC9955 similar to UP|Q84MB4 (Q84MB4) At1g22800, partial (10%) 26 5.0
AV426492 26 5.0
TC10114 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-lik... 26 6.5
TC14320 similar to UP|Q41111 (Q41111) Dehydrin, partial (56%) 25 8.5
BP053014 25 8.5
TC8935 similar to UP|Q9SE94 (Q9SE94) Methylenetetrahydrofolate r... 25 8.5
AV775007 25 8.5
TC13571 similar to PIR|F88545|F88545 protein F59B2.11 [imported]... 25 8.5
>BP032305
Length = 467
Score = 80.9 bits (198), Expect = 2e-16
Identities = 39/41 (95%), Positives = 41/41 (99%)
Frame = -2
Query: 164 PSRIKKLIDVEALGLSRLQVLVLDMHPDVKGYSLFTLPQVR 204
PSRIKKLID+EALGLSRLQVLVLDMHPDVKGYSLF+LPQVR
Sbjct: 466 PSRIKKLIDIEALGLSRLQVLVLDMHPDVKGYSLFSLPQVR 344
>TC18826 similar to UP|Q9LUW5 (Q9LUW5) DEAD-box RNA helicase-like protein,
partial (29%)
Length = 782
Score = 31.6 bits (70), Expect = 0.12
Identities = 16/42 (38%), Positives = 25/42 (59%)
Frame = +3
Query: 146 LQEQISLLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLD 187
+ +Q+ L V++ GTP RI L++ AL L +Q +VLD
Sbjct: 579 ISQQMRQLDYGVDVVVGTPGRIIDLLNRGALNLKEVQFMVLD 704
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial (89%)
Length = 1672
Score = 30.4 bits (67), Expect = 0.26
Identities = 42/151 (27%), Positives = 67/151 (43%), Gaps = 5/151 (3%)
Frame = +3
Query: 74 VKLLGGDIKGAFGNCWRQVLCESEVVEGKIPPGSPSVLIVSPSALRSIHLLKGFRFMTKQ 133
V+ LGG I+ + N V + K+P G+PS LI P R + L + F+
Sbjct: 969 VEDLGGYIEDGYLNGVLSVTRALGDWDMKLPKGAPSPLIAEPE-FRQMVLTEDDEFLIIG 1145
Query: 134 CSAVKLFSKHIKLQEQISLLKNRVNIASGTPSRIKKLIDVEALGLS---RLQVLVLDMHP 190
C + + Q +SL++ + P R + + +EAL L+ L V+++
Sbjct: 1146 CDGI---WDVMSSQHAVSLVRKGLR-RHDDPERCARDLVMEALRLNTFDNLTVIIVCFSS 1313
Query: 191 DVKGYSLFTLPQ--VRCASLSSPYLFCLDSL 219
S + PQ +RC SLS+ L L SL
Sbjct: 1314 LDHRESEPSPPQRKLRCCSLSAEALCSLRSL 1406
>TC17926 similar to UP|IF43_NICPL (P41380) Eukaryotic initiation factor 4A-3
(eIF4A-3) (eIF-4A-3), partial (72%)
Length = 982
Score = 30.4 bits (67), Expect = 0.26
Identities = 15/40 (37%), Positives = 24/40 (59%)
Frame = +2
Query: 148 EQISLLKNRVNIASGTPSRIKKLIDVEALGLSRLQVLVLD 187
E I L++ V++ SGTP R+ +I L +++LVLD
Sbjct: 497 EDIRKLEHGVHVVSGTPGRVCDMIKRRTLRTRTIKMLVLD 616
>CB828084
Length = 528
Score = 28.9 bits (63), Expect = 0.76
Identities = 12/27 (44%), Positives = 20/27 (73%)
Frame = -3
Query: 3 KRKGRENNDDRRKKKKKKTETTGSEQL 29
+RK R +++RKKKKK +ET+ S ++
Sbjct: 115 RRKKRWRGEEQRKKKKKSSETSRSVEI 35
>BP056513
Length = 451
Score = 28.9 bits (63), Expect = 0.76
Identities = 16/42 (38%), Positives = 22/42 (52%), Gaps = 8/42 (19%)
Frame = +2
Query: 3 KRKGRENNDDRRKKKK--------KKTETTGSEQLRFFVNEF 36
+R+ RE D+RRKKKK KKT + G E+ + F
