

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.3 + phase: 0 /pseudo
(523 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
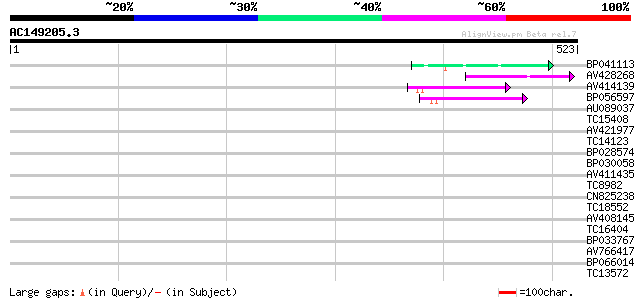
Score E
Sequences producing significant alignments: (bits) Value
BP041113 50 1e-06
AV428268 47 8e-06
AV414139 47 1e-05
BP056597 44 5e-05
AU089037 37 0.008
TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane pro... 36 0.018
AV421977 33 0.087
TC14123 similar to UP|Q93X25 (Q93X25) 2-Cys peroxiredoxin, parti... 33 0.11
BP028574 33 0.15
BP030058 32 0.19
AV411435 32 0.19
TC8982 similar to UP|Q84QW4 (Q84QW4) OJ1191_A10.2 protein, parti... 31 0.43
CN825238 31 0.57
TC18552 similar to UP|Q9FIM2 (Q9FIM2) Cell division protein FtsH... 30 1.3
AV408145 30 1.3
TC16404 similar to UP|P93320 (P93320) Cdc2MsC protein, partial (... 29 1.6
BP033767 28 2.8
AV766417 28 2.8
BP066014 28 2.8
TC13572 28 3.7
>BP041113
Length = 503
Score = 49.7 bits (117), Expect = 1e-06
Identities = 45/140 (32%), Positives = 53/140 (37%), Gaps = 9/140 (6%)
Frame = +1
Query: 371 GGW-WGAWQWRRHLWVWEEDLV-EECGDLLLT------VSLQELETDRWMWLSDQIRGYT 422
GGW G W W+ W LV E G L L V L E D+W+WL YT
Sbjct: 100 GGWNHGVWTWKLS---WRRALVGRELGWLELMMKDLSGVCLSEGVEDKWLWLPGGT--YT 264
Query: 423 VRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITF-VE 481
V Y L + IW P V A R L RLPT NL R +
Sbjct: 265 VNSAYSFLQAPTLADTDPIFATIWRTVAP-SVKAFAWRCLLGRLPTYDNLIKRQVVVDPA 441
Query: 482 DRICVTGCGHVEDTTHMFLS 501
+C G VE TH+ +
Sbjct: 442 MTVCKFCQGEVESVTHLLFA 501
>AV428268
Length = 429
Score = 47.0 bits (110), Expect = 8e-06
Identities = 31/101 (30%), Positives = 46/101 (44%)
Frame = +3
Query: 421 YTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFV 480
Y+V+ YD L +L+W +VP L+ ++L +R+ TK NL RG+ V
Sbjct: 6 YSVKSAYDFLHGCANAQWDNIFKLLWSVKVPSNAIALSWKVLINRVQTKVNLNRRGV--V 179
Query: 481 EDRICVTGCGHVEDTTHMFLSCPIFGALLSMVQTWVGVEGV 521
+C E T H+ SCPI + S + GV V
Sbjct: 180 ISNVCPLCSLDEESTDHLLFSCPIV*RIWSKISE*FGVFSV 302
>AV414139
Length = 309
Score = 46.6 bits (109), Expect = 1e-05
Identities = 33/101 (32%), Positives = 42/101 (40%), Gaps = 6/101 (5%)
Frame = -2
Query: 368 VGGGGWW--GAWQW----RRHLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGY 421
+GG G W G W+W RR L V E+ + L V LQ+ DRW W D Y
Sbjct: 308 IGGMGQWHNGVWEWEFRWRRELSVREQRQEADLIQALAGVCLQQGTEDRWKWTLDTEDIY 129
Query: 422 TVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLL 462
TVR Y +L + E +W P + RLL
Sbjct: 128 TVRSAY*LLLDPIADSDSEVYEKVWVGIAPSNANAFVWRLL 6
>BP056597
Length = 414
Score = 44.3 bits (103), Expect = 5e-05
Identities = 27/110 (24%), Positives = 49/110 (44%), Gaps = 11/110 (10%)
Frame = +3
Query: 379 WRRHLWVWE---------EDLVE--ECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVY 427
W + WVW+ ED+++ ++ + SL + D W+W+ + Y+V+ Y
Sbjct: 69 WLENKWVWDFSWKNPISGEDVLKLGVMKQIVSSFSLINGKKDTWVWILEGDGQYSVKSAY 248
Query: 428 DMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPTKSNLADRGI 477
