

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.4 - phase: 0 /pseudo
(640 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
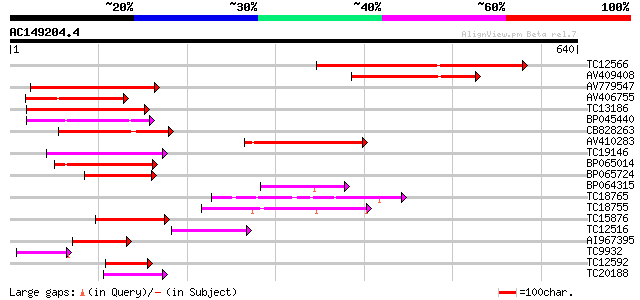
Score E
Sequences producing significant alignments: (bits) Value
TC12566 similar to UP|Q84XI1 (Q84XI1) RCa1 (Fragment), partial (... 193 6e-50
AV409408 173 9e-44
AV779547 142 2e-34
AV406755 129 2e-30
TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidat... 121 4e-28
BP045440 114 4e-26
CB828263 113 8e-26
AV410283 113 1e-25
TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-contain... 102 2e-22
BP065014 100 9e-22
BP065724 87 1e-17
BP064315 84 7e-17
TC18765 weakly similar to UP|Q9LVT1 (Q9LVT1) Disease resistance ... 73 2e-13
TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fr... 70 1e-12
TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 69 2e-12
TC12516 similar to UP|Q9M5V7 (Q9M5V7) Disease resistance-like pr... 65 3e-11
AI967395 59 3e-09
TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like... 57 9e-09
TC12592 54 8e-08
TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 49 3e-06
>TC12566 similar to UP|Q84XI1 (Q84XI1) RCa1 (Fragment), partial (28%)
Length = 918
Score = 193 bits (491), Expect = 6e-50
Identities = 102/239 (42%), Positives = 149/239 (61%), Gaps = 1/239 (0%)
Frame = +1
Query: 347 YKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNID 406
Y+ + D+L LF L AFK + P E Y+DLSK+++ Y G+PLAL+VLG +LYGR
Sbjct: 13 YEVKEMSFEDSLQLFSLNAFKRNDPLETYIDLSKKLLSYAKGIPLALKVLGLFLYGRERK 192
Query: 407 VWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGY 466
W S + KL P P + + LK+SYD LD +KDIFLDIACF+ G + V L+SCG+
Sbjct: 193 AWESQLVKLEKLPDPEIINVLKLSYDGLDDDQKDIFLDIACFYLGHLEESVAQTLDSCGF 372
Query: 467 FPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDR 526
F IG+++L +R LI++ + ++ MHDL++ MG++IV Q+ PNDP +RSRLW+ +
Sbjct: 373 FADIGMEVLKDRCLISI--LEGRIVMHDLIEVMGKEIVRQQCPNDPGKRSRLWNDDQTYD 546
Query: 527 VLTKNKGTEAINSIDMKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQL-PLGLSCLPSSL 584
VL KNKGT+AI I + + + E F + L+ L + ++ L L C PS +
Sbjct: 547 VLGKNKGTDAIQCIFLYTCPIKIVNLHAETFKYMTNLRMLQFYKSRVCSLALECDPSCI 723
Score = 37.4 bits (85), Expect = 0.007
Identities = 17/60 (28%), Positives = 31/60 (51%)
Frame = +2
Query: 573 LPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDITLSHSKIEQLWQGVKVIDSLTKN 632
LP L LP+ L+ L W G PL+ P+ LV + + S +E + +++ +L ++
Sbjct: 713 LPASLGSLPNHLQFLRWDGFPLRYFPLDFCPQNLVKLDMRDSHLEHFGRKIRIYQTLKRH 892
>AV409408
Length = 430
Score = 173 bits (438), Expect = 9e-44
Identities = 86/145 (59%), Positives = 106/145 (72%)
Frame = +2
Query: 387 GGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIA 446
GGLPLALEVLGS+L GR++ W A+ K++ PH + + L+ISYD L+ K +FLDIA
Sbjct: 2 GGLPLALEVLGSFLCGRSLSDWEDALIKIKKVPHDDILNKLRISYDMLEDEHKTLFLDIA 181
Query: 447 CFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRDIVFQ 506
