

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
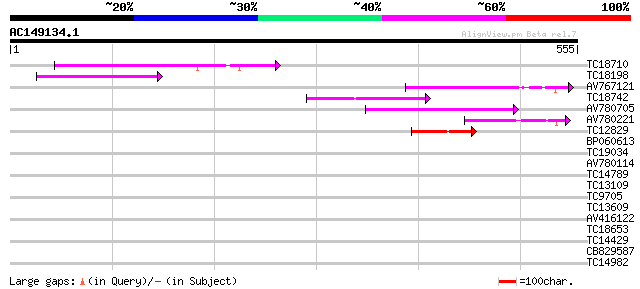
Query= AC149134.1 - phase: 0
(555 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18710 133 7e-32
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 94 8e-20
AV767121 93 1e-19
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 72 2e-13
AV780705 63 1e-10
AV780221 45 3e-05
TC12829 42 3e-04
BP060613 36 0.019
TC19034 32 0.21
AV780114 29 1.7
TC14789 similar to UP|O04873 (O04873) L-ascorbate peroxidase, ch... 28 3.9
TC13109 similar to UP|Q8L4J2 (Q8L4J2) Cleavage stimulation facto... 28 3.9
TC9705 similar to UP|Q8LDM2 (Q8LDM2) Chalcone synthase-like prot... 27 6.6
TC13609 27 8.7
AV416122 27 8.7
TC18653 similar to UP|Q8LFK0 (Q8LFK0) Beta-N-acetylhexosaminidas... 27 8.7
TC14429 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan pro... 27 8.7
CB829587 27 8.7
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 27 8.7
>TC18710
Length = 843
Score = 133 bits (335), Expect = 7e-32
Identities = 77/226 (34%), Positives = 120/226 (53%), Gaps = 5/226 (2%)
Frame = +2
Query: 45 KEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSI 104
+++++ F L + G D F+Q ++VK V A+L F + +PN N +
Sbjct: 137 QKKLRLLFFLLETSQPRGMDGMNGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETLV 316
Query: 105 ILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDC 164
LIPK P+ +S++Q+R I+ F +K+I+K+ RL +P +IS Q GF+QGR I+D
Sbjct: 317 TLIPKVPHPESINQFRPISGCTFLYKVISKIFVARLKDAMPDLISPMQSGFIQGRQIQDN 496
Query: 165 IALTSEAINVLDNKSFGG--NLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTI 222
+ + EA + ++ G + +K+D+ KA+D + W FL L FGF+ NW+K I
Sbjct: 497 LLIVQEAFHAINRPGALGRNHSIIKLDMNKAYDRVEWKFLESSLLAFGFS---TNWVKMI 667
Query: 223 L---HSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLS 265
+ +NG RG+RQGDP SP LF EVLS
Sbjct: 668 MILVSGVSYNYKINGVVGPKLLPQRGLRQGDPFSPYLFLFTMEVLS 805
Score = 30.4 bits (67), Expect = 0.78
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = +1
Query: 25 EAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPG 62
+ +P LV N+ L T EE+K VF L APG
Sbjct: 76 DVVPSLVSQDLNQKLESQVTSEEIKAVVFSLGDLTAPG 189
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 93.6 bits (231), Expect = 8e-20
Identities = 49/123 (39%), Positives = 72/123 (57%)
Frame = -3
Query: 27 IPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAV 86
IPKLV N L + +E+K AVF + APG D F+Q IVK DV A+
Sbjct: 377 IPKLVTDRLNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQQNREIVKNDVNNAI 198
Query: 87 LDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPS 146
LDFF +G LP N + LIPK P+A+++ Q+R I+ +F +K+I+K++ RL + +
Sbjct: 197 LDFFDHGELPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKIIVARLKPDMVN 18
Query: 147 IIS 149
+IS
Sbjct: 17 LIS 9
>AV767121
Length = 525
Score = 92.8 bits (229), Expect = 1e-19
Identities = 55/173 (31%), Positives = 87/173 (49%), Gaps = 8/173 (4%)
Frame = +2
Query: 388 LAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRK 447
L+KWK +S GRI L++SV+ ++ + +S + PI + K + ++NF+W G + K
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 448 MVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWN-LMQSDEQWANIIRSRVIRDHRC 506
+ V W +C E GGLG+K L N+A K W L + D W +I ++ DH
Sbjct: 182 IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQ--PDH-- 349
Query: 507 MNHHISSSIWSGVKAEFHVIKDNCSW-------IVGDGKQINFWSDSWCGQVL 552
+ SS W+ + + +D W +VG+G Q FWS+ W G L
Sbjct: 350 --YSCGSSWWNDILS--LCPEDADGWFSSGLKKLVGEGDQTKFWSEDWLGSGL 496
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 72.0 bits (175), Expect = 2e-13
