

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149209.11 - phase: 0
(607 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
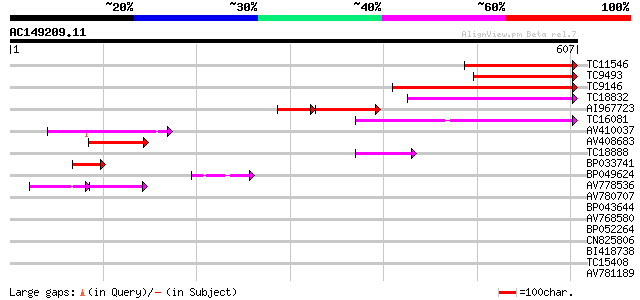
TC11546 homologue to UP|Q940S0 (Q940S0) T5E21.14/T5E21.14 (At1g1... 234 3e-62
TC9493 homologue to UP|Q940S0 (Q940S0) T5E21.14/T5E21.14 (At1g14... 218 3e-57
TC9146 homologue to UP|Q9LIC2 (Q9LIC2) Multispanning membrane pr... 174 3e-44
TC18832 homologue to UP|O04091 (O04091) Endomembrane protein EMP... 171 3e-43
AI967723 103 2e-39
TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), par... 150 4e-37
AV410037 110 5e-25
AV408683 96 2e-20
TC18888 similar to UP|Q9FYQ8 (Q9FYQ8) Endosomal protein-like, pa... 47 7e-06
BP033741 46 2e-05
BP049624 45 3e-05
AV778536 35 4e-05
AV780707 40 0.001
BP043644 35 0.035
AV768580 33 0.17
BP052264 31 0.66
CN825806 30 0.86
BI418738 30 0.86
TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane pro... 30 1.1
AV781189 29 1.9
>TC11546 homologue to UP|Q940S0 (Q940S0) T5E21.14/T5E21.14
(At1g14670/T5E21.14), partial (20%)
Length = 567
Score = 234 bits (597), Expect = 3e-62
Identities = 107/120 (89%), Positives = 114/120 (94%)
Frame = +2
Query: 488 ASVWGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFI 547
ASVWGH+IYTIYSILFIVFIILLIVTAF VALTYFQLAAEDHEWWWRSFLCGGSTGLFI
Sbjct: 2 ASVWGHRIYTIYSILFIVFIILLIVTAFITVALTYFQLAAEDHEWWWRSFLCGGSTGLFI 181
Query: 548 YGYCMFFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
YGYC+++Y+ARSDMSGFM TSFFFGYMACICYGFFLMLG V FRASL+FVRHIYRS+ CE
Sbjct: 182 YGYCLYYYYARSDMSGFMPTSFFFGYMACICYGFFLMLGAVGFRASLLFVRHIYRSITCE 361
>TC9493 homologue to UP|Q940S0 (Q940S0) T5E21.14/T5E21.14
(At1g14670/T5E21.14), partial (19%)
Length = 680
Score = 218 bits (554), Expect = 3e-57
Identities = 100/111 (90%), Positives = 106/111 (95%)
Frame = +1
Query: 497 TIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYGYCMFFYH 556
TIYSILFIVFIILLIVTAF VALTYFQLAAEDHEWWWRSFLCGGSTGLFIY YC+++Y+
Sbjct: 16 TIYSILFIVFIILLIVTAFITVALTYFQLAAEDHEWWWRSFLCGGSTGLFIYAYCLYYYY 195
Query: 557 ARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
ARSDMSGFMQTSFFFGYMACICYGFFLMLGTV FRA+L FVRHIYRS+KCE
Sbjct: 196 ARSDMSGFMQTSFFFGYMACICYGFFLMLGTVGFRAALFFVRHIYRSIKCE 348
>TC9146 homologue to UP|Q9LIC2 (Q9LIC2) Multispanning membrane
protein-like, partial (31%)
Length = 980
Score = 174 bits (442), Expect = 3e-44
Identities = 78/198 (39%), Positives = 123/198 (61%)
Frame = +2
Query: 410 FGTIVVIFLIWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNKYPREIPKLPWYRKTLAQMA 469
FGT+ + +W ++ PL+ +G G + + P +TNK PR++P+ WY + +
Sbjct: 2 FGTMFALVCLWFGISVPLVFVGSYLGFKKPA-IEDPVKTNKIPRQVPEQAWYMTPVFSIL 178
Query: 470 MAGFLPFSAIFIELYYIFASVWGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAAED 529
+ G LPF A+FIEL++I S+W +Q Y I+ LFIVF+ILLI A V L YFQL +ED
Sbjct: 179 IGGILPFGAVFIELFFILTSIWLNQFYYIFGFLFIVFVILLITCAEITVVLCYFQLCSED 358