D+L+ + +W P L R+ DR+ TK NL+ R +
Sbjct: 249 DLLSGLDTTIGISVFSKLWKACAPSNAVALGWRVFLDRIQTKDNLSRRHV 398
>AU089037
Length = 245
Score = 37.0 bits (84), Expect = 0.008
Identities = 18/61 (29%), Positives = 29/61 (47%), Gaps = 1/61 (1%)
Frame = -1
Query: 445 IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTT-HMFLSCP 503
+W P A RL+ DR+ T+ NL R + + C E+++ H+F SCP
Sbjct: 191 LWKVPAPSNAVSFAWRLILDRVQTRGNLRRRQVIQQSEEALCPMCSQCEESSSHLFFSCP 12
Query: 504 I 504
+
Sbjct: 11 L 9
>TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane protein-like,
partial (10%)
Length = 879
Score = 35.8 bits (81), Expect = 0.018
Identities = 22/76 (28%), Positives = 31/76 (39%), Gaps = 4/76 (5%)
Frame = -1
Query: 375 GAWQW----RRHLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDML 430
G W W RR+ E LV + +LL +V L E D W W ++ YTV+ Y L
Sbjct: 774 GVWTWSFIRRRNPSTRENLLVADLCNLLESVPLSEDTNDSWSWTANPEGQYTVQTAYKFL 595
Query: 431 TSQE*PHVHQNMELIW 446
+ +W
Sbjct: 594 RASSNEFTDPIFHFVW 547
>AV421977
Length = 535
Score = 33.5 bits (75), Expect = 0.087
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = -1
Query: 369 GGGGWWGAWQWRRHLWVWEEDLVEECGDLLL 399
GGGGW G W+WR+ WV EE +++
Sbjct: 271 GGGGWRGWWKWRKAGWVSGSPEEEEVNRVMI 179
>TC14123 similar to UP|Q93X25 (Q93X25) 2-Cys peroxiredoxin, partial (89%)
Length = 1130
Score = 33.1 bits (74), Expect = 0.11
Identities = 17/45 (37%), Positives = 20/45 (43%), Gaps = 8/45 (17%)
Frame = -1
Query: 373 WW----GAWQWRRHLWVWEE----DLVEECGDLLLTVSLQELETD 409
WW G+W WR WVWE L E + L L LET+
Sbjct: 173 WWREG*GSWGWRERWWVWERRGV*GLASERREKLQNTPLTVLETE 39
>BP028574
Length = 454
Score = 32.7 bits (73), Expect = 0.15
Identities = 16/64 (25%), Positives = 28/64 (43%)
Frame = +3
Query: 409 DRWMWLSDQIRGYTVRGVYDMLTSQE*PHVHQNMELIWHKQVPLKVSILALRLLRDRLPT 468
D W+W+ D +TV+ Y+ L + +W + P LA R+ +R+P
Sbjct: 261 DSWVWIKDGSGSFTVKSAYEELHGEIQEIEDNFFRKLWQSKAPSNSIALAWRVGLNRVPC 440
Query: 469 KSNL 472
+ L
Sbjct: 441 MAEL 452
>BP030058
Length = 455
Score = 32.3 bits (72), Expect = 0.19
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = +3
Query: 373 WWGAWQWRRHLWVWE 387
WWG W+W R W W+
Sbjct: 324 WWGRWRWWRWRWAWQ 368
>AV411435
Length = 426
Score = 32.3 bits (72), Expect = 0.19
Identities = 20/58 (34%), Positives = 24/58 (40%)
Frame = +3
Query: 445 IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGHVEDTTHMFLSC 502
IW QVP KV R + +P K NL R + D C ED H L+C
Sbjct: 177 IWGAQVPKKVRSFLWRAANNAIPVKRNLKRRNMG--RDDFCPICNKGPEDINHALLAC 344
>TC8982 similar to UP|Q84QW4 (Q84QW4) OJ1191_A10.2 protein, partial (10%)
Length = 901
Score = 31.2 bits (69), Expect = 0.43
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = +1
Query: 362 CEEHVLVGGGGWWGAWQWRRHLWVWE 387
C + VGGG W+W+ + W WE
Sbjct: 538 CGQGAAVGGGHDSSGWRWKHYQWEWE 615
>CN825238
Length = 692
Score = 30.8 bits (68), Expect = 0.57
Identities = 17/47 (36%), Positives = 24/47 (50%)