Identities = 41/122 (33%), Positives = 63/122 (51%)
Frame = -3
Query: 291 HCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNNIVNI 350
H + D+LMVF + S+ ++ ++ + SG N KS +F + +T N I N+
Sbjct: 367 HLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSEVFIL-VKETTKNQITNM 191
Query: 351 LGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQ 410
LG+ G LP LG P K K + + D+ +KL W LS AGR+QL+ S++
Sbjct: 190 LGYKEGKLPVR*LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLINSILF 11
Query: 411 SM 412
SM
Sbjct: 10 SM 5
>AV780705
Length = 524
Score = 63.2 bits (152), Expect = 1e-10
Identities = 43/152 (28%), Positives = 72/152 (47%), Gaps = 2/152 (1%)
Frame = -2
Query: 349 NILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSV 408
N+L + P YLG P G+ K I +KVK KL WK +LL+ AG+ L+K++
Sbjct: 523 NLLHMPIWDDPGHYLGLPAIWGRNKSHSLVWIEEKVKEKLEGWKETLLNQAGKEVLIKAI 344
Query: 409 VQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVA-WRKICADYEEGGLGV 467
+Q++ + M+I +P + + +F W T A K E+GG+G
Sbjct: 343 IQAIPSYAMTIVHFPKTFCNSLYALVADFWWKSQGEGAVEYTGAVGTK*PLSKEKGGVGF 164
Query: 468 KSLICLNEATNLKICWNLMQSDEQ-WANIIRS 498
+ L N A + W ++ + E W +++S
Sbjct: 163 RDLRTQNLAFLARQAWRVLTNPEALWVRVMKS 68
>AV780221
Length = 461
Score = 45.1 bits (105), Expect = 3e-05
Identities = 27/112 (24%), Positives = 54/112 (48%), Gaps = 8/112 (7%)
Frame = +3
Query: 446 RKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLM-QSDEQWANIIRSRVIRDH 504
+K+ V W +C +EGGLGV++L N+A K W ++ + D W ++ + +
Sbjct: 138 KKIAWVKWSTVCRPKDEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVL----LIKY 305
Query: 505 RCMNHHISSSIWSGVKAEFHVIKDNCSWI-------VGDGKQINFWSDSWCG 549
+ +SS W+ + + W+ +G+G ++ FW+++W G
Sbjct: 306 KNAIPQSASSWWNDLHSTCFE-DGGGGWMQRGLCRRIGEGTEVKFWNENWLG 458
>TC12829
Length = 448
Score = 42.0 bits (97), Expect = 3e-04
Identities = 19/64 (29%), Positives = 39/64 (60%)
Frame = +1
Query: 394 SLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAW 453
S + G+ L + + QS L +++ + +L+ ++ W+ +FIWSGDV++R + + W
Sbjct: 259 SCIEREGKF*LSR*LKQSQLTSCLALLFLILFVLR-LKAWLVSFIWSGDVSRRSIHWLGW 435
Query: 454 RKIC 457
+K+C
Sbjct: 436 KKLC 447
>BP060613
Length = 378
Score = 35.8 bits (81), Expect = 0.019
Identities = 27/109 (24%), Positives = 46/109 (41%), Gaps = 7/109 (6%)
Frame = +1
Query: 439 WSGDVTKRKMVTVAWRK---ICADYEEGGLGVKSLICLNEATNLKICWNLMQS-DEQWAN 494
WS T+ V WRK + +EGGLG K N+A K W ++ + D W
Sbjct: 49 WSSKGTRG----VHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQ 216
Query: 495 IIRSRVIRDH---RCMNHHISSSIWSGVKAEFHVIKDNCSWIVGDGKQI 540
I+++ H + +S +WS + ++ W +G G+ +
Sbjct: 217 ILKALYFPHHDFLQTTKRTGASWVWSSLLHGRELLLKQGQWQLGSGETV 363
>TC19034
Length = 843
Score = 32.3 bits (72), Expect = 0.21
Identities = 23/72 (31%), Positives = 39/72 (53%)
Frame = -1
Query: 94 WLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQR 153
+L FN+NSI+L N V RT ++V+F F +I ++LA+ L I+ + +
Sbjct: 795 FLLRQFNSNSILLAIIVWNMFLVCIMRTFSIVHFFFVLIYRLLANLLKTIVLFGLKLGSQ 616
Query: 154 GFVQGRNIRDCI 165
GF+ N + C+
Sbjct: 615 GFL--NNFKLCL 586
>AV780114
Length = 535
Score = 29.3 bits (64), Expect = 1.7
Identities = 18/59 (30%), Positives = 29/59 (48%)
Frame = +2
Query: 463 GGLGVKSLICLNEATNLKICWNLMQSDEQWANIIRSRVIRDHRCMNHHISSSIWSGVKA 521
GG V + L+ + + W L++ E ++ V+RDH +N +SI S VKA
Sbjct: 152 GGRNVVVGVKLDSPSKELLTWALVKVAEPGDIVVAVHVLRDHEIVNGDGKTSILSRVKA 328
>TC14789 similar to UP|O04873 (O04873) L-ascorbate peroxidase, chloroplast
precursor , partial (22%)
Length = 615
Score = 28.1 bits (61), Expect = 3.9