CFFKG KVI +L SCG P +GI +LIE+SL+T D +GMHDLL+EMG+ VF
Sbjct: 182 CFFKGWYKHKVIQLLGSCGLHPTVGINVLIEKSLLTFD--GRVIGMHDLLEEMGKTEVFL 355
Query: 507 ESPNDPCRRSRLWSQEDIDRVLTKN 531
ESPNDP RRSRLWS EDID+VL KN
Sbjct: 356 ESPNDPGRRSRLWSLEDIDQVLRKN 430
>AV779547
Length = 556
Score = 142 bits (357), Expect = 2e-34
Identities = 76/148 (51%), Positives = 92/148 (61%), Gaps = 2/148 (1%)
Frame = +1
Query: 24 LSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAITILSP 83
+SFRGEDTR FTDHL AL KG TFKDD L +G+ IS +LI AI+ S I + S
Sbjct: 109 VSFRGEDTRNNFTDHLLGALYGKGFVTFKDDTMLRKGKNISTELIQAIEGSQILIVVFSK 288
Query: 84 DYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHS 143
YASSTWCL EL I + VLPV V PS+VR Q G + EAF KH+E+F +
Sbjct: 289 YYASSTWCLQELAKIADGIVGKRQTVLPVVCDVTPSEVRKQSGNYGEAFLKHEERFKEDL 468
Query: 144 DRVDRWRDAFTQVASYSGWD--SKGQHE 169
+ V RWR A +VA SGWD +K +HE
Sbjct: 469 EMVQRWRKALAEVADLSGWDVTNKPEHE 552
>AV406755
Length = 425
Score = 129 bits (323), Expect = 2e-30
Identities = 64/116 (55%), Positives = 80/116 (68%)
Frame = +3
Query: 19 TYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAI 78
+Y VFLSFRGE++R+ FT HL AL+ GI F D+ +L+RG+ IS L+ AI+DS AI
Sbjct: 81 SYDVFLSFRGEESRRSFTSHLYTALKNAGIKVFMDN-ELQRGEDISSSLLKAIEDSRIAI 257
Query: 79 TILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRK 134
I S +Y S WCLDEL+ I+EC V+PVFY VDPSD+R QRG EAFRK
Sbjct: 258 IIFSTNYTGSKWCLDELEKIIECQRTIGQEVMPVFYNVDPSDIRKQRGTVGEAFRK 425
Score = 33.5 bits (75), Expect = 0.11
Identities = 14/21 (66%), Positives = 18/21 (85%)
Frame = +3
Query: 487 NNKLGMHDLLQEMGRDIVFQE 507
NNKL MHDLL+EMGR I+ ++
Sbjct: 3 NNKLQMHDLLKEMGRGIIVKK 65
>TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (6%)
Length = 586
Score = 121 bits (303), Expect = 4e-28
Identities = 64/138 (46%), Positives = 87/138 (62%)
Frame = +1
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
Y VF+S DTR GF+ L AL KGI TF DD ++S AI+ S AI
Sbjct: 103 YDVFISLITSDTRFGFSGDLYTALSDKGILTFIDDGLPRPDDIMSTVNSKAIESSRIAIV 282
Query: 80 ILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKF 139
++S +YASS++CLDEL I++ + L + PVFY V+PSDVR+Q+G F +AF H+E+F
Sbjct: 283 VISENYASSSFCLDELMYILKRREEKGLLIWPVFYQVEPSDVRYQKGGFGKAFASHEERF 462
Query: 140 GQHSDRVDRWRDAFTQVA 157
G S +V +WR+A QVA
Sbjct: 463 GSDSAKVRKWRNALKQVA 516
>BP045440
Length = 535
Score = 114 bits (286), Expect = 4e-26
Identities = 66/146 (45%), Positives = 87/146 (59%), Gaps = 2/146 (1%)
Frame = -3
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
Y +F+SFRG DTR FT L L R+G TFKDD+ LE G I EKL++ I++S FAI
Sbjct: 527 YQIFISFRGMDTRDDFTKPLYQGLCREGFVTFKDDESLEGGSPI-EKLLDDIEESRFAII 351
Query: 80 ILSPDYASSTWCLDELQMIMECSSK--NNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQE 137
ILS +YA S WCL EL I+EC + N VLP V P++VRH + +A KH E
Sbjct: 350 ILSKNYAESEWCLKELSKILECKDQEGKNQLVLPSSTEVTPTEVRHVINRYADAMAKH-E 174
Query: 138 KFGQHSDRVDRWRDAFTQVASYSGWD 163
K G S + W+ A V + +G++
Sbjct: 173 KNGIDSKTIQAWKKALFDVCTLTGFE 96
