

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.13 - phase: 0
(163 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
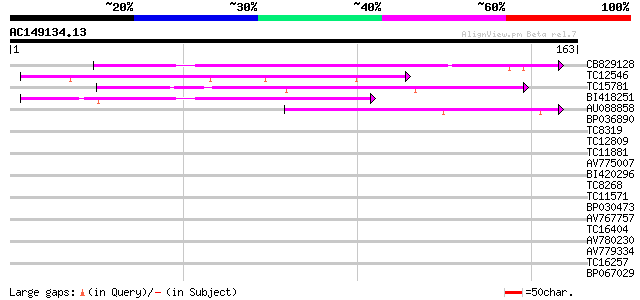
Score E
Sequences producing significant alignments: (bits) Value
CB829128 67 2e-12
TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat ... 58 7e-10
TC15781 50 1e-07
BI418251 49 3e-07
AU088858 46 4e-06
BP036890 30 0.27
TC8319 weakly similar to UP|RK4_TOBAC (O80361) 50S ribosomal pro... 29 0.35
TC12809 similar to UP|Q9ZR88 (Q9ZR88) Bifunctional nuclease (Fra... 27 1.3
TC11881 similar to PIR|T48303|T48303 meiosis specific-like prote... 27 2.3
AV775007 26 3.0
BI420296 26 3.0
TC8268 similar to UP|RS8_ORYSA (P49199) 40S ribosomal protein S8... 26 3.0
TC11571 similar to UP|Q940U9 (Q940U9) AT5g17440/K3M16_10, partia... 26 3.9
BP030473 25 5.1
AV767757 25 5.1
TC16404 similar to UP|P93320 (P93320) Cdc2MsC protein, partial (... 25 6.6
AV780230 25 6.6
AV779334 25 6.6
TC16257 similar to UP|Q9FJD4 (Q9FJD4) Importin beta, partial (16%) 25 6.6
BP067029 25 6.6
>CB829128
Length = 463
Score = 67.0 bits (162), Expect = 2e-12
Identities = 54/138 (39%), Positives = 73/138 (52%), Gaps = 3/138 (2%)
Frame = +3
Query: 25 DIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFVNLP 84
D+ +L AIQR VENL I+ + S +P + TL ILKL +T + V LP
Sbjct: 21 DLKVWLNAAIQRQVENLEIELTDS-----QMPCSILRCTTLVILKLDSVTFDAFTSVYLP 185
Query: 85 SLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVELPYSERPVISLSNLIRANI-C 143
SLK L+L V + ++ LL GCPIL+ L T + + E V +LS L+RA++
Sbjct: 186 SLKTLHLFEVDLVKAQCLIDLLYGCPILEDLKTQIYFRDGSF-EGKVKTLSKLVRADVFL 362
Query: 144 DIH--IEFDWLQNVERLR 159
D H I QNVE LR
Sbjct: 363 DGHFIIPVKAFQNVEFLR 416
>TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat shock
transcription factor HSF30, partial (9%)
Length = 755
Score = 58.2 bits (139), Expect = 7e-10
Identities = 40/119 (33%), Positives = 64/119 (53%), Gaps = 7/119 (5%)
Frame = +2
Query: 4 RDITLPILSFHLQ----CWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLP-SF 58
RD LPI F+L+ C YD ++ ++ +QR +E+L+I + + S S
Sbjct: 386 RDFQLPIKKFYLRLNDVCQPFYDPANVSVWVNAVVQRQLEHLDISLPYPMLSTPRANLSS 565
Query: 59 VFSSKTLSILKLKQ-ITLNEVPFVNLPSLKALYLDVVTFTY-YELILKLLSGCPILQYL 115
+FS +TL +LKL+ + L P V+ P LK L+L + + +LLSGCP+L+ L
Sbjct: 566 IFSCRTLVVLKLRGGLELKRFPSVHFPCLKVLHLQGALLLHDVPYLAELLSGCPVLENL 742
>TC15781
Length = 621
Score = 50.4 bits (119), Expect = 1e-07
Identities = 43/128 (33%), Positives = 66/128 (50%), Gaps = 4/128 (3%)
Frame = +2
Query: 26 IYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEV--PFVNL 83
++++L + R +ENL I L + L ++S +TL +LKLK L V L
Sbjct: 122 VHDWLNTVMPRRIENLEICIP-PLSRGIGLK--IYSCRTLVVLKLKGYPLGVYVSSHVEL 292
Query: 84 PSLKALYLDVVTFTYYELILKLLSGCPILQYL--GTNNLVVELPYSERPVISLSNLIRAN 141
PSLK+L+L+ V F ++ LKL+ GCP+L+ L GT + + E SL L+
Sbjct: 293 PSLKSLHLEKVHFVDFQSFLKLICGCPLLEDLEAGTVSYLDTSFDQEFRFRSLPKLVSVV 472
Query: 142 ICDIHIEF 149
I D + F
Sbjct: 473 INDYNFRF 496
>BI418251
Length = 546
Score = 49.3 bits (116), Expect = 3e-07
Identities = 37/104 (35%), Positives = 57/104 (54%), Gaps = 2/104 (1%)
Frame = +1
Query: 4 RDITLPILSFHLQCWNDYDCR--DIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFS 61
R + PI +F L ++ +C D+ +L A+QR VENL+I S S TLP + S
