

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.7 + phase: 0
(600 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
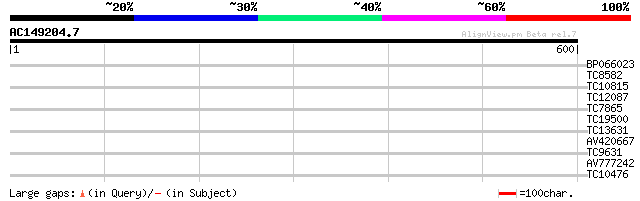
Score E
Sequences producing significant alignments: (bits) Value
BP066023 38 0.004
TC8582 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 33 0.13
TC10815 similar to UP|Q940Y3 (Q940Y3) At2g17400 (At2g17400/At2g1... 32 0.23
TC12087 30 1.1
TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinas... 30 1.5
TC19500 30 1.5
TC13631 similar to UP|Q9SGN7 (Q9SGN7) F3M18.17 (At1g28390/F3M18_... 28 4.3
AV420667 27 7.3
TC9631 similar to GB|AAC19270.1|3193286|T14P8 coded for by A. th... 27 7.3
AV777242 27 7.3
TC10476 27 7.3
>BP066023
Length = 507
Score = 38.1 bits (87), Expect = 0.004
Identities = 18/34 (52%), Positives = 21/34 (60%), Gaps = 4/34 (11%)
Frame = -2
Query: 387 IPLGPNHQAEVPEWTGTTHKSDS----KWLGTQI 416
IP+GP Q EVP W T + +S KWLGTQI
Sbjct: 452 IPIGPRFQVEVPNWDVTRNSPNSDEDWKWLGTQI 351
>TC8582 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (65%)
Length = 1341
Score = 33.1 bits (74), Expect = 0.13
Identities = 53/235 (22%), Positives = 94/235 (39%), Gaps = 18/235 (7%)
Frame = +1
Query: 37 EEYGLGVKVGSSVKLVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVKF------------ 84
E +G+ VKVG +VK+ S+ G + GE K+D + V
Sbjct: 64 EFWGVEVKVGQTVKVEPSEPFPGY-IHISQAALGEATKKDKAAEPVILYLKVEDKKYVLG 240
Query: 85 ---GERLTELQAELVLDDYGEGDVGDEVKSVYGCRKKLCDTNRVKVVNPELNASELEKVY 141
+ + L +LVLDD E + SVY C K+ T+ + + E E++
Sbjct: 241 TLNRDDIPHLSMDLVLDDKSELSHNSKGASVYFCGYKVL-TDDGDASDFSESDEEEEELM 417
Query: 142 EYIDGRKSCGTNKMKDTNLVSNMAKKVESEG--LVDVLMQDCKTNETSLRKLVNQNAAME 199
E +G+ T K V+ AK + G V + D K +E + A ++
Sbjct: 418 EQENGKPETKTEDAK----VAKTAKPAAAAGPSAKQVKIVDPKKDE--------EEADLD 561
Query: 200 IMD-DVSDVANSMPGLSDGSKSCANDDANDSGDDVLMLDPSSVNRESFGRKRKRD 253
+ DVS + + + S ++D+++D + P+ + G+KR +
Sbjct: 562 VSSPDVSGYEDDLISADEDSDDESDDESDDEEE-----TPTPAKKVDQGKKRPNE 711
>TC10815 similar to UP|Q940Y3 (Q940Y3) At2g17400 (At2g17400/At2g17400),
partial (18%)
Length = 1027
Score = 32.3 bits (72), Expect = 0.23
Identities = 21/71 (29%), Positives = 34/71 (47%), Gaps = 3/71 (4%)
Frame = +2
Query: 6 QTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEY---GLGVKVGSSVKLVYSKYLSGLET 62
+ L+ KL+ V GGYDVV ++LW VGE + V + ++ Y K L E
Sbjct: 614 EPLNCLKLWRAVIRLGGYDVVTGSKLWRQVGESFHPPKTCTTVSWTFRIFYEKALLEYEK 793
Query: 63 PLKNVVDGEFP 73
+ + + + P
Sbjct: 794 HKREIGELQLP 826
>TC12087
Length = 374
Score = 30.0 bits (66), Expect = 1.1
Identities = 17/47 (36%), Positives = 24/47 (50%)
Frame = +1
Query: 447 SVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKK 493
SV+ F AE++ ++ V F N K GEE RCW +E +K
Sbjct: 133 SVQFRVFRSAEQAIYHSVKSRVLFRSVNASKNGEERCRCWRRQEGRK 273
>TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinase
(S-adenosyl-L-homocysteine hydrolase) (AdoHcyase) ,
complete