>CB828263
Length = 570
Score = 113 bits (283), Expect = 8e-26
Identities = 56/130 (43%), Positives = 82/130 (63%), Gaps = 1/130 (0%)
Frame = +2
Query: 56 DLERGQVISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYG 115
DL+RG IS L+ AI+++ ++ + S +Y +S WCLDEL I+EC VLPVFY
Sbjct: 5 DLQRGDEISSSLLRAIEEAKLSVIVFSKNYGNSKWCLDELVKILECKRTRGQIVLPVFYD 184
Query: 116 VDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWD-SKGQHEASLVE 174
+DPS VR+Q G + EAF KH + D+V +WR+A + A+ SGWD S + E+ ++E
Sbjct: 185 IDPSHVRNQTGTYAEAFVKHGQ-----VDKVQKWREALREAANLSGWDCSVNRMESEVIE 349
Query: 175 NIAQHIHRKL 184
IA+ + KL
Sbjct: 350 KIAKDVLEKL 379
>AV410283
Length = 426
Score = 113 bits (282), Expect = 1e-25
Identities = 63/138 (45%), Positives = 88/138 (63%)
Frame = +1
Query: 266 QRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPG 325
Q ++L LS S +D+ DG I+ L KVLL+LDDV+E+ QL+ L+G ++WF G
Sbjct: 1 QNRILGDLS-SGGTVNDVNDGVSAIKRVLQGNKVLLILDDVDEIQQLDFLMGNREWFHKG 177
Query: 326 SRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDY 385
SRV+ITTR+ +L V Y+ L AL LFC A + KP EG+ +LSK++V
Sbjct: 178 SRVVITTRNTQVLPESYVDMFYEVRELELSAALALFCHHAMRRKKPAEGFSNLSKQIVKK 357
Query: 386 CGGLPLALEVLGSYLYGR 403
GGLPLALEV+GS+L+ +
Sbjct: 358 TGGLPLALEVIGSFLFDK 411
>TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing protein
TSDC, partial (12%)
Length = 701
Score = 102 bits (254), Expect = 2e-22
Identities = 51/138 (36%), Positives = 79/138 (56%), Gaps = 1/138 (0%)
Frame = +1
Query: 42 ALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMEC 101
AL R+G TF DD+ L G I++ L++A + S +I + S +Y S+WCLDEL I+EC
Sbjct: 10 ALRREGFKTFMDDEGLVGGNEIAKSLLDASEKSRLSIIVFSENYGYSSWCLDELVKIVEC 189
Query: 102 SSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSG 161
V P+FY V+ SDV Q +E A H++ +G+ SD V +WR A + +A G
Sbjct: 190 METKKQLVWPIFYKVEKSDVTSQENSYEHAMIGHEKNYGKDSDEVKKWRSALSAMAKLEG 369
Query: 162 WDSK-GQHEASLVENIAQ 178
+ K ++E ++ I +
Sbjct: 370 HNVKENKYEYEFIKKIVE 423
>BP065014
Length = 498
Score = 100 bits (248), Expect = 9e-22
Identities = 49/116 (42%), Positives = 72/116 (61%)
Frame = -2
Query: 51 FKDDKDLERGQVISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVL 110
FKDD LE G I E+L+ AI++S AI +LS ++A S WCL EL I++C K VL
Sbjct: 497 FKDDLSLEDGAPI-EELLGAIEESRVAIVVLSENFADSKWCLKELVKILDCRKKKKPLVL 321
Query: 111 PVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG 166
P+FY V+P+++RH +G + EA +H++K G+ S V W A + ++ G KG
Sbjct: 320 PIFYKVEPTNIRHLKGRYGEAMAEHEKKLGKDSQTVREWTQALSDISYLKGEPYKG 153
>BP065724
Length = 495
Score = 86.7 bits (213), Expect = 1e-17
Identities = 36/81 (44%), Positives = 55/81 (67%)
Frame = -1
Query: 85 YASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSD 144
YA S+WCL+E+ I+EC KN+ V P+FY V+PSDVRHQR +E+A K + ++G+ S+
Sbjct: 495 YADSSWCLEEVVKILECKQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKQETRYGKDSE 316
Query: 145 RVDRWRDAFTQVASYSGWDSK 165
+V +WR + +QV + K