Sbjct: 253 RALHQPIKNFTLSVISE-ECPYPDVKVWLNAAMQRQVENLDICLSLS-----TLPCSILS 414
Query: 62 SKTLSILKLKQITLNEVPFVNLPSLKALYLDVVTFTYYELILKL 105
TL +LKL + + V+LPSLK L+L++V + ++ L
Sbjct: 415 CTTLVVLKLSDVKFHVFSCVDLPSLKTLHLELVVILNPQSLMDL 546
>AU088858
Length = 765
Score = 45.8 bits (107), Expect = 4e-06
Identities = 33/84 (39%), Positives = 47/84 (55%), Gaps = 4/84 (4%)
Frame = +3
Query: 80 FVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNNLVVE--LPYSERPVISLSNL 137
FV+LPSLK L L + F + +++LLSGCPIL+Y+ + + S V SL+ L
Sbjct: 12 FVHLPSLKTLQLYCLIFPEPQYVMELLSGCPILEYMHLSRMRYADCSSPSNTNVKSLTKL 191
Query: 138 IRANICDIHIEFDW--LQNVERLR 159
+ A+I I E + NVE LR
Sbjct: 192 VSADISFIGFEIPLKVICNVEFLR 263
>BP036890
Length = 454
Score = 29.6 bits (65), Expect = 0.27
Identities = 14/36 (38%), Positives = 24/36 (65%)
Frame = +1
Query: 42 NIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNE 77
+I++++ LFS +T +F+FS LSI K+ + L E
Sbjct: 49 SINYTNILFSSLTCSTFIFSKMKLSIHKINKKLLCE 156
>TC8319 weakly similar to UP|RK4_TOBAC (O80361) 50S ribosomal protein L4,
chloroplast precursor (R-protein L4), partial (30%)
Length = 650
Score = 29.3 bits (64), Expect = 0.35
Identities = 16/39 (41%), Positives = 20/39 (51%), Gaps = 4/39 (10%)
Frame = -1
Query: 21 YDCRD----IYNFLYIAIQRGVENLNIDFSHSLFSQMTL 55
YDCR I NFLY +QR ++ LN H Q+ L
Sbjct: 425 YDCRPSTPCISNFLYKMVQRRLQTLNSPHQHHPLCQVLL 309
>TC12809 similar to UP|Q9ZR88 (Q9ZR88) Bifunctional nuclease (Fragment),
partial (16%)
Length = 471
Score = 27.3 bits (59), Expect = 1.3
Identities = 18/54 (33%), Positives = 29/54 (53%), Gaps = 2/54 (3%)
Frame = -1
Query: 30 LYIAIQRGVENLNIDFSHSLFSQMTLPSFV--FSSKTLSILKLKQITLNEVPFV 81
L+ I+ + NL + SL + TLP + KTL +L QI+L+ +PF+
Sbjct: 366 LFSTIKSWISNLILSSQ*SLVNYSTLPKILSLVPLKTLIVLIELQISLHLIPFL 205
>TC11881 similar to PIR|T48303|T48303 meiosis specific-like protein -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(34%)
Length = 631
Score = 26.6 bits (57), Expect = 2.3
Identities = 26/115 (22%), Positives = 53/115 (45%), Gaps = 10/115 (8%)
Frame = +2
Query: 1 MAMRDITLPILSFHLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVF 60
+ ++D++LP + CR++ + ++Q +E+L +HS+ +T F F
Sbjct: 218 LTLQDLSLP-----------FRCREVADLSLSSVQSNIESLARKIAHSI---LTGNGFAF 355
Query: 61 ------SSKTLSILKLKQITLNEV----PFVNLPSLKALYLDVVTFTYYELILKL 105
S+ L + +L +I L + PF N+ +++ +T ELI +L
Sbjct: 356 DVPSRSSANQLYVPELDRIVLKDKSSLRPFGNISTVRK---STITARILELIHQL 511
>AV775007
Length = 469
Score = 26.2 bits (56), Expect = 3.0
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = -1
Query: 74 TLNEVPFVNLPSLKALYLDVVTFTYYELILKLLSGCPILQYLGTNN 119
TL +N P + V F+ + L+LL+GCP L+ G+ +
Sbjct: 445 TLRIFLVLNFPHSQLCI*TVFIFSMLNI*LELLNGCPNLEIFGSTS 308
>BI420296
Length = 592
Score = 26.2 bits (56), Expect = 3.0
Identities = 14/38 (36%), Positives = 22/38 (57%)
Frame = +2
Query: 33 AIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKL 70
A+ V++L I+F+ L L +FS +TL+ LKL
Sbjct: 479 AVSHDVQSLTIEFNLYLKLGFKLHPCIFSCRTLTYLKL 592
>TC8268 similar to UP|RS8_ORYSA (P49199) 40S ribosomal protein S8, partial
(95%)
Length = 765
Score = 26.2 bits (56), Expect = 3.0
Identities = 11/30 (36%), Positives = 18/30 (59%)
Frame = -2
Query: 107 SGCPILQYLGTNNLVVELPYSERPVISLSN 136
S C +L N +++E+P ER I+L+N