Sbjct: 308 QRRRREEEDERRKKKKKRFRRRTEKKTNSEGEEKKGWLARVF 433
>TC10788 weakly similar to UP|Q8YG29 (Q8YG29) Cytochrome C-type biogenesis
protein, partial (25%)
Length = 656
Score = 28.5 bits (62), Expect = 1.00
Identities = 18/55 (32%), Positives = 25/55 (44%), Gaps = 2/55 (3%)
Frame = +3
Query: 104 PPGSPSVLIVSPSAL--RSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNR 156
PP PS+L P AL R L G RF++ KH+ + + L+NR
Sbjct: 204 PPPPPSLLASLPEALSTRQYFALSGHRFLSTARRQPPRSKKHVDIGARARQLQNR 368
>TC14871 similar to UP|Q85HQ0 (Q85HQ0) NADH dehydrogenase subunit 4, partial
(4%)
Length = 588
Score = 28.1 bits (61), Expect = 1.3
Identities = 11/24 (45%), Positives = 19/24 (78%)
Frame = -2
Query: 3 KRKGRENNDDRRKKKKKKTETTGS 26
KR+ R+N + +KKK+K+ +TTG+
Sbjct: 485 KRQKRKNKKNGKKKKRKERKTTGT 414
>TC19696 similar to UP|Q9ZQT8 (Q9ZQT8) WREBP-2 protein, partial (12%)
Length = 496
Score = 27.3 bits (59), Expect = 2.2
Identities = 23/71 (32%), Positives = 34/71 (47%), Gaps = 13/71 (18%)
Frame = -1
Query: 7 RENNDDRRKKKKK--KTETTGSEQLRF---FVNEFQSANDVQLSSIEL--------ESLK 53
REN+ R K+KKK + E ++ F + E QS++ V SS L +L
Sbjct: 364 RENDISRNKRKKKTGRKEREREREIEFTGAAITENQSSSSVSSSSSSLFPAYLQPSSTLS 185
Query: 54 ADSSILELSNS 64
SS+ LS+S
Sbjct: 184 RSSSVSSLSSS 152
>TC8128 homologue to UP|TBA3_ELEIN (O22349) Tubulin alpha-3 chain (Alpha-3
tubulin), complete
Length = 1615
Score = 26.9 bits (58), Expect = 2.9
Identities = 28/110 (25%), Positives = 48/110 (43%)
Frame = +1
Query: 107 SPSVLIVSPSALRSIHLLKGFRFMTKQCSAVKLFSKHIKLQEQISLLKNRVNIASGTPSR 166
SP + P+ RS+ L G ++ C+ LF + L S L+ ++ G P
Sbjct: 979 SPMRCLSLPA*WRSVILGMGSTWLAV*CTGEMLFRRMSMLLLPPSKLRGLFSLLIGAPL- 1155
Query: 167 IKKLIDVEALGLSRLQVLVLDMHPDVKGYSLFTLPQVRCASLSSPYLFCL 216
+ ++ +L L V+ L + S+ TLP +RC+ +P L L
Sbjct: 1156 VSNVVSTTSL-LQLYLVVTLLRCSGLFA*SVTTLPLLRCSLALTPSLISL 1302
>AV408622
Length = 415
Score = 26.2 bits (56), Expect = 5.0
Identities = 10/31 (32%), Positives = 14/31 (44%)
Frame = -3
Query: 78 GGDIKGAFGNCWRQVLCESEVVEGKIPPGSP 108
GG++ G G WR E ++P G P
Sbjct: 356 GGEVAGLRGGFWRGCFLGDGATEDRVPKGFP 264
>TC10698
Length = 538
Score = 26.2 bits (56), Expect = 5.0
Identities = 24/81 (29%), Positives = 36/81 (43%)
Frame = -2
Query: 12 DRRKKKKKKTETTGSEQLRFFVNEFQSANDVQLSSIELESLKADSSILELSNSQNSRDTD 71
D KKKKKK T ++ + F+ F+ + Q + + E+ S +LS +
Sbjct: 471 DYLKKKKKKQSTCYAQDIFMFLKAFRISRYSQ-TVFDHETQSLRQSTRKLSCRFDIGYKF 295
Query: 72 LDVKLLGGDIKGAFGNCWRQV 92
DV + GAF N W QV
Sbjct: 294 KDVTQRSSSMAGAFSN-WFQV 235
>TC9955 similar to UP|Q84MB4 (Q84MB4) At1g22800, partial (10%)
Length = 387
Score = 26.2 bits (56), Expect = 5.0
Identities = 14/50 (28%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Frame = +2