Length = 1852
Score = 29.6 bits (65), Expect = 1.5
Identities = 16/46 (34%), Positives = 20/46 (42%)
Frame = -3
Query: 540 YQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSNSFTLLSPKKPQP 585
Y+N H P N +S KG +Q+ NSF LL QP
Sbjct: 1670 YKNHHNPTNFPPQIIKSLMKIRKGKQNNQSGSQNSFFLLPESHHQP 1533
>TC19500
Length = 448
Score = 29.6 bits (65), Expect = 1.5
Identities = 12/34 (35%), Positives = 16/34 (46%)
Frame = -2
Query: 378 ILDAPPTVNIPLGPNHQAEVPEWTGTTHKSDSKW 411
+L PP + PL P H +P + TT S W
Sbjct: 429 LLPTPPPLLFPLKPRHSRRIPSYHNTT*TSTFPW 328
>TC13631 similar to UP|Q9SGN7 (Q9SGN7) F3M18.17 (At1g28390/F3M18_17),
partial (8%)
Length = 656
Score = 28.1 bits (61), Expect = 4.3
Identities = 14/47 (29%), Positives = 23/47 (48%)
Frame = -3
Query: 392 NHQAEVPEWTGTTHKSDSKWLGTQIWPLQIVKSKLLEGEPVGKGRQD 438
NH T TTHK++S+ + + L LL+ +P+ +QD
Sbjct: 303 NHSMNEEAPTYTTHKANSQSPSWKAFSLYKTSHTLLQSQPIASSQQD 163
>AV420667
Length = 392
Score = 27.3 bits (59), Expect = 7.3
Identities = 18/53 (33%), Positives = 27/53 (49%), Gaps = 9/53 (16%)
Frame = +3
Query: 53 YSKYLSGLETPLKNVVDGEFPKRDLVGD---------RVKFGERLTELQAELV 96
+S+Y+SG+ PLK +DGE R L + + +R EL A+LV
Sbjct: 222 HSQYVSGVLRPLKVFLDGERTMRYLASETKNEILLCAATEITDRYYELAADLV 380
>TC9631 similar to GB|AAC19270.1|3193286|T14P8 coded for by A. thaliana
cDNA H76583 {Arabidopsis thaliana;}, partial (30%)
Length = 624
Score = 27.3 bits (59), Expect = 7.3
Identities = 18/85 (21%), Positives = 33/85 (38%), Gaps = 13/85 (15%)
Frame = -3
Query: 527 SYYFNVFLLQRRGYQNR-------------HTPDNIDSDDEESEFTPLKGSFGHQTDKSN 573
S+YF+++ L R+ + R T + I+ P + TDK +
Sbjct: 433 SFYFHIYNLPRKSQKERMKSWINAEMQNSNKTSNEINQSSVHGHIGPSSRNMAMVTDKCS 254
Query: 574 SFTLLSPKKPQPTPHTKGRQLVNGL 598
+ L KP+ P + R + G+
Sbjct: 253 NHRLPIIMKPETPPQAESRVQIPGV 179
>AV777242
Length = 407
Score = 27.3 bits (59), Expect = 7.3
Identities = 14/48 (29%), Positives = 24/48 (49%)
Frame = +1
Query: 215 SDGSKSCANDDANDSGDDVLMLDPSSVNRESFGRKRKRDDLMSEMQSW 262
++G +SC D+ S DDV+ +N++SFG + E+ W
Sbjct: 247 NNGYRSCQVDNTK-SRDDVMRTRIEKLNQKSFGSSSSTPVMCLEISKW 387
>TC10476
Length = 649
Score = 27.3 bits (59), Expect = 7.3
Identities = 12/37 (32%), Positives = 22/37 (59%)
Frame = -2
Query: 298 LFREAAFLKKDFGSDCEKLSWLAQRMHPSMYDVNLGV 334
++ E + +K+DF S +SW +Q S +V+LG+
Sbjct: 114 IYAEISVMKRDFSSR*WTMSWNSQYPXSSFNEVSLGI 4
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,500,007
Number of Sequences: 28460
Number of extensions: 146991
Number of successful extensions: 583
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 579
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 582
length of query: 600
length of database: 4,897,600
effective HSP length: 95
effective length of query: 505
effective length of database: 2,193,900
effective search space: 1107919500
effective search space used: 1107919500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149204.7