Sbjct: 315 KVKKWRSSLSQVCDLKAFHYK 253
>BP064315
Length = 350
Score = 84.0 bits (206), Expect = 7e-17
Identities = 47/104 (45%), Positives = 60/104 (57%), Gaps = 4/104 (3%)
Frame = +1
Query: 284 YDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHG- 342
+ G I+ L K+VLLVLDDV+++ QL+ L G DWFG GSR+IITTRD +L H
Sbjct: 37 FRGDFEIKRRLSNKRVLLVLDDVDKIKQLQALAGGGDWFGSGSRIIITTRDTTMLDKHKM 216
Query: 343 ---VHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVV 383
+ K YK L HD+L LFC AF P E + +S V
Sbjct: 217 DDVIMKKYKMEELNDHDSLELFCWHAFNMSTPAENFEGVSSHAV 348
>TC18765 weakly similar to UP|Q9LVT1 (Q9LVT1) Disease resistance
protein-like, partial (18%)
Length = 688
Score = 72.8 bits (177), Expect = 2e-13
Identities = 73/233 (31%), Positives = 114/233 (48%), Gaps = 13/233 (5%)
Frame = +2
Query: 229 KSTIARAVY--ETIRCEF-ELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYD 285
K+T+AR V + +RC F E FL +S++ + L+ ++ H+ +R + Y
Sbjct: 2 KTTLAREVCRDDQVRCHFKERILFLT----VSQSPNVEELRAKIFGHIMGNRG-LNANYA 166
Query: 286 GKKTIQNSLCRKK--VLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGV 343
+ + C+ + +L+VLDDV L LE LV + PG + ++ +R K +
Sbjct: 167 VPQWMPQFECQSQSQILVVLDDVWSLPVLEQLVLRV----PGCKYLVVSRFKFQRI---F 325
Query: 344 HKTYKTGMLCKHDALVLFCLKAFKGDK--PQEGYLDLSKEVVDYCGGLPLALEVLGSYLY 401
+ TY +L + DAL LFC AF G K P +L K+VV CG LPLAL+V+G+ L
Sbjct: 326 NDTYDVELLSEGDALSLFCHHAF-GHKSIPFGANQNLIKQVVAECGRLPLALKVIGASLR 502
Query: 402 GRNIDVWHSAVKKLR------SFPHPRVQDNLKISYDSLDTMEKDIFLDIACF 448
+N W S +L + D + IS + L K+ FLD+ F
Sbjct: 503 DQNEMFWLSVKTRLSQGLSIGESYEVNLIDRMAISTNYLPEKVKECFLDLCAF 661
>TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fragment),
complete
Length = 880
Score = 69.7 bits (169), Expect = 1e-12
Identities = 62/206 (30%), Positives = 92/206 (44%), Gaps = 14/206 (6%)
Frame = +3
Query: 217 RFIGIWGMGGIGKSTIARAVYET-IRCEFELTCFLENVREISETNGLVH-LQRQLLSH-- 272
+ I + GMGG+GK+T+ + VY+ + + C V + E L+ L RQL S
Sbjct: 261 KVISVTGMGGMGKTTLVKQVYDDPVVIKHFRACAWITVSQSCEIGELLRDLARQLFSEIR 440
Query: 273 ----LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRV 328
L + L K I++ L R++ L+V DDV + + E + GSR+
Sbjct: 441 RPVPLGLENMRCDRL---KMIIKDLLQRRRYLVVFDDVWHVREWEAVKYALPDNNCGSRI 611
Query: 329 IITTRDKHLLMTHGVH---KTYKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDY 385
+ITTR L T K Y L + +A LFC K F GD + + ++
Sbjct: 612 MITTRRSDLAFTSSTESKGKVYNLQPLKEDEAWELFCRKTFHGDSCPSHLIGICTYILRK 791
Query: 386 CGGLPLALEVLGSYLY---GRNIDVW 408
C GLPLA+ + L R ID W
Sbjct: 792 CEGLPLAIVAISGVLATKDKRRIDEW 869
>TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (13%)
Length = 558
Score = 68.9 bits (167), Expect = 2e-12
Identities = 31/85 (36%), Positives = 53/85 (61%), Gaps = 2/85 (2%)
Frame = +1
Query: 98 IMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVA 157
I+EC KN+ V P+FY V+PSDVRHQR +E+A K + ++G+ S++V +WR + +QV
Sbjct: 16 ILECXQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKPETRYGKDSEKVKKWRSSLSQVC 195