Query: 105 PGSPSVLIVSPSALRSIHLLKGFRFMTKQCS-AVKLFSKHIKLQEQISLL 153
P SP ++ +IHL + +TK CS ++ L + + +++Q+SL+
Sbjct: 194 PSSPYATCLNNVCPLNIHLKSQWIILTKHCSGSISLQYETLNVEDQLSLV 343
>AV426492
Length = 431
Score = 26.2 bits (56), Expect = 5.0
Identities = 12/37 (32%), Positives = 20/37 (53%)
Frame = -2
Query: 88 CWRQVLCESEVVEGKIPPGSPSVLIVSPSALRSIHLL 124
C +V+C V+ G +PP PSV++ + + LL
Sbjct: 265 CQVEVMCRFWVLLGLLPPALPSVVVALKRLVEVLVLL 155
>TC10114 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-like, partial
(35%)
Length = 633
Score = 25.8 bits (55), Expect = 6.5
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = +3
Query: 73 DVKLLGGDIKGAFGNCWRQVLCESE 97
D++LL GD GAFG W CE E
Sbjct: 126 DLELLQGD--GAFGEDWSSEPCEHE 194
>TC14320 similar to UP|Q41111 (Q41111) Dehydrin, partial (56%)
Length = 1231
Score = 25.4 bits (54), Expect = 8.5
Identities = 8/26 (30%), Positives = 18/26 (68%)
Frame = +3
Query: 3 KRKGRENNDDRRKKKKKKTETTGSEQ 28
+ +G+E + ++KKKK+K E +++
Sbjct: 621 EEEGKEGEEKKKKKKKEKKEVKSTDE 698
>BP053014
Length = 441
Score = 25.4 bits (54), Expect = 8.5
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +3
Query: 96 SEVVEGKIPPGSPSVLIVSPSALRSIH 122
S ++GK PP S +V P LRS H
Sbjct: 228 SSSLQGKPPPSPSSSRVVHPWLLRSTH 308
>TC8935 similar to UP|Q9SE94 (Q9SE94) Methylenetetrahydrofolate reductase
, partial (24%)
Length = 580
Score = 25.4 bits (54), Expect = 8.5
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = -3
Query: 195 YSLFTLPQVRCASLSSPYLFCL 216
+S L QVRC +S+P +FC+
Sbjct: 359 FSTGMLVQVRCIMVSTPTVFCI 294
>AV775007
Length = 469
Score = 25.4 bits (54), Expect = 8.5
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = -3
Query: 190 PDVKGYSLFTLPQVRCASLSSPYLFCLDSLTGAA 223
P+ K +S LP + L+ Y F LTGA+
Sbjct: 449 PNFKDFSSVELPSLTALHLNGVYFFNAQHLTGAS 348
>TC13571 similar to PIR|F88545|F88545 protein F59B2.11 [imported] -
Caenorhabditis elegans {Caenorhabditis elegans;},
partial (6%)
Length = 556
Score = 25.4 bits (54), Expect = 8.5
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = -2
Query: 3 KRKGRENNDDRRKKKKKKT 21
+R+ RE ++ KKKKKKT
Sbjct: 198 RREQRERDESEEKKKKKKT 142
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,466,881
Number of Sequences: 28460
Number of extensions: 42872
Number of successful extensions: 376
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 332
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 360
length of query: 227
length of database: 4,897,600
effective HSP length: 87
effective length of query: 140
effective length of database: 2,421,580
effective search space: 339021200
effective search space used: 339021200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC149471.20