Query: 530 HEWWWRSFLCGGSTGLFIYGYCMFFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVS 589
+ WWWR++L GS+ L+++ Y +F++ + +++ + +FGYM Y FF++ GT+
Sbjct: 359 YNWWWRAYLTAGSSALYLFLYSIFYFFTKLEITKLVSGILYFGYMVIGSYAFFVLTGTIG 538
Query: 590 FRASLIFVRHIYRSLKCE 607
F A FVR IY S+K +
Sbjct: 539 FYACFWFVRKIYSSVKID 592
>TC18832 homologue to UP|O04091 (O04091) Endomembrane protein EMP70 precusor
isolog, 68664-64364, partial (31%)
Length = 795
Score = 171 bits (433), Expect = 3e-43
Identities = 74/181 (40%), Positives = 110/181 (59%)
Frame = +3
Query: 427 LLVLGGIAGKNSQSEFQAPCRTNKYPREIPKLPWYRKTLAQMAMAGFLPFSAIFIELYYI 486
L +LG G+N PCR PR IP+ WY M G LPF +IFIE+Y++
Sbjct: 3 LALLGTAVGRNFSGAPNNPCRVKTIPRPIPEKKWYLTPSVVSLMGGLLPFGSIFIEMYFV 182
Query: 487 FASVWGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLF 546
F S W +++Y +Y + +VF+IL+IVT + TYF L AE++ W W SF ST ++
Sbjct: 183 FTSFWNYKVYYVYGFMLLVFLILVIVTVCVTIVGTYFLLNAENYHWRWTSFFSAASTAVY 362
Query: 547 IYGYCMFFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKC 606
+Y Y +++Y+ ++ MSGF QTSF+ GY +C G ++ G V + S +FVR IYR++KC
Sbjct: 363 VYLYSIYYYYVKTKMSGFFQTSFYLGYTLMLCLGLGILCGAVGYLGSNLFVRRIYRNIKC 542
Query: 607 E 607
+
Sbjct: 543 D 545
>AI967723
Length = 346
Score = 103 bits (256), Expect(2) = 2e-39
Identities = 51/70 (72%), Positives = 57/70 (80%)
Frame = +1
Query: 328 LALVGVFYPYNRGALFTALVVIYALTSGIASYSAASFYYMIEGKNWVKILVLTGSLFSGP 387
LALVGVFYPYNRGALFTALVVIYALTSGIA Y+A SFY +EG NWV+ L+LTG LF GP
Sbjct: 136 LALVGVFYPYNRGALFTALVVIYALTSGIAGYTATSFYIQLEGSNWVRNLLLTGCLFCGP 315
Query: 388 LFFTFSFLNT 397
F + S +T
Sbjct: 316 SFSSCSASST 345
Score = 77.0 bits (188), Expect(2) = 2e-39
Identities = 34/41 (82%), Positives = 39/41 (94%)
Frame = +3
Query: 287 EETGWKYIHGDVFRYPRFKSLFAAALGTGTQLFTLVIFIFM 327
EETGWKYIHGDVFR+P++KSLFAAALG+G+QLFTL I IFM
Sbjct: 12 EETGWKYIHGDVFRFPKYKSLFAAALGSGSQLFTLTISIFM 134
>TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), partial (37%)
Length = 1189
Score = 150 bits (380), Expect = 4e-37
Identities = 78/240 (32%), Positives = 124/240 (51%), Gaps = 3/240 (1%)
Frame = +3
Query: 371 KNWVKILVLTGSLFSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVL 430
+ W I F G F + LN + + NST A+P +F +W ++ PL ++
Sbjct: 27 EGWRSISWSAACFFPGIAFIILTILNFILWSSNSTGAIPISLYFELFFLWFCISVPLTLI 206
Query: 431 GGIAGKNSQSEFQAPCRTNKYPREIP--KLPWYRKTLAQMAMAGFLPFSAIFIELYYIFA 488
GG G ++ + P RTN+ PREIP K P + L AG LPF +FIEL++I +
Sbjct: 207 GGFMGTKAEP-IEFPVRTNQIPREIPARKYPSWLLVLG----AGTLPFGTLFIELFFILS 371
Query: 489 SVWGHQIYTIYSILFIVFIILLIVTAFGNVALTYFQLAAEDHEWWWRSFLCGGSTGLFIY 548
S+W + Y ++ L +V ++L+IV A +V LTY L ED WWW++F GS L+++
Sbjct: 372 SIWLGRFYYVFGFLLVVLLLLVIVCAEVSVVLTYMHLCVEDWRWWWKAFFASGSVALYVF 551
Query: 549 GYCM-FFYHARSDMSGFMQTSFFFGYMACICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
Y + + +SG + + GY + L GT+ F S FV +++ S+K +
Sbjct: 552 LYSVNYLVFDLQSLSGPVSAVLYLGYSLLMAVAIMLSTGTIGFLMSFYFVHYLFSSVKID 731
