

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.17 + phase: 0 /pseudo
(626 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
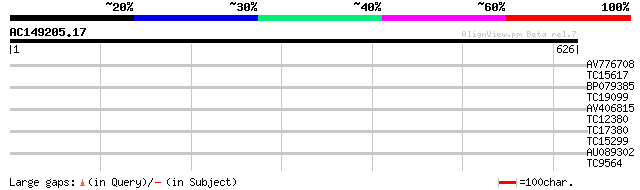
Score E
Sequences producing significant alignments: (bits) Value
AV776708 32 0.23
TC15617 similar to GB|AAN28738.1|23505757|AY143799 At2g02050/F14... 29 2.0
BP079385 29 2.0
TC19099 similar to UP|QUA1_ARATH (Q9LSG3) Glycosyltransferase QU... 28 5.8
AV406815 27 7.5
TC12380 similar to GB|BAC55960.1|28071265|AB087408 actin binding... 27 7.5
TC17380 similar to UP|Q84UH4 (Q84UH4) Dehydroascorbate reductase... 27 7.5
TC15299 similar to UP|Q84MJ5 (Q84MJ5) Methylmalonate semialdehyd... 27 9.8
AU089302 27 9.8
TC9564 27 9.8
>AV776708
Length = 603
Score = 32.3 bits (72), Expect = 0.23
Identities = 21/56 (37%), Positives = 30/56 (53%)
Frame = -2
Query: 479 KCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPKLLKT 534
K E +PP FP ++VG KS DDGS + + + D D P G +PKL+ +
Sbjct: 569 KVENDVDVPPGFPP-QVVGTV----KSFDDGSRSVKVEN--DADVPPGFLPKLVSS 423
>TC15617 similar to GB|AAN28738.1|23505757|AY143799 At2g02050/F14H20.12
{Arabidopsis thaliana;}, partial (85%)
Length = 573
Score = 29.3 bits (64), Expect = 2.0
Identities = 25/85 (29%), Positives = 35/85 (40%), Gaps = 8/85 (9%)
Frame = +3
Query: 538 EKSVEDGLLAEKYVDGGAPLPAKDNTP-------TPLIFVDDCKHVLEDGNESEEARLST 590
E VE LL +K G TP PL + D+C H+L N+ +A
Sbjct: 12 ESGVELRLLKKKMATGEGSSKKMIATPEEMVEARVPLAYRDECAHLLIPLNKCRQAEFYL 191
Query: 591 DTICQSENQTESYSYLS-ELIVEEL 614
C EN+ SY EL++E +
Sbjct: 192 PWKC--ENERHSYEKCEYELVMERM 260
>BP079385
Length = 271
Score = 29.3 bits (64), Expect = 2.0
Identities = 14/35 (40%), Positives = 20/35 (57%)
Frame = +3
Query: 149 FGGILSLWSSCFSPNLKIRRLERWNRSSSALKAKR 183
F LSLW +C S ++ I E+W +S K+KR
Sbjct: 141 FHSKLSLWLTCRS*SVTINIKEKWGKSWGTQKSKR 245
>TC19099 similar to UP|QUA1_ARATH (Q9LSG3) Glycosyltransferase QUASIMODO1 ,
partial (35%)
Length = 597
Score = 27.7 bits (60), Expect = 5.8
Identities = 16/38 (42%), Positives = 18/38 (47%)
Frame = +2
Query: 527 SIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTP 564
SIPK L +S E AEKY D G P P + P
Sbjct: 191 SIPKSLHCLSMRLMEERIAHAEKYSDEGKPTPPEVEDP 304
>AV406815
Length = 426
Score = 27.3 bits (59), Expect = 7.5
Identities = 9/27 (33%), Positives = 15/27 (55%), Gaps = 3/27 (11%)
Frame = +1
Query: 279 CWTLMYNHPFTWFK---FGFGKGSKIC 302
CWT+ + HP W K G+ +++C
Sbjct: 175 CWTIQHFHPLAWIKDVEKSMGRWTQLC 255
>TC12380 similar to GB|BAC55960.1|28071265|AB087408 actin binding protein
{Arabidopsis thaliana;} , partial (9%)
Length = 800
Score = 27.3 bits (59), Expect = 7.5
Identities = 14/43 (32%), Positives = 23/43 (52%)
Frame = +1
Query: 574 KHVLEDGNESEEARLSTDTICQSENQTESYSYLSELIVEELNR 616
KH E+G E E+ + TD I + +++ E +LI +NR
Sbjct: 433 KHQDEEGTEEEQFKSFTDAINEEDSEEEEEKEEVKLISSIINR 561
>TC17380 similar to UP|Q84UH4 (Q84UH4) Dehydroascorbate reductase, complete
Length = 875
Score = 27.3 bits (59), Expect = 7.5
Identities = 16/36 (44%), Positives = 18/36 (49%), Gaps = 2/36 (5%)
Frame = +2
Query: 481 EPSAKIPPEFPS--HKLVGCTVTIGKSSDDGSNTSQ 514
EPS PPEF S K+ G V+ KS D T Q
Sbjct: 344 EPSLVTPPEFASVGSKIFGSFVSFLKSKDANDGTEQ 451
>TC15299 similar to UP|Q84MJ5 (Q84MJ5) Methylmalonate semialdehyde
dehydrogenase (Fragment), partial (31%)
Length = 570
Score = 26.9 bits (58), Expect = 9.8
Identities = 11/21 (52%), Positives = 12/21 (56%), Gaps = 3/21 (14%)
Frame = +2
Query: 141 NHFGRCHCFGGIL---SLWSS 158
NH RCHC G+L WSS
Sbjct: 2 NHLSRCHCRHGVLQGGDFWSS 64
>AU089302
Length = 327
Score = 26.9 bits (58), Expect = 9.8
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = -2
Query: 140 NNHFGRCHCFGGILSLW 156
+NH G C C G L+LW
Sbjct: 200 HNHVGFCACLAGYLALW 150
>TC9564
Length = 646
Score = 26.9 bits (58), Expect = 9.8
Identities = 29/94 (30%), Positives = 39/94 (40%), Gaps = 3/94 (3%)
Frame = -3
Query: 397 YHIELLCNSDLIKMFLVNQTDPKAIAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMK 456
Y + L NS I P A + ++SR S S FPS+ +T + M
Sbjct: 485 YQLLLTSNSSTIHSLNSIT*SPSANSCIHFSRSSSLNST-FPSEVGGFSLTISASGQKMT 309
Query: 457 SVLRSQ---GFVTNFEPRKRSASSSKCEPSAKIP 487
S S VTN +KR+ SSS+ PS P
Sbjct: 308 SAFSSDEGNSGVTNKSSKKRNESSSRVFPSMLSP 207
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.350 0.153 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,740,511
Number of Sequences: 28460
Number of extensions: 210042
Number of successful extensions: 1941
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 1918
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1941
length of query: 626
length of database: 4,897,600
effective HSP length: 96
effective length of query: 530
effective length of database: 2,165,440
effective search space: 1147683200
effective search space used: 1147683200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149205.17