Query: 158 SYSGWDSK--GQHEASLVENIAQHI 180
+ K +E + + +I + +
Sbjct: 196 DLKAFHYKENSGYERAFIPDIIKDV 270
>TC12516 similar to UP|Q9M5V7 (Q9M5V7) Disease resistance-like protein
(Fragment), partial (36%)
Length = 859
Score = 65.1 bits (157), Expect = 3e-11
Identities = 33/91 (36%), Positives = 50/91 (54%)
Frame = +3
Query: 183 KLVPKLPSCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRC 242
K + L T+ L+G+ + + L DVR IGIWGMGGIGK+TIA ++
Sbjct: 510 KYIATLVPNTKELIGVDKSIAHLESLLCQESKDVRVIGIWGMGGIGKTTIAEEIFNRRHS 689
Query: 243 EFELTCFLENVREISETNGLVHLQRQLLSHL 273
++E CFL V++ +G+ L+ +L S L
Sbjct: 690 QYEGICFLSKVKDEL*KHGMQSLREKLFSTL 782
>AI967395
Length = 205
Score = 58.5 bits (140), Expect = 3e-09
Identities = 29/67 (43%), Positives = 41/67 (60%)
Frame = +3
Query: 71 IKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEE 130
I++S I + S YA S CL+EL I++ NN V PV+YGV+ ++RHQRG + E
Sbjct: 3 IQESRLIIVVFSEHYAVSASCLNELVNIVDVMIMNNQLVWPVYYGVEAPEIRHQRGRYGE 182
Query: 131 AFRKHQE 137
A K +E
Sbjct: 183 AMAKFEE 203
>TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like, partial
(5%)
Length = 689
Score = 57.0 bits (136), Expect = 9e-09
Identities = 30/65 (46%), Positives = 39/65 (59%), Gaps = 3/65 (4%)
Frame = -2
Query: 8 SCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVIS--- 64
S SSS+ Y VF++FRG+DTR F HL AL G+ F DD++LERG +
Sbjct: 238 SMSSSSSNHQWIYDVFINFRGKDTRHSFVSHLYTALSNAGVNVFLDDENLERGLELEPEL 59
Query: 65 EKLIN 69
+KLIN
Sbjct: 58 QKLIN 44
>TC12592
Length = 534
Score = 53.9 bits (128), Expect = 8e-08
Identities = 22/53 (41%), Positives = 35/53 (65%)
Frame = +3
Query: 109 VLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSG 161
VLPVFYGV+P+D+R+ +G F EA + + KFG+ S+ + W+ A ++ G
Sbjct: 9 VLPVFYGVEPTDIRNLKGKFGEAMAEIENKFGKDSNTIHEWKQALRDISYLKG 167
>TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (11%)
Length = 494
Score = 48.5 bits (114), Expect = 3e-06
Identities = 22/74 (29%), Positives = 42/74 (56%), Gaps = 1/74 (1%)
Frame = +1
Query: 106 NLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWD-S 164
N V P+FY + SDV +Q + A H+ FG+ S++V +W+ A +++A G +
Sbjct: 1 NQLVWPIFYEISXSDVSNQTKSYGHAMLAHENSFGKDSEKVQKWKSALSEMAFLQGEHIT 180
Query: 165 KGQHEASLVENIAQ 178
+ ++E L++ I +
Sbjct: 181 ENEYEYKLIQKIVE 222
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.137 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,126,663
Number of Sequences: 28460
Number of extensions: 175879
Number of successful extensions: 956
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 930
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 945
length of query: 640
length of database: 4,897,600
effective HSP length: 96
effective length of query: 544
effective length of database: 2,165,440
effective search space: 1177999360
effective search space used: 1177999360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149204.4