>AV410037
Length = 425
Score = 110 bits (276), Expect = 5e-25
Identities = 52/137 (37%), Positives = 82/137 (58%), Gaps = 3/137 (2%)
Frame = +3
Query: 41 SDHRYKEGDFVPFYANKVGPFHNPSETYRYFDLPFCSPEN---VEEKKEDLGEVLNGDRL 97
SDH+Y+ D V + NKVGP++NP ETY Y+ LPFC P + K LGEVL G+ L
Sbjct: 15 SDHKYQREDPVTLWVNKVGPYNNPQETYNYYSLPFCHPSSSSSAAHKWGGLGEVLGGNEL 194
Query: 98 VVAPYKLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDLPIWGFLGRFETD 157
+ + +++FL + CQ L+ V QF+ A+ +Y+++ + DDLP+WG++G D
Sbjct: 195 IDSQIEIKFLRNVDKTVFCQIDLSDANVKQFKDAIENNYWFEFFMDDLPLWGYVGELHPD 374
Query: 158 EKDVDTNEATVYLFRNV 174
K+ D + +Y +N+
Sbjct: 375 -KNSDNGKHVLYTHKNI 422
>AV408683
Length = 194
Score = 95.5 bits (236), Expect = 2e-20
Identities = 44/64 (68%), Positives = 51/64 (78%)
Frame = +2
Query: 85 KEDLGEVLNGDRLVVAPYKLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDD 144
KE LGEVLNGDRLV APYKLEF DK +C+K L + +V +FR AV+KDY+YQMYYDD
Sbjct: 2 KEALGEVLNGDRLVSAPYKLEFQRDKDSVVVCKKTLKKDDVVKFRSAVMKDYYYQMYYDD 181
Query: 145 LPIW 148
LPIW
Sbjct: 182 LPIW 193
>TC18888 similar to UP|Q9FYQ8 (Q9FYQ8) Endosomal protein-like, partial (12%)
Length = 264
Score = 47.4 bits (111), Expect = 7e-06
Identities = 22/65 (33%), Positives = 33/65 (49%)
Frame = +1
Query: 371 KNWVKILVLTGSLFSGPLFFTFSFLNTVAVAYNSTAALPFGTIVVIFLIWTLVTSPLLVL 430
K WV + F G F + LN + +ST A+PF VV+ L+W ++ PL ++
Sbjct: 58 KGWVSVAWKAACFFPGIAFLILTSLNFLLWGSHSTGAIPFSLFVVLILLWFCISVPLTLV 237
Query: 431 GGIAG 435
GG G
Sbjct: 238 GGYFG 252
>BP033741
Length = 526
Score = 45.8 bits (107), Expect = 2e-05
Identities = 17/35 (48%), Positives = 25/35 (70%)
Frame = +3
Query: 68 YRYFDLPFCSPENVEEKKEDLGEVLNGDRLVVAPY 102
Y Y+ LP+C P+ + + E+LGEVL GDR+ +PY
Sbjct: 129 YSYYSLPYCRPDQIVDSAENLGEVLRGDRIENSPY 233
>BP049624
Length = 424
Score = 45.1 bits (105), Expect = 3e-05
Identities = 25/68 (36%), Positives = 41/68 (59%)
Frame = -3
Query: 195 NAVVDLTEDREVNVDFTYSVKWIETDIPFEKRLEKYSQTSSLSHHLEIHWFSIINSCVTV 254
N ++ E++E+ FTY V + E+D+ + R + Y L + +IHWFSI+NS + V
Sbjct: 215 NTPQEVEENKEII--FTYDVDFQESDVKWASRWDAYL----LMNDDQIHWFSIVNSLMIV 54
Query: 255 LLLTGFLA 262
L L+G +A
Sbjct: 53 LFLSGMVA 30
>AV778536
Length = 550
Score = 35.0 bits (79), Expect(2) = 4e-05
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +1
Query: 86 EDLGEVLNGDRLVVAPYKLEFLIDKKPESICQKMLTRKEVAQFRHAVLKDYFYQMYYDDL 145
+ LGEVL GDR+ + Y + + +C+ L K F+ + +Y M D+L
Sbjct: 277 KSLGEVLRGDRIENSRYVFKMREPEMCNIVCKIKLDAKTAKAFKEKINDEYRVNMILDNL 456
Query: 146 PI 147
P+
Sbjct: 457 PL 462
Score = 28.9 bits (63), Expect(2) = 4e-05
Identities = 14/65 (21%), Positives = 30/65 (45%)
Frame = +3
Query: 22 SAIVILCCFCCINYVTSDASDHRYKEGDFVPFYANKVGPFHNPSETYRYFDLPFCSPENV 81
SA+++L + + + +GD + NK+ Y Y+ LP+C+P+ +
Sbjct: 87 SAVLLLLIHGAHCFYLPGVAPQDFIKGDQLQVKVNKLSSIKTQLP-YSYYSLPYCAPKKI 263
Query: 82 EEKKE 86
++ E
Sbjct: 264 QDSAE 278
>AV780707