Identities = 21/71 (29%), Positives = 35/71 (48%), Gaps = 7/71 (9%)
Frame = +1
Query: 466 GVKSLICLNEATNL-KICWNLMQSDEQWANIIRSRVIRDH---RCMNHHISSS---IWSG 518
G++ IC + L ++C + Q+ + W+ I SR + +C + SS +W G
Sbjct: 4 GIR*EIC*RSGSIL*RLC*SACQT*QPWSQI*SSRGHCNRWCSKCWGKEVCSSRVLLWKG 183
Query: 519 VKAEFHVIKDN 529
VK E HV+ N
Sbjct: 184 VKNECHVLLYN 216
>TC13109 similar to UP|Q8L4J2 (Q8L4J2) Cleavage stimulation factor 50K chain
(Cleavage stimulation factor 50), partial (7%)
Length = 541
Score = 28.1 bits (61), Expect = 3.9
Identities = 14/32 (43%), Positives = 20/32 (61%)
Frame = -3
Query: 198 NWDFLLLVLKTFGFNELFCNWIKTILHSSKMF 229
++DF L+L FG + FC+ I T L S+K F
Sbjct: 368 SYDFQNLLLLGFGLIQYFCSPIDTKLRSTKRF 273
>TC9705 similar to UP|Q8LDM2 (Q8LDM2) Chalcone synthase-like protein,
partial (35%)
Length = 634
Score = 27.3 bits (59), Expect = 6.6
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = -2
Query: 276 LIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIV 311
++ +I +R CLP+ Y L+ F KA LIV
Sbjct: 606 VLSIIICNRGRCLPYKGLYTSFLLSFIKALNLLLIV 499
>TC13609
Length = 356
Score = 26.9 bits (58), Expect = 8.7
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = +3
Query: 2 WFLLIKTCFFLLTIFLQDHLLVEEAIPKLVDATTNRLL 39
+F I CFF L +FL D L+ + + TNRLL
Sbjct: 219 FFPTIFLCFFFLDLFL*DRLIPRKVSIRSEVVGTNRLL 332
>AV416122
Length = 414
Score = 26.9 bits (58), Expect = 8.7
Identities = 13/35 (37%), Positives = 17/35 (48%)
Frame = -3
Query: 513 SSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
S + SGV A F + CSW G G ++ SW
Sbjct: 184 SGLGSGVAASFGALVGGCSWGSGTGWRLEGNVGSW 80
>TC18653 similar to UP|Q8LFK0 (Q8LFK0) Beta-N-acetylhexosaminidase-like
protein, partial (5%)
Length = 431
Score = 26.9 bits (58), Expect = 8.7
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = -2
Query: 497 RSRVIRDHRCMNHHISSS 514
R+++ RDHR NH ISS+
Sbjct: 205 RNKIYRDHRSSNHSISSA 152
>TC14429 weakly similar to UP|Q8LEJ6 (Q8LEJ6) Arabinogalactan protein-like,
partial (19%)
Length = 530
Score = 26.9 bits (58), Expect = 8.7
Identities = 15/58 (25%), Positives = 22/58 (37%)
Frame = +1
Query: 245 GVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFC 302
GV + DP +FC ++ + L C FH +V L+VFC
Sbjct: 307 GVEEEDPFGSCIFCFQCHIVMHPVFFL---------------CFVFHLHFVICLLVFC 435
>CB829587
Length = 548
Score = 26.9 bits (58), Expect = 8.7
Identities = 12/29 (41%), Positives = 16/29 (54%)
Frame = -2
Query: 225 SSKMFISMNGAQHGFFNCNRGVRQGDPLS 253
SS FI NG+Q+G +N N + G S
Sbjct: 325 SSSRFIHQNGSQYGKWNLNTSIGYGTSFS 239
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 26.9 bits (58), Expect = 8.7
Identities = 21/87 (24%), Positives = 40/87 (45%), Gaps = 1/87 (1%)
Frame = +3
Query: 146 SIISKEQRGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLV 205
S + + + F QGRN+R T A+ + D++S G L + D A + + +
Sbjct: 678 SHVMESESLFRQGRNLRRRRPRTRNAMLMSDSRSGNGRLDEEDDHDAAIEV*TLCYFYFL 857
Query: 206 LKTF-GFNELFCNWIKTILHSSKMFIS 231
L F FN +F ++ + + +I+
Sbjct: 858 LNLFLDFNFIFWDFCSKLYYMIHYYIN 938
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,163,305
Number of Sequences: 28460
Number of extensions: 168606
Number of successful extensions: 1087
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1079
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1083
length of query: 555
length of database: 4,897,600
effective HSP length: 95
effective length of query: 460
effective length of database: 2,193,900
effective search space: 1009194000
effective search space used: 1009194000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149134.1