Frame = +3
Query: 445 IWHKQVPLKVSILALRLLRDRLPTKSNLADRGITFVEDRICVTGCGH 491
+W VP K+ A R+ R+ +P + NL RGI + C T C H
Sbjct: 534 LWRTPVPNKMKHFAYRVARNIIPCRENLQRRGIAVQNE--CPT-CNH 665
>TC18552 similar to UP|Q9FIM2 (Q9FIM2) Cell division protein FtsH, partial
(10%)
Length = 643
Score = 29.6 bits (65), Expect = 1.3
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -2
Query: 369 GGGGWWGAWQWRRHLWVWEEDLVEECGDLLLTVSL 403
G GGWW WQW+ + +E G LLL + +
Sbjct: 282 GKGGWWWRWQWQPII------QAQEIGALLLQLGI 196
>AV408145
Length = 422
Score = 29.6 bits (65), Expect = 1.3
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = -2
Query: 365 HVLVGGGGWWGAWQWR 380
H + GGGGWW W+WR
Sbjct: 91 HHVGGGGGWW-RWEWR 47
>TC16404 similar to UP|P93320 (P93320) Cdc2MsC protein, partial (15%)
Length = 584
Score = 29.3 bits (64), Expect = 1.6
Identities = 15/48 (31%), Positives = 20/48 (41%), Gaps = 5/48 (10%)
Frame = +1
Query: 373 WWGAWQWRRHLWVWEEDLVEECGDLLLTVSLQELETDRW-----MWLS 415
WW +W WR LW W L + S L+ D W +WL+
Sbjct: 100 WWSSW-WRWRLWGWSSKLSSTRRSIWGLSSWSWLKHDGWKSQSTVWLA 240
>BP033767
Length = 541
Score = 28.5 bits (62), Expect = 2.8
Identities = 10/18 (55%), Positives = 10/18 (55%)
Frame = -3
Query: 369 GGGGWWGAWQWRRHLWVW 386
GGGG W W WR W W
Sbjct: 257 GGGGKWVFWVWR*GRW*W 204
>AV766417
Length = 440
Score = 28.5 bits (62), Expect = 2.8
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 2/44 (4%)
Frame = -3
Query: 369 GGGGWWGAWQWRRHLWV--WEEDLVEECGDLLLTVSLQELETDR 410
GGG WW +W R LW+ W LV C D+ + ++L ++R
Sbjct: 243 GGGRWW-CERWGRWLWLGRWMRVLVVGC-DISICITLLAPASER 118
>BP066014
Length = 535
Score = 28.5 bits (62), Expect = 2.8
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -2
Query: 362 CEEHVLVGGGGWWGAWQWRRHLWV 385
C++ GGG WW +WRR++ V
Sbjct: 162 CKQESFEGGGCWWW**EWRRYVRV 91
>TC13572
Length = 534
Score = 28.1 bits (61), Expect = 3.7
Identities = 16/60 (26%), Positives = 27/60 (44%)
Frame = -3
Query: 371 GGWWGAWQWRRHLWVWEEDLVEECGDLLLTVSLQELETDRWMWLSDQIRGYTVRGVYDML 430
G W ++WRR L E D ++ ++ V L D+W+W + Y+V + L
Sbjct: 214 GVWNWDFRWRRPLAGRELDWLQALSLDIVRVPLLAGVPDKWVWKPSEDGSYSVNSAFIFL 35
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.364 0.164 0.688
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,274,551
Number of Sequences: 28460
Number of extensions: 194107
Number of successful extensions: 3774
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 2130
Number of HSP's successfully gapped in prelim test: 118
Number of HSP's that attempted gapping in prelim test: 1261
Number of HSP's gapped (non-prelim): 2637
length of query: 523
length of database: 4,897,600
effective HSP length: 94
effective length of query: 429
effective length of database: 2,222,360
effective search space: 953392440
effective search space used: 953392440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 57 (26.6 bits)
Medicago: description of AC149205.3