Length = 429
Score = 39.7 bits (91), Expect = 0.001
Identities = 21/64 (32%), Positives = 36/64 (55%), Gaps = 1/64 (1%)
Frame = +1
Query: 43 HRYKEGDFVPFYANKVGPFHNPSETYRYFDLPFCSP-ENVEEKKEDLGEVLNGDRLVVAP 101
H+Y GD + N + + + Y+ LPF +P V++ E+LGE+L GDR+ +P
Sbjct: 232 HKYAVGDELSVKVNSLTSI-DTEMPFSYYSLPFRTPLGGVKDSAENLGELLMGDRIENSP 408
Query: 102 YKLE 105
Y+ +
Sbjct: 409 YRFK 420
>BP043644
Length = 500
Score = 35.0 bits (79), Expect = 0.035
Identities = 14/31 (45%), Positives = 21/31 (67%)
Frame = -2
Query: 577 ICYGFFLMLGTVSFRASLIFVRHIYRSLKCE 607
+ YGFF++ GT+ F A +F R IY S+K +
Sbjct: 493 LSYGFFVVTGTIGFYACFLFTRLIYSSVKVD 401
>AV768580
Length = 485
Score = 32.7 bits (73), Expect = 0.17
Identities = 13/28 (46%), Positives = 19/28 (67%)
Frame = -3
Query: 580 GFFLMLGTVSFRASLIFVRHIYRSLKCE 607
G ++ G V F S +FVR IYR++KC+
Sbjct: 474 GLGILCGAVGFLGSNLFVRRIYRNIKCD 391
>BP052264
Length = 526
Score = 30.8 bits (68), Expect = 0.66
Identities = 11/43 (25%), Positives = 22/43 (50%)
Frame = +2
Query: 20 LVSAIVILCCFCCINYVTSDASDHRYKEGDFVPFYANKVGPFH 62
L + ++C FCCI+Y+ ++ ++ E + N +G H
Sbjct: 383 LTGLVSVICFFCCISYINNNRPXGKFSE------FXNNMGTMH 493
>CN825806
Length = 519
Score = 30.4 bits (67), Expect = 0.86
Identities = 15/33 (45%), Positives = 23/33 (69%), Gaps = 1/33 (3%)
Frame = +1
Query: 177 EILYNN-DRIIDVFVKNDPNAVVDLTEDREVNV 208
+I+YNN R++D VK DP+ VVD+ + + V V
Sbjct: 415 KIVYNNIGRLVDHLVKRDPSIVVDIKKIKHVLV 513
>BI418738
Length = 545
Score = 30.4 bits (67), Expect = 0.86
Identities = 13/28 (46%), Positives = 18/28 (63%), Gaps = 2/28 (7%)
Frame = +3
Query: 28 CCFCCINYV--TSDASDHRYKEGDFVPF 53
CC+CC V TSD+ DH ++ +VPF
Sbjct: 27 CCYCCC*VVAETSDSLDHYHQGTRYVPF 110
>TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane protein-like,
partial (10%)
Length = 879
Score = 30.0 bits (66), Expect = 1.1
Identities = 13/46 (28%), Positives = 23/46 (49%)
Frame = +2
Query: 416 IFLIWTLVTSPLLVLGGIAGKNSQSEFQAPCRTNKYPREIPKLPWY 461
+ +W ++ PL+ +GG + T+K P +IP+ PWY
Sbjct: 47 LVFLWFGISVPLVFVGGHIDFTKKKVI*DLVETSKIPWQIPEQPWY 184
>AV781189
Length = 420
Score = 29.3 bits (64), Expect = 1.9
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = -3
Query: 532 WWWRSFLCGGSTGLFIYGYCMF 553
WWW FLC + L IY Y +F
Sbjct: 94 WWWWVFLCSLACTLTIYLYAIF 29
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.329 0.143 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,654,470
Number of Sequences: 28460
Number of extensions: 174242
Number of successful extensions: 1737
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 1706
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1731
length of query: 607
length of database: 4,897,600
effective HSP length: 96
effective length of query: 511
effective length of database: 2,165,440
effective search space: 1106539840
effective search space used: 1106539840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149209.11