Sbjct: 464 SSCTLLLATKVNTIMLEIPLLERSCINLNN 375
>TC11571 similar to UP|Q940U9 (Q940U9) AT5g17440/K3M16_10, partial (67%)
Length = 1011
Score = 25.8 bits (55), Expect = 3.9
Identities = 23/75 (30%), Positives = 33/75 (43%), Gaps = 9/75 (12%)
Frame = -1
Query: 14 HLQCWNDYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPS---------FVFSSKT 64
HL+C + + R+ L + R + L I SHSL S S F F+S +
Sbjct: 516 HLRCLSYFRNRNCIAALASSSSRRLSALPIFLSHSLISLSMTSSNSLS*LSAPFAFAS-S 340
Query: 65 LSILKLKQITLNEVP 79
S L K+ TL + P
Sbjct: 339 YSFLSCKECTLGQGP 295
>BP030473
Length = 532
Score = 25.4 bits (54), Expect = 5.1
Identities = 15/43 (34%), Positives = 26/43 (59%), Gaps = 3/43 (6%)
Frame = -2
Query: 76 NEVPFV---NLPSLKALYLDVVTFTYYELILKLLSGCPILQYL 115
+EV F +P L+ L++ + F+ E + +L GCP+L+YL
Sbjct: 480 DEVAFAIAGTMPKLRHLHVALNRFSN-EGLDAILDGCPLLEYL 355
>AV767757
Length = 494
Score = 25.4 bits (54), Expect = 5.1
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +2
Query: 26 IYNFLYIAIQRGVENLNIDFSHSLFS 51
++NF + QR V ++ I FSHSLF+
Sbjct: 212 VFNFTIGSEQRIVFHVAIQFSHSLFN 289
>TC16404 similar to UP|P93320 (P93320) Cdc2MsC protein, partial (15%)
Length = 584
Score = 25.0 bits (53), Expect = 6.6
Identities = 15/76 (19%), Positives = 35/76 (45%)
Frame = -1
Query: 20 DYDCRDIYNFLYIAIQRGVENLNIDFSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVP 79
+Y R +++F + RG+++ H F ++ + ++ + LSI + I P
Sbjct: 488 NYKIRGLFHFCGPLMTRGIKHYLRCMHHKGFQELQVMGILYLNILLSIETISHIFSGT*P 309
Query: 80 FVNLPSLKALYLDVVT 95
++L + +Y + T
Sbjct: 308 -IDLSDISQIYQSITT 264
>AV780230
Length = 583
Score = 25.0 bits (53), Expect = 6.6
Identities = 15/52 (28%), Positives = 24/52 (45%), Gaps = 6/52 (11%)
Frame = +3
Query: 12 SFHLQCWNDYDCRDIYNFLYIAI------QRGVENLNIDFSHSLFSQMTLPS 57
+FH+QC D D++ F +I Q+ N +I F+ L Q P+
Sbjct: 75 TFHIQCIQDKRTSDLFFFSFILSP*DLNKQKSRRNCDIYFTTELQKQSAPPT 230
>AV779334
Length = 596
Score = 25.0 bits (53), Expect = 6.6
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = -1
Query: 8 LPILSFHLQCWNDYDCRDIYNF 29
+P+ FHLQC N Y +++ F
Sbjct: 251 VPLAGFHLQCQNHYLRFELWRF 186
>TC16257 similar to UP|Q9FJD4 (Q9FJD4) Importin beta, partial (16%)
Length = 562
Score = 25.0 bits (53), Expect = 6.6
Identities = 14/40 (35%), Positives = 18/40 (45%)
Frame = +3
Query: 45 FSHSLFSQMTLPSFVFSSKTLSILKLKQITLNEVPFVNLP 84
FSHSL P F S S+ + + +PFVN P
Sbjct: 12 FSHSLARVSESPRFSLPSTRSSLTH*LTVLPSPIPFVNSP 131
>BP067029
Length = 499
Score = 25.0 bits (53), Expect = 6.6
Identities = 10/31 (32%), Positives = 18/31 (57%)
Frame = -2
Query: 114 YLGTNNLVVELPYSERPVISLSNLIRANICD 144
++G L++ LP +PV+ ++ AN CD
Sbjct: 297 FIGVVVLLLLLPIPAKPVLGMTRDFDANFCD 205
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.142 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,951,908
Number of Sequences: 28460
Number of extensions: 38514
Number of successful extensions: 335
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 333
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 333
length of query: 163
length of database: 4,897,600
effective HSP length: 83
effective length of query: 80
effective length of database: 2,535,420
effective search space: 202833600
effective search space used: 202833600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC149134.13


